Featured
-
-
Article |
 GAGE-seq concurrently profiles multiscale 3D genome organization and gene expression in single cells
GAGE-seq concurrently profiles multiscale 3D genome organization and gene expression in single cells
GAGE-seq is a joint assay for 3D genome and transcriptome in single cells using combinatorial indexing to increase throughput. Applied to complex tissues, GAGE-seq enables the analysis of links between 3D organization and gene expression in rare cell types.
- Tianming Zhou
- , Ruochi Zhang
- & Jian Ma
-
Brief Communication
| Open Access Harnessing clonal gametes in hybrid crops to engineer polyploid genomes
Harnessing clonal gametes in hybrid crops to engineer polyploid genomes
An approach to generate unreduced, clonal gametes in hybrid tomato genotypes enables polyploid genome design through controlled combination of four predefined genome haplotypes, thereby establishing a framework for exploiting progressive heterosis in crops.
- Yazhong Wang
- , Roven Rommel Fuentes
- & Charles J. Underwood
-
Article |
 Near-gapless and haplotype-resolved apple genomes provide insights into the genetic basis of rootstock-induced dwarfing
Near-gapless and haplotype-resolved apple genomes provide insights into the genetic basis of rootstock-induced dwarfing
Near-gapless and haplotype-resolved genome assemblies of the dwarfing ‘M9’ and semi-vigorous ‘MM106’ rootstocks and a major apple cultivar ‘Fuji’ provide insights into the genetic basis of rootstock-induced dwarfing traits.
- Wei Li
- , Chong Chu
- & Zhenhai Han
-
Article
| Open Access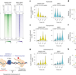 The BAF chromatin remodeler synergizes with RNA polymerase II and transcription factors to evict nucleosomes
The BAF chromatin remodeler synergizes with RNA polymerase II and transcription factors to evict nucleosomes
A chemical-genetic approach coupled with temporally resolved chromatin profiling shows that RNAPII promoter-proximal pausing stabilizes BAF complex occupancy and promotes nucleosome eviction.
- Sandipan Brahma
- & Steven Henikoff
-
News & Views |
 Current approaches to genomic deep learning struggle to fully capture human genetic variation
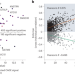
Current approaches to genomic deep learning struggle to fully capture human genetic variation
Deep learning shows promise for predicting gene expression levels from DNA sequences. However, recent studies show that current state-of-the-art models struggle to accurately characterize expression variation from personal genomes, limiting their usefulness in personalized medicine.
- Ziqi Tang
- , Shushan Toneyan
- & Peter K. Koo
-
Article
| Open Access Circular extrachromosomal DNA promotes tumor heterogeneity in high-risk medulloblastoma
Circular extrachromosomal DNA promotes tumor heterogeneity in high-risk medulloblastoma
Circular extrachromosomal DNA in high-risk medulloblastoma contributes to tumor heterogeneity and associates with relapse and survival. Enhancer rewiring events involving known oncogenes are frequent events, affecting transcription and proliferation.
- Owen S. Chapman
- , Jens Luebeck
- & Lukas Chavez
-
Article
| Open Access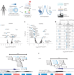 Transcriptional and epigenetic regulators of human CD8+ T cell function identified through orthogonal CRISPR screens
Transcriptional and epigenetic regulators of human CD8+ T cell function identified through orthogonal CRISPR screens
CRISPR activation/interference screens identify transcriptional regulators of human CD8+ T cells, including BATF3. BATF3 overexpression counteracts T cell exhaustion and enhances cancer immunotherapy in in vivo models.
- Sean R. McCutcheon
- , Adam M. Swartz
- & Charles A. Gersbach
-
Article
| Open Access Solanum americanum genome-assisted discovery of immune receptors that detect potato late blight pathogen effectors
Solanum americanum genome-assisted discovery of immune receptors that detect potato late blight pathogen effectors
High-quality genome assemblies of four Solanum Americanum accessions lead to the identification of three NLR-encoding genes, Rpi-amr4, R02860 and R04373, that recognize potato late blight pathogen Phytophthora infestans effectors.
