Featured
-
-
Article
| Open Access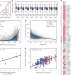 RNA structure drives interaction with proteins
RNA structure drives interaction with proteins
Interactions between protein and RNA occur on a large scale, and are often studied from a proteincentric point of view. Here the authors show that the number and strength of RNA-protein interactions correlate with the amount of RNA structure, which impacts the biological activity of the transcript.
- Natalia Sanchez de Groot
- , Alexandros Armaos
- & Gian Gaetano Tartaglia
-
Article
| Open Access Genetic mapping of cell type specificity for complex traits
Genetic mapping of cell type specificity for complex traits
Tissue- and cell type-specific information helps to interpret findings from genome-wide association studies. Here, the authors leverage multiple single cell expression datasets to infer cell type specificity of traits.
- Kyoko Watanabe
- , Maša Umićević Mirkov
- & Danielle Posthuma
-
Article
| Open Access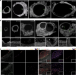 Single-cell reconstruction of follicular remodeling in the human adult ovary
Single-cell reconstruction of follicular remodeling in the human adult ovary
In the ovary, follicular degeneration occurs after folliculogenesis, with one dominant follicle reaching maturity every month. Here, by performing single cell RNA-seq of adult human follicles, the authors identify subclasses of granulosa and theca cell lineages, which correlate with the growth or degeneration of the follicles.
- X. Fan
- , M. Bialecka
- & S. M. Chuva de Sousa Lopes
-
Article
| Open Access Predictive metabolomic profiling of microbial communities using amplicon or metagenomic sequences
Predictive metabolomic profiling of microbial communities using amplicon or metagenomic sequences
Obtaining metabolomic data from microbial communities can be costly and difficult, whereas many microbial community sequence datasets are already available. Here Mallick et al. describe a computational approach to predict metabolic features from microbial DNA sequencing information.
- Himel Mallick
- , Eric A. Franzosa
- & Curtis Huttenhower
-
Article
| Open Access NASC-seq monitors RNA synthesis in single cells
NASC-seq monitors RNA synthesis in single cells
Sequencing of newly synthesised RNA can reveal the transcriptional dynamics in a population of cells. Here the authors develop NASC-seq to bring this sensitivity and temporal resolution to single-cell analysis.
- Gert-Jan Hendriks
- , Lisa A. Jung
- & Rickard Sandberg
-
Article
| Open Access Label-free neuroimaging in vivo using synchronous angular scanning microscopy with single-scattering accumulation algorithm
Label-free neuroimaging in vivo using synchronous angular scanning microscopy with single-scattering accumulation algorithm
A major challenge of in vivo imaging is imaging deeper, including in turbid tissue. The authors report an adaptive optics based microscope that uses coherent single scattering signal to reduce sample-induced aberrations and enable fast deep-tissue imaging of in vivo larval zebrafish brain.
- Moonseok Kim
- , Yonghyeon Jo
- & Wonshik Choi
-
Article
| Open Access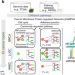 Membrane protein-regulated networks across human cancers
Membrane protein-regulated networks across human cancers
Membrane proteins have been implicated in cancers, but studying the downstream effects of their perturbation remains challenging. Here, the authors map the membrane protein-regulated network of 15 cancers, a resource for prognostic biomarker development and druggable target identification.
- Chun-Yu Lin
- , Chia-Hwa Lee
- & Jinn-Moon Yang
-
Article
| Open Access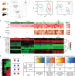 Defective HNF4alpha-dependent gene expression as a driver of hepatocellular failure in alcoholic hepatitis
Defective HNF4alpha-dependent gene expression as a driver of hepatocellular failure in alcoholic hepatitis
Alcoholic hepatitis, a common cause of liver failure, lacks effective treatment. Here, the authors show altered hepatic HNF4a isoform expression and hypermethylation of its target genes in patients. HNF4a dysregulation is improved in vitro by TGFb or PPARg modulation suggesting potential therapeutic avenues.
- Josepmaria Argemi
- , Maria U. Latasa
- & Ramon Bataller
-
Comment
| Open AccessUsing “outbreak science” to strengthen the use of models during epidemics
Infectious disease modeling has played a prominent role in recent outbreaks, yet integrating these analyses into public health decision-making has been challenging. We recommend establishing ‘outbreak science’ as an inter-disciplinary field to improve applied epidemic modeling.
