Featured
-
-
Article
| Open Access Localization of adaptive variants in human genomes using averaged one-dependence estimation
Localization of adaptive variants in human genomes using averaged one-dependence estimation
Selective sweeps are events in which beneficial mutations spread rapidly through a population. Here, Sugden et al. develop SWIF(r), a probabilistic classification framework for detecting and localizing selective sweeps, and apply it to genomic data from the ‡Khomani San.
- Lauren Alpert Sugden
- , Elizabeth G. Atkinson
- & Sohini Ramachandran
-
Article
| Open Access Identifying noncoding risk variants using disease-relevant gene regulatory networks
Identifying noncoding risk variants using disease-relevant gene regulatory networks
Current methods for prioritization of non-coding genetic risk variants are based on sequence and chromatin features. Here, Gao et al. develop ARVIN, which predicts causal regulatory variants using disease-relevant gene-regulatory networks, and validate this approach in reporter gene assays.
- Long Gao
- , Yasin Uzun
- & Kai Tan
-
Article
| Open Access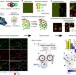 Mapping molecular assemblies with fluorescence microscopy and object-based spatial statistics
Mapping molecular assemblies with fluorescence microscopy and object-based spatial statistics
Elucidating molecular organisation requires precise localisation and analysis. Here the authors develop SODA software for automatic and quantitative mapping of statistically coupled molecules, and use it to unravel spatial organisation of thousands of synaptic proteins in SIM and 3DSTORM microscopy.
- Thibault Lagache
- , Alexandre Grassart
- & Jean-Christophe Olivo-Marin
-
Article
| Open Access A community approach to mortality prediction in sepsis via gene expression analysis
A community approach to mortality prediction in sepsis via gene expression analysis
Sepsis is characterized by deregulated host response to infection. Efficient therapies are still needed but a limitation for sepsis treatment is the heterogeneity in patients. Here Sweeney et al. generate prognostic models based on gene expression to improve risk stratification classification and prediction for 30-day mortality of patients.
- Timothy E. Sweeney
- , Thanneer M. Perumal
- & Raymond J. Langley
-
Article
| Open Access The effects of death and post-mortem cold ischemia on human tissue transcriptomes
The effects of death and post-mortem cold ischemia on human tissue transcriptomes
RNA levels in post-mortem tissue can differ greatly from those before death. Studying the effect of post-mortem interval on the transcriptome in 36 human tissues, Ferreira et al. find that the response to death is largely tissue-specific and develop a model to predict time since death based on RNA data.
- Pedro G. Ferreira
- , Manuel Muñoz-Aguirre
- & Roderic Guigó
-
Article
| Open Access CellCycleTRACER accounts for cell cycle and volume in mass cytometry data
CellCycleTRACER accounts for cell cycle and volume in mass cytometry data
Mass cytometry is a powerful method of single cell analysis, but potential confounding effects of cell cycle and cell volume are not taken into account. Here the authors present a combined experimental and computational method to correct for these effects and reveal features of TNFα stimulation that are otherwise masked.
- Maria Anna Rapsomaniki
- , Xiao-Kang Lun
- & María Rodríguez Martínez
-
Article
| Open Access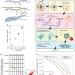 Single-cell full-length total RNA sequencing uncovers dynamics of recursive splicing and enhancer RNAs
Single-cell full-length total RNA sequencing uncovers dynamics of recursive splicing and enhancer RNAs
Total RNA sequencing has been used to profile poly(A) and non-poly(A) RNA expression, processing and the activity of enhancers. Here the authors develop RamDA-seq, a method for full-length total RNA sequencing in single cells.
- Tetsutaro Hayashi
- , Haruka Ozaki
- & Itoshi Nikaido
-
Article
| Open Access Statistical modeling of RNA structure profiling experiments enables parsimonious reconstruction of structure landscapes
Statistical modeling of RNA structure profiling experiments enables parsimonious reconstruction of structure landscapes
Different experimental and computational approaches can be used to study RNA structures. Here, the authors present a computational method for data-directed reconstruction of complex RNA structure landscapes, which predicts a parsimonious set of co-existing structures and estimates their abundances from structure profiling data.
