Featured
-
-
Article
| Open Access An African Salmonella Typhimurium ST313 sublineage with extensive drug-resistance and signatures of host adaptation
An African Salmonella Typhimurium ST313 sublineage with extensive drug-resistance and signatures of host adaptation
Invasive non-typhoidal Salmonella (iNTS) infections are dominated by antibiotic resistant isolates of the sequence type (ST) 313. Here, the authors identify the ST313 sublineage II.1 in the Democratic Republic of the Congo exhibiting extensive drug resistance and genetic signatures potentially associated with host adaptation.
- Sandra Van Puyvelde
- , Derek Pickard
- & Stijn Deborggraeve
-
Article
| Open Access Optimized CRISPR guide RNA design for two high-fidelity Cas9 variants by deep learning
Optimized CRISPR guide RNA design for two high-fidelity Cas9 variants by deep learning
Application of highly specific Cas9 variants can be restricted by the design of the guide RNA. Here the authors present DeepHF, a gRNA activity prediction tool built from genome-scale screens of 50,000 guides covering 20,000 genes.
- Daqi Wang
- , Chengdong Zhang
- & Yongming Wang
-
Article
| Open Access Identification of significant chromatin contacts from HiChIP data by FitHiChIP
Identification of significant chromatin contacts from HiChIP data by FitHiChIP
HiChIP/PLAC-seq assay is popular for profiling 3D genome interactions among regulatory elements at kilobase resolution. Here the authors describe FitHiChIP an empirical null-based, flexible computational method for statistical significance estimation and loop calling from HiChIP data.
- Sourya Bhattacharyya
- , Vivek Chandra
- & Ferhat Ay
-
Article
| Open Access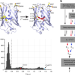 Learning the pattern of epistasis linking genotype and phenotype in a protein
Learning the pattern of epistasis linking genotype and phenotype in a protein
Epistasis underlies the complexity of genotype-phenotype maps. Here, the authors analyze 8,192 mutants that link two phenotypically distinct variants of the Entacmaea quadricolor fluorescent protein, and show the existence, but also the sparsity, of high-order epistatic interactions.
- Frank J. Poelwijk
- , Michael Socolich
- & Rama Ranganathan
-
Article
| Open Access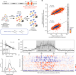 The mutational landscape of a prion-like domain
The mutational landscape of a prion-like domain
TDP43 aggregates are a hallmark of amyotrophic lateral sclerosis. By using deep mutagenesis to measure the toxicity of more than 50,000 mutations in the prion domain of TDP43, the authors conclude that mutations that increase toxicity promote formation of liquid-like condensates, while aggregation of TDP43 is protective for the cell.
- Benedetta Bolognesi
- , Andre J. Faure
- & Ben Lehner
-
Article
| Open Access UPF1/SMG7-dependent microRNA-mediated gene regulation
UPF1/SMG7-dependent microRNA-mediated gene regulation
UPF1 mediates the decay of target mRNA in a 3′ untranslated region (UTR)-length-dependent manner. Here the authors reveal that the 3′UTR-length-dependent regulation of UPF1-dependent mRNA decay occurs through EJC-independent but miRNA-dependent regulation.
- Jungyun Park
- , Jwa-Won Seo
- & Jin-Wu Nam
-
Article
| Open Access The distinction of CPR bacteria from other bacteria based on protein family content
The distinction of CPR bacteria from other bacteria based on protein family content
Recent studies have identified a large, phylogenetically distinct clade of bacteria, the candidate phyla radiation (CPR). Here, Méheust and colleagues analyze almost 3600 genomes to characterize the protein family content of CPR versus other bacteria and archaea.
- Raphaël Méheust
- , David Burstein
- & Jillian F. Banfield
-
Article
| Open Access RuvC uses dynamic probing of the Holliday junction to achieve sequence specificity and efficient resolution
RuvC uses dynamic probing of the Holliday junction to achieve sequence specificity and efficient resolution
Holliday junctions (HJs) are four-way DNA structures that occur in DNA repair by homologous recombination which are removed by nucleases such as RuvC. Here the authors provide structural and biochemical details on the steps required for the completion of the process.