- Caitlin Rivers
- , Jean-Paul Chretien
- & Simon Pollett
-
Article
| Open Access Environmental conditions shape the nature of a minimal bacterial genome
Environmental conditions shape the nature of a minimal bacterial genome
Minimal bacterial genomes still contain hundreds of genes of unknown function. Here the authors use in silico annotation methods and identify the environmental factors shaping a minimal genome.
- Magdalena Antczak
- , Martin Michaelis
- & Mark N. Wass
-
Article
| Open Access Landscape of transcriptomic interactions between breast cancer and its microenvironment
Landscape of transcriptomic interactions between breast cancer and its microenvironment
The transcriptomic profile of tumour-adjacent cells provides important information about tumour context but its clinical utility is unclear. Here, in breast cancer, Fox et al. show that the mRNA abundances of tumour and tumour-adjacent cells hold prognostic information.
- Natalie S. Fox
- , Syed Haider
- & Paul C. Boutros
-
Article
| Open Access Weakly supervised classification of aortic valve malformations using unlabeled cardiac MRI sequences
Weakly supervised classification of aortic valve malformations using unlabeled cardiac MRI sequences
The availability of labelled training data is one of the practical obstacles towards wide application of machine learning models in medicine. Here the authors develop a weakly supervised deep learning model for the classification of aortic malformations using unlabelled cardiac MRI sequences from the UK biobank.
- Jason A. Fries
- , Paroma Varma
- & James R. Priest
-
Article
| Open Access Fourier ring correlation simplifies image restoration in fluorescence microscopy
Fourier ring correlation simplifies image restoration in fluorescence microscopy
Fourier ring correlation (FRC) analysis is commonly used in fluorescence microscopy to measure effective image resolution. Here, the authors demonstrate that FRC can also be leveraged in blind image restoration methods, such as image deconvolution.
- Sami Koho
- , Giorgio Tortarolo
- & Giuseppe Vicidomini
-
Article
| Open Access Detection of cell-type-specific risk-CpG sites in epigenome-wide association studies
Detection of cell-type-specific risk-CpG sites in epigenome-wide association studies
Cellular heterogeneity is one of the major confounding factors in EWAS studies. Here the authors present a statistical method, HIgh REsolution (HIRE), which enables the detection of risk-CpG sites for individual cell types.
- Xiangyu Luo
- , Can Yang
- & Yingying Wei
-
Article
| Open Access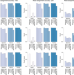 Strain-level metagenomic assignment and compositional estimation for long reads with MetaMaps
Strain-level metagenomic assignment and compositional estimation for long reads with MetaMaps
Sequencing platforms, such as Oxford Nanopore or Pacific Biosciences generate long-read data that preserve long-range genomic information but have high error rates. Here, the authors develop MetaMaps, a computational tool for strain-level metagenomic assignment and compositional estimation using long reads.
- Alexander T. Dilthey
- , Chirag Jain
- & Adam M. Phillippy
-
Article
| Open Access Integrating biomedical research and electronic health records to create knowledge-based biologically meaningful machine-readable embeddings
Integrating biomedical research and electronic health records to create knowledge-based biologically meaningful machine-readable embeddings
The Scalable Precision Medicine Oriented Knowledge Engine (SPOKE) is a heterogeneous knowledge network that integrates information from 29 public databases. Here, Nelson et al. extend SPOKE to embed clinical data from electronic health records to create medically meaningful barcodes for each medical variable.
- Charlotte A. Nelson
- , Atul J. Butte
- & Sergio E. Baranzini
-
Article
| Open Access A systematic approach to orient the human protein–protein interaction network
A systematic approach to orient the human protein–protein interaction network
The directions of most human protein-protein interactions (PPIs) remain unknown. Here, the authors use cancer genomic and drug response data to infer the direction of signal flow in the human PPI network and show that the directed network improves drug target and cancer driver gene prioritization.