- Xiao Lin
- , Yuxin Jia
- & Jonathan D. G. Jones
-
Article
| Open Access Single-molecule genome-wide mutation profiles of cell-free DNA for non-invasive detection of cancer
Single-molecule genome-wide mutation profiles of cell-free DNA for non-invasive detection of cancer
Somatic mutations are identified from circulating cell-free DNA using a single-molecule-based lowpass whole-genome sequencing method. The regional distribution of mutations can identify a tumor-specific mutational profile in patients with cancer and can be used to monitor patients through treatment.
- Daniel C. Bruhm
- , Dimitrios Mathios
- & Victor E. Velculescu
-
Research Briefing |
 Regulation of transposable elements and stem cell fate by crosstalk between RNA and DNA methylation
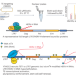
Regulation of transposable elements and stem cell fate by crosstalk between RNA and DNA methylation
How the chromatin states of transposable elements (TEs) are controlled in development and disease is unclear. We present CARGO-BioID, a CRISPR-based proteomic approach to identify TE-associated proteins, and reveal an interplay between RNA N6-methyladenosine (m6A) and DNA methylation that is crucial for regulating TE activation and human embryonic stem cell (hESC) fate.
-
Article |
 Potent and uniform fetal hemoglobin induction via base editing
Potent and uniform fetal hemoglobin induction via base editing
A comparison of fetal hemoglobin gene editing strategies using human sickle cell disease donor cells and in vivo transplantation finds that adenine base editing of the –175A>G site in the γ-globin gene promoters results in durable and potent expression.
- Thiyagaraj Mayuranathan
- , Gregory A. Newby
- & Jonathan S. Yen
-
Article |
 Single-cell multi-omics of mitochondrial DNA disorders reveals dynamics of purifying selection across human immune cells
Single-cell multi-omics of mitochondrial DNA disorders reveals dynamics of purifying selection across human immune cells
Single-cell analyses of immune cells from patients with pathogenic, single large-scale mitochondrial DNA (mtDNA) deletions including Pearson syndrome describe heteroplasmy dynamics consistent with purifying selection, as well as T-cell state-specific regulatory mechanisms and metabolic vulnerabilities.
- Caleb A. Lareau
- , Sonia M. Dubois
- & Leif S. Ludwig
-
-
Letter
| Open Access The wheat stem rust resistance gene Sr43 encodes an unusual protein kinase
The wheat stem rust resistance gene Sr43 encodes an unusual protein kinase
The resistance gene Sr43, which was crossed into bread wheat from the wild grass Thinopyrum elongatum, encodes an unusual protein kinase fusion protein that confers wheat stem rust resistance.
- Guotai Yu
- , Oadi Matny
- & Brande B. H. Wulff
-
Technical Report
| Open Access Single duplex DNA sequencing with CODEC detects mutations with high sensitivity
Single duplex DNA sequencing with CODEC detects mutations with high sensitivity
Concatenating Original Duplex for Error Correction (CODEC) is a method that concatenates both strands of each DNA duplex to enable highly sensitive mutation detection in a range of analytes with fewer reads and lower error rates than current methods.
- Jin H. Bae
- , Ruolin Liu
- & Viktor A. Adalsteinsson
-
Research Briefing |
 Tomato super-pangenome highlights the potential use of wild relatives in tomato breeding
Tomato super-pangenome highlights the potential use of wild relatives in tomato breeding
Genome assembly of nine wild species and two domesticated accessions of tomato generated a super-pangenome for the tomato clade. Comparative analyses revealed the landscape of structural variations in wild and cultivated tomatoes and led to the discovery of a wild tomato gene that has the potential for yield increase in modern breeding.
-
Article
| Open Access Super-pangenome analyses highlight genomic diversity and structural variation across wild and cultivated tomato species
Super-pangenome analyses highlight genomic diversity and structural variation across wild and cultivated tomato species
A tomato super-pangenome constructed using chromosome-scale genomes of nine wild species and two cultivated accessions highlights genomic diversity and structural variation across wild and cultivated tomatoes.