- Karolina Maria Górecka
- , Miroslav Krepl
- & Marcin Nowotny
-
Article
| Open Access Exploring use of unsupervised clustering to associate signaling profiles of GPCR ligands to clinical response
Exploring use of unsupervised clustering to associate signaling profiles of GPCR ligands to clinical response
Identifying ligands which activate the specific effectors driving particular in vivo drug effects remains challenging. Here, the authors apply unsupervised clustering of pharmacodynamic parameters to classify GPCR ligands into different categories with similar signaling profiles and shared frequency of report of side effects.
- Besma Benredjem
- , Jonathan Gallion
- & Graciela Pineyro
-
Article
| Open Access Accurate detection of m6A RNA modifications in native RNA sequences
Accurate detection of m6A RNA modifications in native RNA sequences
We currently lack generic methods to map RNA modifications across the entire transcriptome. Here, the authors demonstrate that m6A RNA modifications can be detected with high accuracy using nanopore direct RNA sequencing.
- Huanle Liu
- , Oguzhan Begik
- & Eva Maria Novoa
-
Article
| Open Access Components of genetic associations across 2,138 phenotypes in the UK Biobank highlight adipocyte biology
Components of genetic associations across 2,138 phenotypes in the UK Biobank highlight adipocyte biology
While many pleiotropic genetic loci have been identified, how they contribute to phenotypes across traits and diseases is unclear. Here, the authors propose decomposition of genetic associations (DeGAs), which uses singular value decomposition, to characterize the underlying latent structure of genetic associations of 2,138 phenotypes.
- Yosuke Tanigawa
- , Jiehan Li
- & Manuel A. Rivas
-
Article
| Open Access Spatially clustered loci with multiple enhancers are frequent targets of HIV-1 integration
Spatially clustered loci with multiple enhancers are frequent targets of HIV-1 integration
HIV-1 usually targets active genes and integrates near the nuclear pore compartment. Here the authors show that recurrently targeted genes are proximal to super-enhancer genomic elements, which cluster in specific spatial compartments of the T cell nucleus, suggesting a role for nuclear organisation in viral infection.
- Bojana Lucic
- , Heng-Chang Chen
- & Marina Lusic
-
Article
| Open Access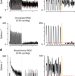 Neural sensitization improves encoding fidelity in the primate retina
Neural sensitization improves encoding fidelity in the primate retina
Light intensity on the retina can fluctuate rapidly during natural vision, posing a challenge for encoding visual information. Here, the authors report that mechanisms of sensitization/facilitation maintain the sensitivity of the numerically dominant neural pathway in the primate retina during dynamic vision.
- Todd R. Appleby
- & Michael B. Manookin
-
Article
| Open Access Translational coupling via termination-reinitiation in archaea and bacteria
Translational coupling via termination-reinitiation in archaea and bacteria
Archaea and bacteria often have gene pairs with overlapping stop and start codons, suggesting translational coupling. Here, Huber et al. analyse overlapping gene pairs from 720 genomes, and validate translational coupling via termination-reinitiation for 14 gene pairs in Haloferax volcanii and Escherichia coli.
- Madeleine Huber
- , Guilhem Faure
- & Jörg Soppa
-
Article
| Open Access Deep learning extends de novo protein modelling coverage of genomes using iteratively predicted structural constraints
Deep learning extends de novo protein modelling coverage of genomes using iteratively predicted structural constraints
Prediction of protein structures on the scale of genomes remains a challenge. Here the authors introduce a protein structure prediction method that uses deep learning to predict inter-atomic distances, torsion angles and hydrogen bonds, and apply it to predict the structures of 1475 Pfam domains.
- Joe G. Greener
- , Shaun M. Kandathil
- & David T. Jones
-
Comment
| Open Access Technology to advance infectious disease forecasting for outbreak management
Technology to advance infectious disease forecasting for outbreak management
Forecasting is beginning to be integrated into decision-making processes for infectious disease outbreak response. We discuss how technologies could accelerate the adoption of forecasting among public health practitioners, improve epidemic management, save lives, and reduce the economic impact of outbreaks.
- Dylan B. George
- , Wendy Taylor
- & Nicholas G. Reich
-
Article
| Open Access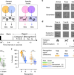 A dual role of prestimulus spontaneous neural activity in visual object recognition
A dual role of prestimulus spontaneous neural activity in visual object recognition
The effect of spontaneous variations in prestimulus neural activity on subsequent perception is incompletely understood. Here, using MEG, the authors identify two distinct neural processes that can influence object recognition in different ways.