- Dana Silverbush
- & Roded Sharan
-
Article
| Open Access Chemical genomics reveals histone deacetylases are required for core regulatory transcription
Chemical genomics reveals histone deacetylases are required for core regulatory transcription
Core regulatory transcription factors are usually regulated by cell-type specific super enhancers (SEs). Here, the authors screen for chemical probes able to distinguish between SE-driven and promoter-driven transcription and find that histone deacetylases are selectively required for core regulatory transcription.
- Berkley E. Gryder
- , Lei Wu
- & Javed Khan
-
Article
| Open Access Accurate estimation of cell-type composition from gene expression data
Accurate estimation of cell-type composition from gene expression data
Bulk RNA-seq data harbors valuable information about gene expression levels from different cell types in tissue samples. Here, the authors develop DWLS, a computational method for estimating cell-type composition of bulk data by leveraging single-cell RNA-seq-derived cell-type signatures.
- Daphne Tsoucas
- , Rui Dong
- & Guo-Cheng Yuan
-
Article
| Open Access PRC1 collaborates with SMCHD1 to fold the X-chromosome and spread Xist RNA between chromosome compartments
PRC1 collaborates with SMCHD1 to fold the X-chromosome and spread Xist RNA between chromosome compartments
The inactive X (Xi)-specific S1/S2 chromosome compartments are merged by SMCHD1, but how the S1/S2 structure is constructed is unclear. The authors find that PRC1 drives the formation of S1/S2s and that the stepwise folding process of the Xi facilitates Xist RNA spreading between Xi compartments.
- Chen-Yu Wang
- , David Colognori
- & Jeannie T. Lee
-
Article
| Open Access Nuclei multiplexing with barcoded antibodies for single-nucleus genomics
Nuclei multiplexing with barcoded antibodies for single-nucleus genomics
Single-nucleus RNA-seq enables interrogation of complex tissues but is limited due to batch effects and processing costs. Here the authors use barcoded antibodies against the nuclear pore complex to label nuclei from distinct samples, and develop a computational tool to assign the sample of origin.
- Jellert T. Gaublomme
- , Bo Li
- & Aviv Regev
-
Article
| Open Access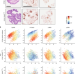 Cell type-dependent differential activation of ERK by oncogenic KRAS in colon cancer and intestinal epithelium
Cell type-dependent differential activation of ERK by oncogenic KRAS in colon cancer and intestinal epithelium
KRASG12V and BRAFV600E are oncogenic mutations that activate ERK signalling. Here, the authors use single cell analysis in intestinal organoids and show that BRAFV600E activates ERK in all intestinal cell types, while KRASG12V induces ERK activation in only a subset of cells, depending on cell differentiation state.
- Raphael Brandt
- , Thomas Sell
- & Markus Morkel
-
Article
| Open Access Improving the diagnostic yield of exome- sequencing by predicting gene–phenotype associations using large-scale gene expression analysis
Improving the diagnostic yield of exome- sequencing by predicting gene–phenotype associations using large-scale gene expression analysis
A genetic diagnosis remains unattainable for many individuals with a rare disease because of incomplete knowledge about the genetic basis of many diseases. Here, the authors present the web-based tool GADO (GeneNetwork Assisted Diagnostic Optimization) that uses public RNA-seq data for prioritization of candidate genes.
- Patrick Deelen
- , Sipko van Dam
- & Lude Franke
-
Article
| Open Access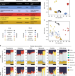 Transcriptional profiling unveils type I and II interferon networks in blood and tissues across diseases
Transcriptional profiling unveils type I and II interferon networks in blood and tissues across diseases
The authors present an extensive profile of host transcriptional respones to a diverse group of pathogens and allergens. In doing so, they identify TH1, type I IFN, TH17, and TH2 responses, that underlie each immune response in both the blood and lung, which represents a global profile of host-pathogen immune responses.
- Akul Singhania
- , Christine M. Graham
- & Anne O’Garra
-
Article
| Open Access Conserved transcriptomic profile between mouse and human colitis allows unsupervised patient stratification
Conserved transcriptomic profile between mouse and human colitis allows unsupervised patient stratification
Clinical and molecular heterogeneity of ulcerative colitis presents unresolved challenges to identify predictive biomarkers of response to therapies. Here, the authors combine mouse colitis time course with patient biopsy transcriptomes, achieving unsupervised clustering of UC patients correlating with therapeutic outcomes in independent data sets.