- Ning Li
- , Qiang He
- & Qinghui Yu
-
Research Briefing |
 sortChIC enables detailed chromatin analysis of rare cell types
sortChIC enables detailed chromatin analysis of rare cell types
Current methods of chromatin analysis focus mainly on the most abundant cell types in a sample. We present a workflow that combines enrichment of rare cell types with high-resolution mapping of histone modifications, which enables us to study chromatin dynamics in rare stem and progenitor cell populations.
-
Research Briefing |
 A scalable technology for measuring cell-type-specific activity of cis-regulatory sequences
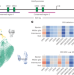
A scalable technology for measuring cell-type-specific activity of cis-regulatory sequences
A high-throughput reporter gene assay for use in living tissues has been developed. The assay allows investigators to quantify, in parallel, the activities of reporter genes in each of the cell types that constitute a tissue.
-
Technical Report |
 A single-cell massively parallel reporter assay detects cell-type-specific gene regulation
A single-cell massively parallel reporter assay detects cell-type-specific gene regulation
A single-cell massively parallel reporter assay is used to compare cis-regulatory sequence activities in cell line models and mouse retinal tissue ex vivo, identifying cell state- and cell-type-specific effects of sequence variation.
- Siqi Zhao
- , Clarice K. Y. Hong
- & Barak A. Cohen
-
Letter |
 Multimodal single-cell and whole-genome sequencing of small, frozen clinical specimens
Multimodal single-cell and whole-genome sequencing of small, frozen clinical specimens
High-quality multimodal single-cell, bulk and spatial genomics data are prepared from low-input, frozen needle biopsy specimens collected during routine clinical procedures.
- Yiping Wang
- , Joy Linyue Fan
- & Benjamin Izar
-
News & Views |
 Accounting for diversity in the design of CRISPR-based therapeutic genome editing
Accounting for diversity in the design of CRISPR-based therapeutic genome editing
CRISPR cell and gene therapy have been designed largely with respect to a single reference human genome. A new study reveals how human genetic diversity could lead to off-target effects and presents a new tool to identify these risks.
- Krishanu Saha
-
Research Briefing |
 Adult human kidney organoids: cellular origin and disease modeling
Adult human kidney organoids: cellular origin and disease modeling
Adult human kidney organoids or tubuloids are derived from an epithelial CD24+ subpopulation in the proximal nephron and can be utilized for advanced disease modeling of the most common hereditary kidney disease: autosomal dominant polycystic kidney disease.
-
Article |
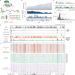 Long-range phasing of dynamic, tissue-specific and allele-specific regulatory elements
Long-range phasing of dynamic, tissue-specific and allele-specific regulatory elements
Nanopore sequencing is used to profile chromatin accessibility and DNA methylation on DNA molecules over 100 kb. Phasing analysis at the H19/IGF2 locus identifies a primate-specific enhancer driving biallelic IGF2 expression in specific cellular contexts.
- Sofia Battaglia
- , Kevin Dong
- & Bradley E. Bernstein
-
Technical Report |
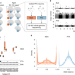 Single-cell genome sequencing of human neurons identifies somatic point mutation and indel enrichment in regulatory elements
Single-cell genome sequencing of human neurons identifies somatic point mutation and indel enrichment in regulatory elements
Single-cell DNA sequencing data are generated from human neurons using primary template-directed amplification and analyzed using SCAN2, an improved genotyping tool. Indels are enriched in neuronal regulatory elements and may be deleterious.
- Lovelace J. Luquette
- , Michael B. Miller
- & Peter J. Park
-
Article |
 Single-cell multi-omics of human clonal hematopoiesis reveals that DNMT3A R882 mutations perturb early progenitor states through selective hypomethylation
Single-cell multi-omics of human clonal hematopoiesis reveals that DNMT3A R882 mutations perturb early progenitor states through selective hypomethylation
Multi-modality single-cell sequencing determines genotype, transcriptome and methylome information in cells from individuals with DNMT3A R882 mutated clonal hematopoiesis, allowing for the comparison of mutant and wild-type cells from the same individuals.
