Abstract
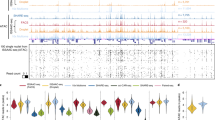
The organization of mammalian genomes features a complex, multiscale three-dimensional (3D) architecture, whose functional significance remains elusive because of limited single-cell technologies that can concurrently profile genome organization and transcriptional activities. Here, we introduce genome architecture and gene expression by sequencing (GAGE-seq), a scalable, robust single-cell co-assay measuring 3D genome structure and transcriptome simultaneously within the same cell. Applied to mouse brain cortex and human bone marrow CD34+ cells, GAGE-seq characterized the intricate relationships between 3D genome and gene expression, showing that multiscale 3D genome features inform cell-type-specific gene expression and link regulatory elements to target genes. Integration with spatial transcriptomic data revealed in situ 3D genome variations in mouse cortex. Observations in human hematopoiesis unveiled discordant changes between 3D genome organization and gene expression, underscoring a complex, temporal interplay at the single-cell level. GAGE-seq provides a powerful, cost-effective approach for exploring genome structure and gene expression relationships at the single-cell level across diverse biological contexts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All sequencing data from this study have been submitted to GEO under the accession number GSE238001. We use the following publicly available datasets in this work: in situ Hi-C datasets from Rao et al.3 (GSE: GSE63525); scHi-C datasets from Nagano et al.17 (GEO: GSE48262), Nagano et al.23 (GEO: GSE94489), Ramani et al.22 (GEO: GSE84920), Kim et al.37 (4DN Data Portal: 4DNES4D5MWEZ, 4DNESUE2NSGS, 4DNESIKGI39T, 4DNES1BK1RMQ and 4DNESTVIP977), Tan et al.26 (GEO: GSE117876), Tan et al.57 (GEO: GSE121791), Tan et al.27 (GEO: GSE162511), Flyamer et al.24 (GEO: GSE80006), Gassler et al.60 (GEO: GSE100569), Stevens et al.25 (GEO: GSE80280), Collombet et al.59 (GEO: GSE129029), Lee et al.44 (GEO: GSE124391), Liu et al.45 (GEO: GSE132489) and Mulqueen et al.58 (GEO: GSE174226); scRNA-seq datasets from Chen et al.62 (GEO: GSE126074), Plongthongkum et al.56 (GEO: GSE157660), Chen et al.55 (GEO: GSE178707), Ma et al.43 (GEO: GSE140203), Xu et al.65 (ArrayExpress: E-MTAB-11264), Xiong et al.66 (GEO: GSE158435), Zhu et al.63 (GEO: GSE130399), Zhu et al.52 (GEO: GSE152020), Cao et al.61 (GEO: GSE117089), Mimitou et al.64 (GEO: GSE126310) and Zhang et al.53 (GEO: GSE137864); HiRES co-assayed scHi-C and scRNA-seq datasets from Liu et al.35 (GEO: GSE223917); MERFISH spatial transcriptome datasets from Zhang et al.49 (Brain Image Library: cf1c1a431ef8d021); Paired-seq co-assayed scRNA-seq and scATAC-seq from Zhu et al.52 (GEO: GSE152020).
Code availability
The source code of the GAGE-seq data processing and analysis workflows can be accessed at: https://github.com/ma-compbio/GAGE-seq, which has also been deposited via Zenedo (https://doi.org/10.5281/zenodo.10888453)72. In our GitHub repository, we have provided notebooks (https://github.com/ma-compbio/GAGE-seq/tree/main/scripts_analysis) that detail the integration between GAGE-seq and Paired-seq data for single-cell joint analysis of 3D genome structure, chromatin accessibility and gene expression.
References
Dekker, J. et al. The 4D nucleome project. Nature 549, 219–226 (2017).
Cremer, T. & Cremer, C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat. Rev. Genet. 2, 292–301 (2001).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Xiong, K. & Ma, J. Revealing Hi-C subcompartments by imputing inter-chromosomal chromatin interactions. Nat. Commun. 10, 5069 (2019).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Beagan, J. A. & Phillips-Cremins, J. E. On the existence and functionality of topologically associating domains. Nat. Genet. 52, 8–16 (2020).
Salameh, T. J. et al. A supervised learning framework for chromatin loop detection in genome-wide contact maps. Nat. Commun. 11, 3428 (2020).
Tang, Z. et al. CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163, 1611–1627 (2015).
Marchal, C., Sima, J. & Gilbert, D. M. Control of DNA replication timing in the 3D genome. Nat. Rev. Mol. Cell Biol. 20, 721–737 (2019).
Ma, J. & Duan, Z. Replication timing becomes intertwined with 3D genome organization. Cell 176, 681–684 (2019).
Zheng, H. & Xie, W. The role of 3D genome organization in development and cell differentiation. Nat. Rev. Mol. Cell Biol. 20, 535–550 (2019).
Misteli, T. The self-organizing genome: principles of genome architecture and function. Cell 183, 28–45 (2020).