- Ella Podvalny
- , Matthew W. Flounders
- & Biyu J. He
-
Article
| Open Access Single-cell transcriptomics reveals multi-step adaptations to endocrine therapy
Single-cell transcriptomics reveals multi-step adaptations to endocrine therapy
The development of resistance to endocrine therapy is a significant, clinical problem in breast cancer. Here, the authors identify a rare subpopulation of cells that drive resistance following transcriptional reprogramming.
- Sung Pil Hong
- , Thalia E. Chan
- & Luca Magnani
-
Article
| Open Access Changes in gene expression predictably shift and switch genetic interactions
Changes in gene expression predictably shift and switch genetic interactions
Non-additive genetic interactions are plastic and can complicate genetic prediction. Here, using deep mutagenesis of the lambda repressor, Li et al. reveal that changes in gene expression can alter the strength and direction of genetic interactions between mutations in many genes and develop mathematical models for predicting them.
- Xianghua Li
- , Jasna Lalić
- & Ben Lehner
-
Article
| Open Access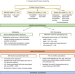 A phenotypic and genomics approach in a multi-ethnic cohort to subtype systemic lupus erythematosus
A phenotypic and genomics approach in a multi-ethnic cohort to subtype systemic lupus erythematosus
Systemic lupus erythematosus (SLE) is an autoimmune disease of substantial phenotypic heterogeneity in different ethnic groups. Here, using data from a multi-ethnic cohort, the authors describe an approach based on clinical and molecular data to subtype SLE patients into three clusters of severity.
- Cristina M. Lanata
- , Ishan Paranjpe
- & Lindsey A. Criswell
-
Article
| Open Access Identification of somatic mutations in single cell DNA-seq using a spatial model of allelic imbalance
Identification of somatic mutations in single cell DNA-seq using a spatial model of allelic imbalance
Single cell whole-genome sequencing data harbors information about somatic genetic variation but is challenging to analyze. Here, the authors develop a spatial model to correct for allelic amplification imbalance and a somatic SNV genotyper SCAN-SNV for analyzing single cell DNA sequencing data.
- Lovelace J. Luquette
- , Craig L. Bohrson
- & Peter J. Park
-
Article
| Open Access Integrative transcriptome imputation reveals tissue-specific and shared biological mechanisms mediating susceptibility to complex traits
Integrative transcriptome imputation reveals tissue-specific and shared biological mechanisms mediating susceptibility to complex traits
PrediXcan is a widely used gene expression imputation method that links genetic variants to gene expression. Here, the authors develop EpiXcan which leverages epigenetic annotations to inform transcriptomic imputation and further use the obtained gene-trait associations for computational drug repurposing.
- Wen Zhang
- , Georgios Voloudakis
- & Panos Roussos
-
Article
| Open Access Deep multiomics profiling of brain tumors identifies signaling networks downstream of cancer driver genes
Deep multiomics profiling of brain tumors identifies signaling networks downstream of cancer driver genes
Multi-omic profiling is a powerful approach to dissecting molecular mechanisms in disease. Here the authors generate whole proteome, phosphoproteome and transcriptome profiles from two mouse models of high-grade glioma driven by different oncogenes, and validate identified master regulators with a CRISPR screen.
- Hong Wang
- , Alexander K. Diaz
- & Junmin Peng
-
Article
| Open Access Cancer-associated mutations in DICER1 RNase IIIa and IIIb domains exert similar effects on miRNA biogenesis
Cancer-associated mutations in DICER1 RNase IIIa and IIIb domains exert similar effects on miRNA biogenesis
DICER is involved in the processing of miRNAs, where the RNase IIIa and IIIb domains are thought to cut the 3p and 5p hairpin arms, respectively. Here, in endometrial cancer, the authors identify an RNase IIIa mutation, which phenocopies mutations in the RNase IIIb domain.
- Jeffrey Vedanayagam
- , Walid K. Chatila
- & Eric C. Lai
-
Article
| Open Access A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing
A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing
The advent of single-cell RNA sequencing has revealed significant transcriptional heterogeneity in cancer, but its relationship to genomic heterogeneity remains unclear. Focusing on acute myeloid leukemia samples, the authors describe a general approach for linking mutation-containing cells to their transcriptional phenotypes using single-cell RNA sequencing data.