- Anna S. Nam
- , Neville Dusaj
- & Dan A. Landau
-
Article |
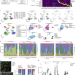 Single-nucleus and spatial transcriptome profiling of pancreatic cancer identifies multicellular dynamics associated with neoadjuvant treatment
Single-nucleus and spatial transcriptome profiling of pancreatic cancer identifies multicellular dynamics associated with neoadjuvant treatment
Single-nucleus and spatial, whole-transcriptome profiling of 43 pancreatic adenocarcinomas provides a refined molecular and cellular classification, highlighting a new neoadjuvant treatment-associated neural-like progenitor tumor cell state.
- William L. Hwang
- , Karthik A. Jagadeesh
- & Aviv Regev
-
Article
| Open Access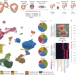 Origin and function of activated fibroblast states during zebrafish heart regeneration
Origin and function of activated fibroblast states during zebrafish heart regeneration
Single-cell RNA sequencing and spatiotemporal analysis of the regenerating zebrafish heart identify transient proregenerative fibroblast-like cells that are derived from the epicardium and the endocardium. Wnt signalling regulates the endocardial fibroblast response.
- Bo Hu
- , Sara Lelek
- & Jan Philipp Junker
-
Comment |
 Democratizing plant genomics to accelerate global food production
Democratizing plant genomics to accelerate global food production
Building on the fundamental discoveries of Mendel, plant genomics has had a major role in advancing the genetic improvement of crops worldwide, particularly in developed economies where the technologies are easily accessible. From cumbersome to more miniaturized high-throughput sequencing technologies, the field continues to evolve, providing vast opportunities for studying plant genomes with varying levels of complexity and potential real-life applications.
- Carol Nkechi Ibe
-
Research Briefing |
 Identification of two intrinsic epithelial subtypes of colorectal cancer
Identification of two intrinsic epithelial subtypes of colorectal cancer
By integrating single-cell and bulk transcriptomic analyses, we found that malignant cells belong to two major intrinsic epithelial subtypes. We propose a refined, three-tiered classification of colorectal cancer subtypes based on intrinsic epithelial subtypes, microsatellite instability status and the presence of fibrosis.
-
Article
| Open Access Single-cell and bulk transcriptome sequencing identifies two epithelial tumor cell states and refines the consensus molecular classification of colorectal cancer
Single-cell and bulk transcriptome sequencing identifies two epithelial tumor cell states and refines the consensus molecular classification of colorectal cancer
A single-cell transcriptomic analysis of 63 patients with colorectal cancer classifies tumor cells into two epithelial subtypes. An improved tumor classification based on epithelial subtype, microsatellite stability and fibrosis reveals differences in pathway activation and metastasis.
- Ignasius Joanito
- , Pratyaksha Wirapati
- & Iain Beehuat Tan
-
News & Views |
 Graph pangenomes find missing heritability
Graph pangenomes find missing heritability
The use of association studies to identify candidate genes for complex biological traits in plants has been challenging due to a reliance on single reference genomes, leading to missing heritability. Graphical pangenomes and the identification of causal variants help overcome this and provide an important advance for crop breeding.
- David Edwards
- & Jacqueline Batley
-
Comment |
 Genome-edited crops for improved food security of smallholder farmers
Genome-edited crops for improved food security of smallholder farmers
Widespread enthusiasm about potential contributions of genome-edited crops to address climate change, food security, nutrition and health, environmental sustainability and diversification of agriculture is dampened by concerns about the associated risks. Analysis of the top seven risks of genome-edited crops finds that the scientific risks are comparable to those of accepted, past and current breeding methods, but failure to address regulatory, legal and trade framework, and the granting of social license, squanders the potential benefits.
- Kevin V. Pixley
- , Jose B. Falck-Zepeda
- & Neal Gutterson
-
Article
| Open Access Chromosome-scale and haplotype-resolved genome assembly of a tetraploid potato cultivar
Chromosome-scale and haplotype-resolved genome assembly of a tetraploid potato cultivar
Haplotype-resolved genome assembly of the tetraploid potato cultivar ‘Otava’ sheds light on functional organization of the tetraploid genome and provides the potential for genomics-assisted breeding.