- Hua Li
- & Sharon Aviran
-
Article
| Open Access High-throughput immune repertoire analysis with IGoR
High-throughput immune repertoire analysis with IGoR
B and T cell receptor diversity can be studied by high-throughput immune receptor sequencing. Here, the authors develop a software tool, IGoR, that calculates the likelihoods of potential V(D)J recombination and somatic hypermutation scenarios from raw immune sequence reads.
- Quentin Marcou
- , Thierry Mora
- & Aleksandra M. Walczak
-
Article
| Open Access Optimal compressed representation of high throughput sequence data via light assembly
Optimal compressed representation of high throughput sequence data via light assembly
Increase in high throughput sequencing (HTS) data warrants compression methods to facilitate better storage and communication. Here, Ginart et al. introduce Assembltrie, a reference-free compression tool which is guaranteed to achieve optimality for error-free reads.
- Antonio A. Ginart
- , Joseph Hui
- & David N. Tse
-
Article
| Open Access High contiguity Arabidopsis thaliana genome assembly with a single nanopore flow cell
High contiguity Arabidopsis thaliana genome assembly with a single nanopore flow cell
Long-read sequencing technologies facilitate efficient and high quality genome assembly. Here Michael et al. achieve a fast reference assembly for Arabidopsis thaliana KBS-Mac-74 accession using the handheld Oxford Nanopore MinION sequencer and consumer computing hardware, and demonstrate its usefulness in resolving complex structural variation.
- Todd P. Michael
- , Florian Jupe
- & Joseph R. Ecker
-
Article
| Open Access Stratification of TAD boundaries reveals preferential insulation of super-enhancers by strong boundaries
Stratification of TAD boundaries reveals preferential insulation of super-enhancers by strong boundaries
Topologically associating domains (TADs) detected by Hi-C technologies are megabase-scale areas of highly interacting chromatin. Here Gong, Lazaris et al. develop a computational approach to improve the reproducibility of Hi-C contact matrices and stratify TAD boundaries based on their insulating strength.
- Yixiao Gong
- , Charalampos Lazaris
- & Aristotelis Tsirigos
-
Article
| Open Access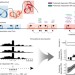 Transcriptional decomposition reveals active chromatin architectures and cell specific regulatory interactions
Transcriptional decomposition reveals active chromatin architectures and cell specific regulatory interactions
Transcriptional regulation is coupled with chromosomal positioning and chromatin architecture. Here the authors develop a transcriptional decomposition approach to separate expression associated with genome structure from independent effects not directly associated with genomic positioning.
- Sarah Rennie
- , Maria Dalby
- & Robin Andersson
-
Article
| Open Access Cell fate in antiviral response arises in the crosstalk of IRF, NF-κB and JAK/STAT pathways
Cell fate in antiviral response arises in the crosstalk of IRF, NF-κB and JAK/STAT pathways
Innate immunity combines intra- and intercellular signalling to develop responses that limit pathogen spread. Here the authors analyse feedback and feedforward loops connecting IRF3, NF-κB and STAT pathways, and suggest they allow coordinating cell fate decisions in cellular populations in response to the virus-mimicking agent poly(I:C).
- Maciej Czerkies
- , Zbigniew Korwek
- & Tomasz Lipniacki
-
Article
| Open Access Probability of phenotypically detectable protein damage by ENU-induced mutations in the Mutagenetix database
Probability of phenotypically detectable protein damage by ENU-induced mutations in the Mutagenetix database
Programs such as PolyPhen-2 predict the relative severity of damage by missense mutations. Here, Wang et al estimate probabilities that putative null or missense alleles would reduce protein function to cause detectable phenotype by analyzing data from ENU-induced mouse mutations.
- Tao Wang
- , Chun Hui Bu
- & Bruce Beutler
-
Article
| Open Access Transcriptomic alterations during ageing reflect the shift from cancer to degenerative diseases in the elderly
Transcriptomic alterations during ageing reflect the shift from cancer to degenerative diseases in the elderly
Ageing is associated with a pronounced shift in mortality from cancer to degenerative diseases. Here, the authors show that in concordance with this shift, conserved transcriptional alterations during ageing across four vertebrates align with degenerative diseases but are opposite to those in cancer.