- Allegra A. Petti
- , Stephen R. Williams
- & Timothy J. Ley
-
Article
| Open Access Smu1 and RED are required for activation of spliceosomal B complexes assembled on short introns
Smu1 and RED are required for activation of spliceosomal B complexes assembled on short introns
Human spliceosome components Smu1 and RED regulate alternative splicing. Here the authors show that Smu1 and RED are also required for constitutive splicing of short introns.
- Sandra Keiper
- , Panagiotis Papasaikas
- & Reinhard Lührmann
-
Article
| Open Access The pause-initiation limit restricts transcription activation in human cells
The pause-initiation limit restricts transcription activation in human cells
Gene activation requires an increase of successful initiation events. Here, by employing a genome-wide kinetic analysis of transcription, the authors showed that gene activation generally requires a decrease in RNA Polymerase II (Pol II) promoter-proximal pausing while transcription of enhancer elements is not limited by Pol II pausing.
- Saskia Gressel
- , Björn Schwalb
- & Patrick Cramer
-
Article
| Open Access Past–future information bottleneck for sampling molecular reaction coordinate simultaneously with thermodynamics and kinetics
Past–future information bottleneck for sampling molecular reaction coordinate simultaneously with thermodynamics and kinetics
Efficient sampling of rare events in all-atom molecular dynamics simulations remains a challenge. Here, the authors adapt the Predictive Information Bottleneck framework to sample biomolecular structure and dynamics through iterative rounds of biased simulations and deep learning.
- Yihang Wang
- , João Marcelo Lamim Ribeiro
- & Pratyush Tiwary
-
Article
| Open Access Proteogenomic landscape of squamous cell lung cancer
Proteogenomic landscape of squamous cell lung cancer
Squamous cell lung cancer has dismal prognosis due to the dearth of effective treatments. Here, the authors perform an integrated proteogenomic analysis of the disease, revealing three proteomics-based subtypes and suggesting potential therapeutic opportunities.
- Paul A. Stewart
- , Eric A. Welsh
- & Eric B. Haura
-
Article
| Open Access Comprehensive transcriptomic analysis of cell lines as models of primary tumors across 22 tumor types
Comprehensive transcriptomic analysis of cell lines as models of primary tumors across 22 tumor types
Cell lines are used ubiquitously in cancer research but how well they represent the tumor type they were derived from is variable. Here, the authors compare transcriptomic profiles of 22 tumor types and cell lines and propose a new comprehensive cell line panel for pan-cancer studies.
- K. Yu
- , B. Chen
- & M. Sirota
-
Article
| Open Access Saturation mutagenesis of twenty disease-associated regulatory elements at single base-pair resolution
Saturation mutagenesis of twenty disease-associated regulatory elements at single base-pair resolution
Interpreting genetic variation in the noncoding genome remains challenging, with functional effects difficult to predict. Here, the authors perform saturation mutagenesis combined with massively parallel reporter assays for 20 disease-associated regulatory elements, quantifying the effects of over 30,000 variants.
- Martin Kircher
- , Chenling Xiong
- & Nadav Ahituv
-
Article
| Open Access A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism
A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism
Genome-scale metabolic models provide a platform to study metabolism through simulations and analysis of omics data. Here the authors introduce Yeast8 with its model ecosystem, a comprehensive computational resource for simulating the metabolism of Saccharomyces cerevisiae.
- Hongzhong Lu
- , Feiran Li
- & Jens Nielsen
-
Article
| Open Access Quantifying the impact of public omics data
Quantifying the impact of public omics data
Increasing amount of public omics data are important and valuable resources for the research community. Here, the authors develop a set of metrics to quantify the attention and impact of biomedical datasets and integrate them into the framework of Omics Discovery Index (OmicsDI).