- Hequan Sun
- , Wen-Biao Jiao
- & Korbinian Schneeberger
-
Article |
 Paralog knockout profiling identifies DUSP4 and DUSP6 as a digenic dependence in MAPK pathway-driven cancers
Paralog knockout profiling identifies DUSP4 and DUSP6 as a digenic dependence in MAPK pathway-driven cancers
A CRISPR paralog targeting library profiling 815 paralog families across 11 cell lines identifies DUSP4 and DUSP6 as paralog pairs whose combined inactivation confers sensitivity to cells resistant to MAPK inhibitors or cells harboring NRAS or BRAF mutations.
- Takahiro Ito
- , Michael J. Young
- & William R. Sellers
-
News & Views |
 Reversing polycystic kidney disease
Reversing polycystic kidney disease
A new study shows that re-expressing PKD genes early in the course of the disease can fully reverse polycystic kidney disease in mice. These results reveal an unexpected ability of the kidney to regenerate following genetic rescue of polycystin function.
- Alessandra Boletta
-
Article
| Open Access High-quality genome assembly and resequencing of modern cotton cultivars provide resources for crop improvement
High-quality genome assembly and resequencing of modern cotton cultivars provide resources for crop improvement
High-quality genomes of two cultivated tetraploid cottons Gossypium hirsutum cv. NDM8 and Gossypium barbadense acc. Pima90 and resequencing of 1,081 G. hirsutum accessions provide insights into the role of structural variations.
- Zhiying Ma
- , Yan Zhang
- & Xingfen Wang
-
Comment |
 A call for direct sequencing of full-length RNAs to identify all modifications
A call for direct sequencing of full-length RNAs to identify all modifications
For most organisms, DNA sequences are available, but the complete RNA sequences are not. Here, we call for technologies to sequence full-length RNAs with all their modifications.
- Juan D. Alfonzo
- , Jessica A. Brown
- & Robert L. Ross
-
Letter |
 Long-read sequencing of 3,622 Icelanders provides insight into the role of structural variants in human diseases and other traits
Long-read sequencing of 3,622 Icelanders provides insight into the role of structural variants in human diseases and other traits
Analysis of long-read sequencing data from 3,622 Icelanders identifies a set of high-confidence structural variants and provides insights into their effect on human traits and diseases.
- Doruk Beyter
- , Helga Ingimundardottir
- & Kari Stefansson
-
Article |
 Population-scale single-cell RNA-seq profiling across dopaminergic neuron differentiation
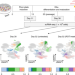
Population-scale single-cell RNA-seq profiling across dopaminergic neuron differentiation
Single-cell RNA-seq analysis of iPSC neural differentiation identifies markers that predict line-to-line differences in cell fate potential and eQTLs that are specific to different stages of differentiation and that overlap with GWAS risk variants for neurological traits.
- Julie Jerber
- , Daniel D. Seaton
- & Oliver Stegle
-
Comment |
 The UCSC SARS-CoV-2 Genome Browser
The UCSC SARS-CoV-2 Genome Browser
The UCSC SARS-CoV-2 Genome Browser (https://genome.ucsc.edu/covid19.html) is an adaptation of our popular genome-browser visualization tool for this virus, containing many annotation tracks and new features, including conservation with similar viruses, immune epitopes, RT–PCR and sequencing primers and CRISPR guides. We invite all investigators to contribute to this resource to accelerate research and development activities globally.
- Jason D. Fernandes
- , Angie S. Hinrichs
- & Maximilian Haeussler
-
Article |
 Subclonal reconstruction of tumors by using machine learning and population genetics
Subclonal reconstruction of tumors by using machine learning and population genetics
MOBSTER is an approach for subclonal reconstruction of tumors from cancer genomics data on the basis of models that combine machine learning with evolutionary theory, thus leading to more accurate evolutionary histories of tumors.