- Peer Aramillo Irizar
- , Sascha Schäuble
- & Christoph Kaleta
-
Article
| Open Access Subcortical evidence for a contribution of arousal to fMRI studies of brain activity
Subcortical evidence for a contribution of arousal to fMRI studies of brain activity
Resting cortical activity fluctuates, but it is unclear what underlies these variations in activity. Here, the authors show that large-scale fluctuations in fMRI cortical activity are associated with momentary decreases in cortical arousal and opposite activity changes in the basal forebrain and thalamus.
- Xiao Liu
- , Jacco A. de Zwart
- & Jeff H. Duyn
-
Article
| Open Access Automated NMR resonance assignments and structure determination using a minimal set of 4D spectra
Automated NMR resonance assignments and structure determination using a minimal set of 4D spectra
Further automation of NMR structure determination is needed to increase the throughput and accessibility of this method. Here the authors present 4D-CHAINS/autoNOE-Rosetta, a complete pipeline that allows rapid and fully automated structure determination from two highly complementary NMR datasets.
- Thomas Evangelidis
- , Santrupti Nerli
- & Konstantinos Tripsianes
-
Article
| Open Access Single-cell variability in multicellular organisms
Single-cell variability in multicellular organisms
While gene expression noise in single-celled organisms is well understood, it is less so in the context of tissues. Here the authors show that coupling between cells in tissues can increase or decrease cell-to-cell variability depending on the level of noise intrinsic to the regulatory networks.
- Stephen Smith
- & Ramon Grima
-
Article
| Open Access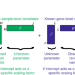 A general and flexible method for signal extraction from single-cell RNA-seq data
A general and flexible method for signal extraction from single-cell RNA-seq data
Single-cell RNA sequencing (scRNA-seq) data provides information on transcriptomic heterogeneity within cell populations. Here, Risso et al develop ZINB-WaVE for low-dimensional representations of scRNA-seq data that account for zero inflation, over-dispersion, and the count nature of the data.
- Davide Risso
- , Fanny Perraudeau
- & Jean-Philippe Vert
-
Article
| Open Access Intelligent image-based in situ single-cell isolation
Intelligent image-based in situ single-cell isolation
The isolation of single cells while retaining context is important for quantifying cellular heterogeneity but technically challenging. Here, the authors develop a high-throughput, scalable workflow for microscopy-based single cell isolation using machine-learning, high-throughput microscopy and laser capture microdissection.
- Csilla Brasko
- , Kevin Smith
- & Peter Horvath
-
Article
| Open Access High-resolution TADs reveal DNA sequences underlying genome organization in flies
High-resolution TADs reveal DNA sequences underlying genome organization in flies
Although topologically associating domains (TADs) have been extensively investigated, it is not clear to what extent DNA sequence contributes to their formation. Here the authors develop software to identify high-resolution TAD boundaries and reveal their relationship to underlying DNA motifs.
- Fidel Ramírez
- , Vivek Bhardwaj
- & Thomas Manke
-
Article
| Open Access Monitoring single-cell gene regulation under dynamically controllable conditions with integrated microfluidics and software
Monitoring single-cell gene regulation under dynamically controllable conditions with integrated microfluidics and software
How gene regulatory pathways control cell fate decisions in single cells is not fully understood. Here the authors present an integrated dual-input microfluidic chip and a linked analysis software, enabling tracking of gene regulatory responses of single bacterial cells to changing conditions.
- Matthias Kaiser
- , Florian Jug
- & Erik van Nimwegen
-
Article
| Open Access Causal associations between risk factors and common diseases inferred from GWAS summary data
Causal associations between risk factors and common diseases inferred from GWAS summary data
Genetic methods are useful to test whether risk factors are causal for or consequence of disease. Here, Zhu et al. develop a generalized summary-based Mendelian Randomization (GSMR) method which uses summary-level data from GWAS to test for causal associations of health risk factors with common diseases.
- Zhihong Zhu
- , Zhili Zheng
- & Jian Yang
-
Article
| Open Access Pathway design using de novo steps through uncharted biochemical spaces
Pathway design using de novo steps through uncharted biochemical spaces
Existing pathway design tools make use of existing reactions from databases or successively apply retrosynthetic rules. novoStoic provides an integrated optimization-based framework combining known reactions with novel steps in pathway design allowing for constraints on thermodynamic feasibility, product yield, pathway length and number of novel steps.