- Yasset Perez-Riverol
- , Andrey Zorin
- & Henning Hermjakob
-
Article
| Open Access Pluripotency reprogramming by competent and incompetent POU factors uncovers temporal dependency for Oct4 and Sox2
Pluripotency reprogramming by competent and incompetent POU factors uncovers temporal dependency for Oct4 and Sox2
Oct4, along with Sox2 and Klf4 can induce pluripotency, but structurally similar factors like Oct6 cannot. Here, using pluripotency competent and incompetent factors, the authors show that Sox2 plays a dominant role in facilitating chromatin opening at Oct4 bound DNA early during reprogramming to pluripotency.
- Vikas Malik
- , Laura V. Glaser
- & Ralf Jauch
-
Article
| Open Access TeraVR empowers precise reconstruction of complete 3-D neuronal morphology in the whole brain
TeraVR empowers precise reconstruction of complete 3-D neuronal morphology in the whole brain
Reconstructing the full shape of neurons is a major informatics challenge as it requires handling huge whole-brain imaging datasets. Here the authors present an open-source virtual reality annotation system for precise and efficient data production of neuronal shapes reconstructed from whole brains.
- Yimin Wang
- , Qi Li
- & Hanchuan Peng
-
Article
| Open Access Cell-type-specific resolution epigenetics without the need for cell sorting or single-cell biology
Cell-type-specific resolution epigenetics without the need for cell sorting or single-cell biology
Compared to bulk data, cell-type-specific DNA methylation data provide higher resolution of epigenetic variation. Here, the authors introduce Tensor Composition Analysis, a novel computational approach for learning cell-type-specific DNA methylation from tissue-level bulk data, and show its application in epigenome-wide association studies.
- Elior Rahmani
- , Regev Schweiger
- & Eran Halperin
-
Article
| Open Access Fast and covariate-adaptive method amplifies detection power in large-scale multiple hypothesis testing
Fast and covariate-adaptive method amplifies detection power in large-scale multiple hypothesis testing
Side information in addition to the p-values is often available in modern applications of multiple hypothesis testing. Here, the authors develop
AdaFDR , a new statistical method for multiple hypothesis testing that adaptively learns the decision threshold and amplifies the discovery power.- Martin J. Zhang
- , Fei Xia
- & James Zou
-
Article
| Open Access MAPCap allows high-resolution detection and differential expression analysis of transcription start sites
MAPCap allows high-resolution detection and differential expression analysis of transcription start sites
The position, shape and number of transcription start sites (TSS) regulate gene expression. Here authors present MAPCap, a method for high-resolution detection and differential expression analysis of TSS, and apply MAPCap to early fly development, detecting stage and sex-specific promoter and enhancer activity.
- Vivek Bhardwaj
- , Giuseppe Semplicio
- & Asifa Akhtar
-
Article
| Open Access Profiling host ANP32A splicing landscapes to predict influenza A virus polymerase adaptation
Profiling host ANP32A splicing landscapes to predict influenza A virus polymerase adaptation
Polymorphisms in the avian influenza A virus (IAV) polymerase restrict its host range during transmission from birds to mammals. Here, the authors investigate differences in the host chromatin regulator ANP32A regarding IAV polymerase adaptation, and profile ANP32A splicing to predict avian species associated with pre-adaptive human-signatures in the virus.
- Patricia Domingues
- , Davide Eletto
- & Benjamin G. Hale
-
Article
| Open Access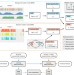 Detailed modeling of positive selection improves detection of cancer driver genes
Detailed modeling of positive selection improves detection of cancer driver genes
Finding driver genes sheds lights on the biological mechanisms propelling the development of a tumour, and can suggest therapeutic strategies. Here, the authors develop driverMAPS, a model-based approach to identify driver genes, and apply it to TCGA datasets.
- Siming Zhao
- , Jun Liu
- & Xin He
-
Article
| Open Access A machine-compiled database of genome-wide association studies
A machine-compiled database of genome-wide association studies
Most databases of genotype-phenotype associations are manually curated. Here, Kuleshov et al. describe a machine curation system that extracts such relationships from the GWAS literature and synthesizes them into a structured knowledge base called GWASkb that can complement manually curated databases.
- Volodymyr Kuleshov
- , Jialin Ding
- & Michael Snyder
-
Article
| Open Access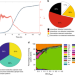 Regulatory mechanisms underlying coordination of amino acid and glucose catabolism in Escherichia coli
Regulatory mechanisms underlying coordination of amino acid and glucose catabolism in Escherichia coli
Bacteria must adapt their metabolism in the face of dynamically changing nutrient availability. Here, using their constraint-based modeling approach the authors analyze E. coli exometabolome data during growth in complex medium, revealing temporal coordination of glucose and amino acid catabolism.