- Paulo Czarnewski
- , Sara M. Parigi
- & Eduardo J. Villablanca
-
Article
| Open Access MorphoNet: an interactive online morphological browser to explore complex multi-scale data
MorphoNet: an interactive online morphological browser to explore complex multi-scale data
Most morphological visualization platforms are not designed to share research data, or are limited to data visualization. Here the authors present MorphoNet, an open-source, web-based tool for interactive visualization and sharing of complex morphodynamic datasets, onto which users can project their own data.
- Bruno Leggio
- , Julien Laussu
- & Emmanuel Faure
-
Article
| Open Access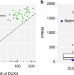 Long-read sequencing unveils IGH-DUX4 translocation into the silenced IGH allele in B-cell acute lymphoblastic leukemia
Long-read sequencing unveils IGH-DUX4 translocation into the silenced IGH allele in B-cell acute lymphoblastic leukemia
The IGH@ proto-oncogene translocation is a known genomic driver in several blood cancers. Here, the authors show that IGH-DUX4 translocation occurs on the silenced IGH allele avoiding toxic high-level expression of DUX4 in B-ALL.
- Liqing Tian
- , Ying Shao
- & Jinghui Zhang
-
Article
| Open Access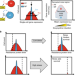 Role of network-mediated stochasticity in mammalian drug resistance
Role of network-mediated stochasticity in mammalian drug resistance
The role of gene expression noise in the evolution of drug resistance in mammalian cells is unclear. Here, by uncoupling noise from mean expression of a drug resistance gene in CHO cells the authors show that noisy expression aids adaptation to high drug levels, but delays it at low drug levels.
- Kevin S. Farquhar
- , Daniel A. Charlebois
- & Gábor Balázsi
-
Article
| Open Access Pathologic gene network rewiring implicates PPP1R3A as a central regulator in pressure overload heart failure
Pathologic gene network rewiring implicates PPP1R3A as a central regulator in pressure overload heart failure
The genetic and pathogenetic basis of heart failure is incompletely understood. Here, the authors present a high-fidelity tissue collection from rapidly preserved failing and non-failing control hearts which are used for eQTL mapping and network analysis, resulting in the prioritization of PPP1R3A as a heart failure gene.
- Pablo Cordero
- , Victoria N. Parikh
- & Euan A. Ashley
-
Article
| Open Access Minimal exposure of lipid II cycle intermediates triggers cell wall antibiotic resistance
Minimal exposure of lipid II cycle intermediates triggers cell wall antibiotic resistance
Antibiotics targeting cell wall synthesis display an unexplained gap between in vivo efficacy and in vitro binding affinity for their target. Here, Piepenbreier et al. develop a model for bacterial cell wall biosynthesis, show how it is affected by antibiotics, and use it to predict in vivo efficacy of antibiotics.
- Hannah Piepenbreier
- , Angelika Diehl
- & Georg Fritz
-
Article
| Open Access Learning cellular morphology with neural networks
Learning cellular morphology with neural networks
Volume electron microscopy data of brain tissue can tell us much about neural circuits, but increasingly large data sets demand automation of analysis. Here, the authors introduce cellular morphology neural networks and successfully automate a range of morphological analysis tasks.
- Philipp J. Schubert
- , Sven Dorkenwald
- & Joergen Kornfeld
-
Article
| Open Access Integrative inference of subclonal tumour evolution from single-cell and bulk sequencing data
Integrative inference of subclonal tumour evolution from single-cell and bulk sequencing data
Intra-tumour heterogeneity provides important information about subclonal tumour evolution. Here, the authors develop B-SCITE, a computational method for inferring tumour phylogenies from combined single-cell and bulk sequencing data.
- Salem Malikic
- , Katharina Jahn
- & Niko Beerenwinkel
-
Article
| Open Access Probabilistic controllability approach to metabolic fluxes in normal and cancer tissues
Probabilistic controllability approach to metabolic fluxes in normal and cancer tissues
Metabolic rewiring is a feature of many cancers. Here, the authors combine control theory and flux correlation analysis to study the transition of healthy metabolic networks to cancer states, and find that cancer metabolism is characterized by more streamlined flux distributions.