- Akhil Kumar
- , Lin Wang
- & Costas D. Maranas
-
Article
| Open Access Experimental noise cutoff boosts inferability of transcriptional networks in large-scale gene-deletion studies
Experimental noise cutoff boosts inferability of transcriptional networks in large-scale gene-deletion studies
Reliable inference of gene interactions from perturbation experiments remains a challenge. Here, the authors quantify the upper limits of transcriptional network inference from knockout screens, identify the key determinants of accuracy, and introduce an unbiased and scalable inference algorithm.
- C. F. Blum
- , N. Heramvand
- & M. Kollmann
-
Article
| Open Access A machine learning approach to integrate big data for precision medicine in acute myeloid leukemia
A machine learning approach to integrate big data for precision medicine in acute myeloid leukemia
Identification of markers of drug response is essential for precision therapy. Here the authors introduce an algorithm that uses prior information about each gene’s importance in AML to identify the most predictive gene-drug associations from transcriptome and drug response data from 30 AML samples.
- Su-In Lee
- , Safiye Celik
- & Pamela S. Becker
-
Article
| Open Access Targeting immune checkpoints potentiates immunoediting and changes the dynamics of tumor evolution
Targeting immune checkpoints potentiates immunoediting and changes the dynamics of tumor evolution
The cancer immunoediting hypothesis assumes the immune system sculpts the cancer genome. Here the authors show, in a mouse model, that neutral evolution outweighs the effects of immunoselection and that immune checkpoint blockade potentiates the immunoediting, switching the system to non-neutral evolution.
- Mirjana Efremova
- , Dietmar Rieder
- & Zlatko Trajanoski
-
Article
| Open Access Perturbation-response genes reveal signaling footprints in cancer gene expression
Perturbation-response genes reveal signaling footprints in cancer gene expression
Deregulation of signalling is responsible for many cancer phenotypes. Leveraging available perturbation data, here the authors assess large-scale pathway activity patterns based on consensus downstream readout genes, enabling accurate prediction of the effects of mutations and small molecules.
- Michael Schubert
- , Bertram Klinger
- & Julio Saez-Rodriguez
-
Article
| Open Access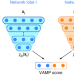 VAMPnets for deep learning of molecular kinetics
VAMPnets for deep learning of molecular kinetics
Extracting kinetic models from high-throughput molecular dynamics (MD) simulations is laborious and prone to human error. Here the authors introduce a deep learning framework that automates construction of Markov state models from MD simulation data.
- Andreas Mardt
- , Luca Pasquali
- & Frank Noé
-
Article
| Open Access Strain profiling and epidemiology of bacterial species from metagenomic sequencing
Strain profiling and epidemiology of bacterial species from metagenomic sequencing
Microbiota is often a complex mixture of multiple coexisting species and strains with high level of phenotypic and genomic variability. Here, Albanese and Donati develop StrainEst for estimating the number and identity of coexisting strains and their relative abundances in mixed metagenomic samples.
- Davide Albanese
- & Claudio Donati
-
Article
| Open Access Promoter-enhancer interactions identified from Hi-C data using probabilistic models and hierarchical topological domains
Promoter-enhancer interactions identified from Hi-C data using probabilistic models and hierarchical topological domains
Proximity-ligation methods like Hi-C map DNA-DNA interactions and reveal its organization into topologically associating domains (TADs). Here the authors describe PSYCHIC, a computational approach for analysing Hi-C data that allows the identification of promoter-enhancer interactions.
- Gil Ron
- , Yuval Globerson
- & Tommy Kaplan
-
Article
| Open Access Model-free inference of direct network interactions from nonlinear collective dynamics
Model-free inference of direct network interactions from nonlinear collective dynamics
Network dynamical systems can represent the interactions involved in the collective dynamics of gene regulatory networks or metabolic circuits. Here Casadiego et al. present a method for inferring these types of interactions directly from observed time series without relying on their model.