- Giulio Caravagna
- , Timon Heide
- & Andrea Sottoriva
-
Article |
 Identification of cancer driver genes based on nucleotide context
Identification of cancer driver genes based on nucleotide context
MutPanning is a new method to detect cancer driver genes that identifies genes with an excess of mutations in unusual nucleotide contexts. Applying this to whole-exome sequencing data from 11,873 tumor–normal pairs identifies 460 driver genes.
- Felix Dietlein
- , Donate Weghorn
- & Shamil R. Sunyaev
-
Article |
 A pooled single-cell genetic screen identifies regulatory checkpoints in the continuum of the epithelial-to-mesenchymal transition
A pooled single-cell genetic screen identifies regulatory checkpoints in the continuum of the epithelial-to-mesenchymal transition
Transcriptional profiling of 61,052 cells undergoing epithelial-to-mesenchymal transition (EMT) shows continuous waves of gene regulation, not discrete stages. A CRISPR screen orders EMT regulators along EMT progression.
- José L. McFaline-Figueroa
- , Andrew J. Hill
- & Cole Trapnell
-
Article |
 NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements
NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements
A new technique called NET-CAGE identifies 5′ ends of nascent RNAs with high sensitivity, including enhancer-derived RNAs. This approach shows that enhancer–promoter pairs are generally activated simultaneously.
- Shigeki Hirabayashi
- , Shruti Bhagat
- & Yasuhiro Murakawa
-
Article |
 High-throughput identification of human SNPs affecting regulatory element activity
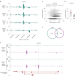
High-throughput identification of human SNPs affecting regulatory element activity
Application of SuRE reporter technology to survey the effect of 5.9 million SNPs in the human genome on enhancer and promoter activity identifies over 30,000 SNPs that alter the activity of putative regulatory elements.
- Joris van Arensbergen
- , Ludo Pagie
- & Bas van Steensel
-
Article |
 Rational targeting of a NuRD subcomplex guided by comprehensive in situ mutagenesis
Rational targeting of a NuRD subcomplex guided by comprehensive in situ mutagenesis
Comprehensive CRISPR mutagenesis targeting all members of the NuRD complex identifies a specific subcomplex required for fetal globin silencing and informs a rational targeting strategy for elevating globin levels while avoiding cytotoxicity.
- Falak Sher
- , Mir Hossain
- & Daniel E. Bauer
-
Article
| Open Access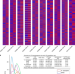 The genome of cultivated peanut provides insight into legume karyotypes, polyploid evolution and crop domestication
The genome of cultivated peanut provides insight into legume karyotypes, polyploid evolution and crop domestication
High-quality genome sequence of cultivated peanut comprising 2.54 Gb with 20 pseudomolecules and 83,709 protein-coding gene models provides insights into genome evolution and the genetic mechanisms underlying seed size and leaf resistance in peanut.
- Weijian Zhuang
- , Hua Chen
- & Rajeev K. Varshney
-
Article |
 Resequencing of 429 chickpea accessions from 45 countries provides insights into genome diversity, domestication and agronomic traits
Resequencing of 429 chickpea accessions from 45 countries provides insights into genome diversity, domestication and agronomic traits
The authors performed whole-genome resequencing of 429 chickpea lines sampled from 45 countries. They identified 122 candidate regions (204 genes) under selection during chickpea breeding.
- Rajeev K. Varshney
- , Mahendar Thudi
- & Xin Liu
Browse broader subjects
Browse narrower subjects
- Animal biotechnology
- Applied immunology
- Assay systems
- Biologics
- Biomaterials
- Biomimetics
- Cell delivery
- Environmental biotechnology
- Expression systems
- Functional genomics
- Gene delivery
- Gene therapy
- Genomics
- Industrial microbiology
- Metabolic engineering
- Metabolomics
- Molecular engineering
- Nanobiotechnology
- Nucleic-acid therapeutics
- Oligo delivery
- Peptide delivery
- Plant biotechnology
- Protein delivery
- Proteomics
- Regenerative medicine
- Sequencing
- Stem-cell biotechnology
- Tissue engineering

 Cicer super-pangenome provides insights into species evolution and agronomic trait loci for crop improvement in chickpea
Cicer super-pangenome provides insights into species evolution and agronomic trait loci for crop improvement in chickpea