- Jean-Marc Schwartz
- , Hiroaki Otokuni
- & Jose C. Nacher
-
Article
| Open Access Establishing microbial composition measurement standards with reference frames
Establishing microbial composition measurement standards with reference frames
Most microbiome studies make conclusions based on changes in relative abundance of taxa, inferred from sequencing data. Here, the authors highlight common pitfalls in comparing relative abundance across samples, and identify solutions that reveal microbial changes without the need to estimate total microbial load.
- James T. Morton
- , Clarisse Marotz
- & Rob Knight
-
Article
| Open Access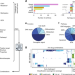 Community assessment to advance computational prediction of cancer drug combinations in a pharmacogenomic screen
Community assessment to advance computational prediction of cancer drug combinations in a pharmacogenomic screen
Resistance to first line treatment is a major hurdle in cancer treatment, that can be overcome with drug combinations. Here, the authors provide a large drug combination screen across cancer cell lines to benchmark crowdsourced methods and to computationally predict drug synergies.
- Michael P. Menden
- , Dennis Wang
- & Julio Saez-Rodriguez
-
Article
| Open Access Genomic signatures of heterokaryosis in the oomycete pathogen Bremia lactucae
Genomic signatures of heterokaryosis in the oomycete pathogen Bremia lactucae
The oomycete Bremia lactucae is a highly variable pathogen that causes lettuce downy mildew. Here, the authors generate a high-quality genome assembly for B. lactucae, detect a high prevalence of heterokaryosis, and investigate its pathogenic consequences.
- Kyle Fletcher
- , Juliana Gil
- & Richard Michelmore
-
Article
| Open Access Statistics of chromatin organization during cell differentiation revealed by heterogeneous cross-linked polymers
Statistics of chromatin organization during cell differentiation revealed by heterogeneous cross-linked polymers
Chromatin is folded into Topologically Associating domains (TADs), with the organization and folding hierarchy of the TADs being highly dynamic. Here the authors develop a parsimonious randomly cross-linked (RCL) polymer model that maps high frequency encounters present in Hi-C data within and between TADs and reconstruct TADs across cell differentiation, revealing local chromatin re-organization.
- O. Shukron
- , V. Piras
- & D. Holcman
-
Article
| Open Access Orchestrated ensemble activities constitute a hippocampal memory engram
Orchestrated ensemble activities constitute a hippocampal memory engram
The brain stores memories through a set of neurons known as engram cells. Here, the authors show that engram cells in the mouse hippocampus are organized into sub-ensembles representing distinct pieces of information, which are then orchestrated to constitute an entire memory.
- Khaled Ghandour
- , Noriaki Ohkawa
- & Kaoru Inokuchi
-
Article
| Open Access Simulating multiple faceted variability in single cell RNA sequencing
Simulating multiple faceted variability in single cell RNA sequencing
Simulated single cell RNA sequencing data is useful for method development and comparison. Here, the authors developed SymSim, a simulator that explicitly models the main factors of variation in single cell data.
- Xiuwei Zhang
- , Chenling Xu
- & Nir Yosef
-
Article
| Open Access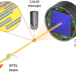 Coherent diffractive imaging of microtubules using an X-ray laser
Coherent diffractive imaging of microtubules using an X-ray laser
XFEL radiation is providing new opportunities for probing biological systems. Here the authors perform nanoscale x-ray imaging of microtubules with helical symmetry, by using imaging sorting and reconstruction techniques.
- Gisela Brändén
- , Greger Hammarin
- & Richard Neutze
-
Article
| Open Access Facial recognition from DNA using face-to-DNA classifiers
Facial recognition from DNA using face-to-DNA classifiers
Prediction of face from DNA followed by matching to facial images has been proposed for forensic applications. Here, Sero et al. present a different approach that can establish facial identity from DNA without directly predicting the face but is based on classifying given faces by individual DNA-encoded traits.
- Dzemila Sero
- , Arslan Zaidi
- & Peter Claes
-
Article
| Open Access Comparative analysis of mRNA and protein degradation in prostate tissues indicates high stability of proteins
Comparative analysis of mRNA and protein degradation in prostate tissues indicates high stability of proteins
Protein degradation in clinical samples is largely unexplored. Here, the authors analyze the transcriptome and proteome of clinical tissue samples and develop an algorithm to assess protein degradation, showing that protein degradation is negligible in most tissue samples and does not correlate with transcript degradation.