- Jose Casadiego
- , Mor Nitzan
- & Marc Timme
-
Article
| Open Access PAX7 target genes are globally repressed in facioscapulohumeral muscular dystrophy skeletal muscle
PAX7 target genes are globally repressed in facioscapulohumeral muscular dystrophy skeletal muscle
Facioscapulohumeral muscular dystrophy is a myopathy linked to ectopic expression of the DUX4 transcription factor. The authors show that the suppression of targets genes of the myogenesis regulator PAX7 is a signature of FSHD, and might explain oxidative stress sensitivity and epigenetic changes.
- Christopher R. S. Banerji
- , Maryna Panamarova
- & Peter S. Zammit
-
Article
| Open Access Evolutionary action and structural basis of the allosteric switch controlling β2AR functional selectivity
Evolutionary action and structural basis of the allosteric switch controlling β2AR functional selectivity
Ligand-induced biased signaling is thought to result in part from ligand-specific receptor conformations that cause the engagement of distinct effectors. Here the authors trace and evaluate the impact of mutations of the β2–adrenergic receptor on multiple signaling outputs to provide structural-level insight into the determinants of GPCR functional selectivity.
- Anne-Marie Schönegge
- , Jonathan Gallion
- & Michel Bouvier
-
Article
| Open Access Cell shape information is transduced through tension-independent mechanisms
Cell shape information is transduced through tension-independent mechanisms
It is not known whether the shape of a cell can regulate cellular phenotype independently. Here, the authors show that culturing kidney podocytes or smooth muscle cells on 3-D biomimetic surfaces results in phenotypic changes and that cell shape is sensed by integrin β3 in a tension-independent manner.
- Amit Ron
- , Evren U. Azeloglu
- & Ravi Iyengar
-
Article
| Open Access A general pharmacodynamic interaction model identifies perpetrators and victims in drug interactions
A general pharmacodynamic interaction model identifies perpetrators and victims in drug interactions
Assessment of pharmacodynamic interactions is at the heart of combination therapy development. Here the authors introduce a general drug interaction scoring model that enables quantification of synergistic and antagonistic interactions and determination of the directionality of the interactions.
- Sebastian G. Wicha
- , Chunli Chen
- & Ulrika S. H. Simonsson
-
Article
| Open Access Post-transcriptional 3´-UTR cleavage of mRNA transcripts generates thousands of stable uncapped autonomous RNA fragments
Post-transcriptional 3´-UTR cleavage of mRNA transcripts generates thousands of stable uncapped autonomous RNA fragments
Most mammalian genes contain alternative polyadenylation sites. Here, the authors provide evidence that mRNA can be cleaved post-transcriptionally to generate mRNAs with shorter 3-´UTRs and stable autonomous uncapped 3´-UTR sequences.
- Yuval Malka
- , Avital Steiman-Shimony
- & Michael Berger
-
Article
| Open Access Estimation of immune cell content in tumour tissue using single-cell RNA-seq data
Estimation of immune cell content in tumour tissue using single-cell RNA-seq data
Mathematical approaches can be used to assess immune cell composition from the tumour's bulk expression data. Here the authors optimise the CYBERSORT-based deconvolution algorithm by including cell type-specific reference gene expression profiles generated from tumour-derived single-cell RNA sequencing data.
- Max Schelker
- , Sonia Feau
- & Andreas Raue
-
Article
| Open Access Differentiation dynamics of mammary epithelial cells revealed by single-cell RNA sequencing
Differentiation dynamics of mammary epithelial cells revealed by single-cell RNA sequencing
There is a need to understand how mammary epithelial cells respond to changes at various developmental stages. Here, the authors use single-cell RNA sequencing of mammary epithelial cells at different adult developmental stages, identifying different cell types and charting their developmental trajectory.
- Karsten Bach
- , Sara Pensa
- & Walid T. Khaled
-
Article
| Open Access Integrative transcriptomic analysis reveals key drivers of acute peanut allergic reactions
Integrative transcriptomic analysis reveals key drivers of acute peanut allergic reactions
Rising rates of peanut allergy pose a public health problem. Here, the authors profile blood transcriptomes during double-blind, placebo-controlled oral challenge in peanut-allergic children to identify gene and cell composition changes, and construct causal networks to detect key allergic reaction drivers.