- Mattia Zampieri
- , Manuel Hörl
- & Uwe Sauer
-
Article
| Open Access FDA-ARGOS is a database with public quality-controlled reference genomes for diagnostic use and regulatory science
FDA-ARGOS is a database with public quality-controlled reference genomes for diagnostic use and regulatory science
To be able to use infectious disease next generation sequencing as a diagnostic tool, appropriate reference datasets are required. Here, Sichtig et al. describe FDA-ARGOS, a reference database for high-quality microbial reference genomes, and demonstrate its utility on the example of two use cases.
- Heike Sichtig
- , Timothy Minogue
- & Uwe Scherf
-
Article
| Open Access Glycine, serine and threonine metabolism confounds efficacy of complement-mediated killing
Glycine, serine and threonine metabolism confounds efficacy of complement-mediated killing
Serum-resistant bacteria can escape complement killing in the bloodstream. Here, using metabolomics and metabolite perturbations, the authors describe an altered metabolic state in serum-resistant Escherichia coli and show that exogenous glycine potentiates elimination of pathogenic bacteria in vivo.
- Zhi-xue Cheng
- , Chang Guo
- & Xuan-xian Peng
-
Article
| Open Access Mendelian randomization integrating GWAS and eQTL data reveals genetic determinants of complex and clinical traits
Mendelian randomization integrating GWAS and eQTL data reveals genetic determinants of complex and clinical traits
Many genetic variants identified in genome-wide association studies are associated with gene expression. Here, Porcu et al. propose a transcriptome-wide summary statistics-based Mendelian randomization approach (TWMR) that, applied to 43 human traits, uncovers hundreds of previously unreported gene–trait associations.
- Eleonora Porcu
- , Sina Rüeger
- & Zoltán Kutalik
-
Review Article
| Open Access Towards a standardized bioinformatics infrastructure for N- and O-glycomics
Towards a standardized bioinformatics infrastructure for N- and O-glycomics
Glycomics is gaining momentum in basic, translational and clinical research. Here, the authors review current reporting standards and analysis tools for mass-spectrometry-based glycomics, and propose an e-infrastructure for standardized reporting and online deposition of glycomics data.
- Miguel A. Rojas-Macias
- , Julien Mariethoz
- & Niclas G. Karlsson
-
Article
| Open Access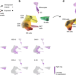 Predicting bacterial infection outcomes using single cell RNA-sequencing analysis of human immune cells
Predicting bacterial infection outcomes using single cell RNA-sequencing analysis of human immune cells
Complex interactions between different host immune cell types can determine the outcome of pathogen infections. Here, Avraham and colleagues present a deconvolution algorithm that uses single-cell RNA and bulk RNA sequencing measurements of pathogen-infected cells to predict disease risk outcomes.
- Noa Bossel Ben-Moshe
- , Shelly Hen-Avivi
- & Roi Avraham
-
Article
| Open Access Estimating dispensable content in the human interactome
Estimating dispensable content in the human interactome
The fraction of protein-protein interactions (PPIs) that can be disrupted without fitness effect is unknown. Here, the authors model how disease-causing mutations and common mutations carried by healthy people perturb the interactome, and estimate that <20% of human PPIs are completely dispensable.
- Mohamed Ghadie
- & Yu Xia
-
Article
| Open Access Comprehensive evaluation and characterisation of short read general-purpose structural variant calling software
Comprehensive evaluation and characterisation of short read general-purpose structural variant calling software
A number of computational methods have been developed for calling structural variants (SVs) using short read sequencing data. Here, the authors perform a comprehensive benchmarking analysis comparing 10 general-purpose callers and provide recommendations for both users and methods developers.
- Daniel L. Cameron
- , Leon Di Stefano
- & Anthony T. Papenfuss
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Enhancing and shaping the immunogenicity of native-like HIV-1 envelope trimers with a two-component protein nanoparticle
Enhancing and shaping the immunogenicity of native-like HIV-1 envelope trimers with a two-component protein nanoparticle