- Wenguang Shao
- , Tiannan Guo
- & Ruedi Aebersold
-
Article
| Open Access Detection of DNA base modifications by deep recurrent neural network on Oxford Nanopore sequencing data
Detection of DNA base modifications by deep recurrent neural network on Oxford Nanopore sequencing data
DNA modification generates unique electric signals in Oxford Nanopore sequencing data but the signals can be complicated to decipher. Here, the authors develop a deep learning framework, DeepMod, to detect DNA base modifications including 5mC and 6mA using Nanopore sequencing data
- Qian Liu
- , Li Fang
- & Kai Wang
-
Article
| Open Access Feed-forward regulation adaptively evolves via dynamics rather than topology when there is intrinsic noise
Feed-forward regulation adaptively evolves via dynamics rather than topology when there is intrinsic noise
Feed‐forward loops (FFLs) can filter out noise, but whether their overrepresentation in GRNs reflects adaptive evolution for this function is debated. Here, the authors develop a null model of regulatory evolution and find that FFLs evolve readily under selection for the noise filtering function.
- Kun Xiong
- , Alex K. Lancaster
- & Joanna Masel
-
Article
| Open Access A comprehensive single cell transcriptional landscape of human hematopoietic progenitors
A comprehensive single cell transcriptional landscape of human hematopoietic progenitors
Human Hematopoietic stem and progenitor cells (HSPCs) are commonly defined by CD34 expression. Here, the authors map single-cell RNA states both inside and outside the CD34 compartment, uncovering previously unappreciated branchpoints and validating CD164 as an efficient marker for early HSPCs.
- Danilo Pellin
- , Mariana Loperfido
- & Luca Biasco
-
Article
| Open Access Bivariate causal mixture model quantifies polygenic overlap between complex traits beyond genetic correlation
Bivariate causal mixture model quantifies polygenic overlap between complex traits beyond genetic correlation
To better understand the phenotypic relationships of complex traits it is also important to understand their genetic overlap. Here, Frei et al. develop MiXeR which uses GWAS summary statistics to evaluate the polygenic overlap between two traits irrespective of their genetic correlation.
- Oleksandr Frei
- , Dominic Holland
- & Anders M. Dale
-
Article
| Open Access A tessellation-based colocalization analysis approach for single-molecule localization microscopy
A tessellation-based colocalization analysis approach for single-molecule localization microscopy
Multicolour single-molecule localization microscopy lacks a standard analysis method. Here Levet et al. introduce Coloc-Tesseler, a parameter-free colocalisation analysis method based on tessellation analysis for the efficient analysis of multicolour SMLM data.
- Florian Levet
- , Guillaume Julien
- & Jean-Baptiste Sibarita
-
Article
| Open Access Flexible and scalable diagnostic filtering of genomic variants using G2P with Ensembl VEP
Flexible and scalable diagnostic filtering of genomic variants using G2P with Ensembl VEP
Diagnostic filtering is an important step to analyze the functional and clinical significance of the large number of genetic variants identified from next-generation genome sequencing data. Here, the authors develop a flexible and scalable system for diagnostic filtering of genetic variants using G2P with Ensembl VEP.
- Anja Thormann
- , Mihail Halachev
- & David R. FitzPatrick
-
Article
| Open Access Increasing species sampling in chelicerate genomic-scale datasets provides support for monophyly of Acari and Arachnida
Increasing species sampling in chelicerate genomic-scale datasets provides support for monophyly of Acari and Arachnida
Morphological and molecular data have led to conflicting phylogenetic hypotheses for the Chelicerata. Here, the authors reconstruct the phylogeny of the Chelicerata using genomic-scale datasets, finding evidence for a monophyletic Acari and a single terrestrialisation of Arachnida.
- Jesus Lozano-Fernandez
- , Alastair R. Tanner
- & Davide Pisani
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening
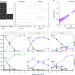
 Stochastic search and joint fine-mapping increases accuracy and identifies previously unreported associations in immune-mediated diseases
Stochastic search and joint fine-mapping increases accuracy and identifies previously unreported associations in immune-mediated diseases