- C. T. Watson
- , A. T. Cohain
- & S. Bunyavanich
-
Article
| Open Access Meta-analysis of gut microbiome studies identifies disease-specific and shared responses
Meta-analysis of gut microbiome studies identifies disease-specific and shared responses
Reported associations between the human microbiome and disease are often inconsistent. Here, Duvallet et al. perform a meta-analysis of 28 gut microbiome studies spanning ten diseases, and find associations that are likely not disease-specific but potentially part of a shared response to disease.
- Claire Duvallet
- , Sean M. Gibbons
- & Eric J. Alm
-
Article
| Open Access Network dynamics-based cancer panel stratification for systemic prediction of anticancer drug response
Network dynamics-based cancer panel stratification for systemic prediction of anticancer drug response
Genomic alterations underlie the variability of drug responses between cancers, but our mechanistic understanding is limited. Here the authors use the p53 network to study how rewiring of signalling networks by genomic alterations impact their dynamic response to pharmacological perturbation.
- Minsoo Choi
- , Jue Shi
- & Kwang-Hyun Cho
-
Article
| Open Access Inference of differentiation time for single cell transcriptomes using cell population reference data
Inference of differentiation time for single cell transcriptomes using cell population reference data
Single cell transcriptome data can be used to determine developmental lineage trajectories. Here the authors map single cell transcriptomes onto a differentiation trajectory defined by cell population transcriptomes and show that cell cycle regulators have a role in differentiation timing.
- Na Sun
- , Xiaoming Yu
- & Jing-Dong J. Han
-
Article
| Open Access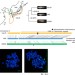 Centromere evolution and CpG methylation during vertebrate speciation
Centromere evolution and CpG methylation during vertebrate speciation
Centromeres and large-scale structural variants evolve and contribute to genome diversity during vertebrate speciation. Here Ichikawa et al perform de novo long-read genome assembly of three inbred medaka strains, and report long-range structure of centromeres and their methylation as well as correlation of structural variants with differential gene expression.
- Kazuki Ichikawa
- , Shingo Tomioka
- & Shinich Morishita
-
Article
| Open Access Functional mapping and annotation of genetic associations with FUMA
Functional mapping and annotation of genetic associations with FUMA
Prioritizing genetic variants is a major challenge in genome-wide association studies. Here, the authors develop FUMA, a web-based bioinformatics tool that uses a combination of positional, eQTL and chromatin interaction mapping to prioritize likely causal variants and genes.
- Kyoko Watanabe
- , Erdogan Taskesen
- & Danielle Posthuma
-
Article
| Open Access NSD1- and NSD2-damaging mutations define a subset of laryngeal tumors with favorable prognosis
NSD1- and NSD2-damaging mutations define a subset of laryngeal tumors with favorable prognosis
The authors use an integrative clustering approach to identify two laryngeal cancer clusters with distinct prognosis and show that mutations damaging the NSD1 and NSD2 methyltransferases segregate to the cluster with favorable prognosis, and independently predict longer survival in patients with laryngeal, but not other head and neck cancers.
- Suraj Peri
- , Evgeny Izumchenko
- & Erica A. Golemis
-
Article
| Open Access Single-cell absolute contact probability detection reveals chromosomes are organized by multiple low-frequency yet specific interactions
Single-cell absolute contact probability detection reveals chromosomes are organized by multiple low-frequency yet specific interactions
Eukaryotic genomes are partitioned into self-interacting modules or topologically associated domains (TADs) that exist at the kilo-megabase scale. Here Cattoni et al. combine super-resolution microscopy with DNA-labeling methods to quantify absolute frequencies of interactions within TADs.
- Diego I. Cattoni
- , Andrés M. Cardozo Gizzi
- & Marcelo Nollmann
-
Article
| Open Access Visual analysis of mass cytometry data by hierarchical stochastic neighbour embedding reveals rare cell types
Visual analysis of mass cytometry data by hierarchical stochastic neighbour embedding reveals rare cell types
Single cell profiling yields high dimensional data of very large numbers of cells, posing challenges of visualization and analysis. Here the authors introduce a method for analysis of mass cytometry data that can handle very large datasets and allows their intuitive and hierarchical exploration.
- Vincent van Unen
- , Thomas Höllt
- & Boudewijn P. F. Lelieveldt
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Clonal dynamics towards the development of venetoclax resistance in chronic lymphocytic leukemia
Clonal dynamics towards the development of venetoclax resistance in chronic lymphocytic leukemia