Spielmann, M., Lupiáñez, D. G. & Mundlos, S. Structural variation in the 3D genome. Nat. Rev. Genet. 19, 453–467 (2018).
Oudelaar, A. M. & Higgs, D. R. The relationship between genome structure and function. Nat. Rev. Genet. 22, 154–168 (2021).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Zhou, T., Zhang, R. & Ma, J. The 3D genome structure of single cells. Annu. Rev. Biomed. Data Sci. 4, 21–41 (2021).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019).
Cao, J. et al. A human cell atlas of fetal gene expression. Science 370, eaba7721 (2020).
Calderon, D. et al. The continuum of Drosophila embryonic development at single-cell resolution. Science 377, eabn5800 (2022).
Ramani, V. et al. Massively multiplex single-cell Hi-C. Nat. Methods 14, 263–266 (2017).
Nagano, T. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017).
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017).
Stevens, T. J. et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544, 59–64 (2017).
Tan, L., Xing, D., Chang, C.-H., Li, H. & Xie, X. S. Three-dimensional genome structures of single diploid human cells. Science 361, 924–928 (2018).
Tan, L. et al. Changes in genome architecture and transcriptional dynamics progress independently of sensory experience during post-natal brain development. Cell 184, 741–758.e17 (2021).
Li, G. et al. Joint profiling of DNA methylation and chromatin architecture in single cells. Nat. Methods 16, 991–993 (2019).
Zhang, R., Zhou, T. & Ma, J. Multiscale and integrative single-cell Hi-C analysis with Higashi. Nat. Biotechnol. 40, 254–261 (2022).
Luo, C. et al. Single nucleus multi-omics identifies human cortical cell regulatory genome diversity.Cell Genom. 2, 100107 (2022).
Cardozo Gizzi, A. M. et al. Microscopy-based chromosome conformation capture enables simultaneous visualization of genome organization and transcription in intact organisms. Mol. Cell 74, 212–222.e5 (2019).
Mateo, L. J. et al. Visualizing DNA folding and RNA in embryos at single-cell resolution. Nature 568, 49–54 (2019).
Su, J.-H., Zheng, P., Kinrot, S. S., Bintu, B. & Zhuang, X. Genome-scale imaging of the 3D organization and transcriptional activity of chromatin. Cell 182, 1641–1659.e26 (2020).
Takei, Y. et al. Integrated spatial genomics reveals global architecture of single nuclei. Nature 590, 344–350 (2021).
Liu, Z. et al. Linking genome structures to functions by simultaneous single-cell Hi-C and RNA-seq. Science 380, 1070–1076 (2023).
Ramani, V. et al. Sci-Hi-C: a single-cell Hi-C method for mapping 3D genome organization in large number of single cells. Methods 170, 61–68 (2020).
Kim, H.-J. et al. Capturing cell type-specific chromatin compartment patterns by applying topic modeling to single-cell Hi-C data. PLoS Comput. Biol. 16, e1008173 (2020).
Bonora, G. et al. Single-cell landscape of nuclear configuration and gene expression during stem cell differentiation and X inactivation. Genome Biol. 22, 279 (2021).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Cusanovich, D. A. et al. Multiplex single cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Cao, J. et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357, 661–667 (2017).
Rosenberg, A. B. et al. Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360, 176–182 (2018).
Ma, S. et al. Chromatin potential identified by shared single-cell profiling of RNA and chromatin. Cell 183, 1103–1116.e20 (2020).
Lee, D.-S. et al. Simultaneous profiling of 3D genome structure and DNA methylation in single human cells. Nat. Methods 16, 999–1006 (2019).
Liu, H. et al. DNA methylation atlas of the mouse brain at single-cell resolution. Nature 598, 120–128 (2021).
Zhang, R., Zhou, T. & Ma, J. Ultrafast and interpretable single-cell 3D genome analysis with Fast-Higashi. Cell Syst. 13, 798–807.e6 (2022).
Winick-Ng, W. et al. Cell-type specialization is encoded by specific chromatin topologies. Nature 599, 684–691 (2021).
Heffel, M. G. et al. Epigenomic and chromosomal architectural reconfiguration in developing human frontal cortex and hippocampus. Preprint at bioRxiv https://doi.org/10.1101/2022.10.07.511350 (2022).
Zhang, M. et al. Spatially resolved cell atlas of the mouse primary motor cortex by MERFISH. Nature 598, 137–143 (2021).
Chidester, B., Zhou, T., Alam, S. & Ma, J. SPICEMIX enables integrative single-cell spatial modeling of cell identity. Nat. Genet. 55, 78–88 (2023).
Law, A. J., Kleinman, J. E., Weinberger, D. R. & Weickert, C. S. Disease-associated intronic variants in the ErbB4 gene are related to altered ErbB4 splice-variant expression in the brain in schizophrenia. Hum. Mol. Genet. 16, 129–141 (2007).
Zhu, C. et al. Joint profiling of histone modifications and transcriptome in single cells from mouse brain. Nat. Methods 18, 283–292 (2021).
Zhang, Y. et al. Temporal molecular program of human hematopoietic stem and progenitor cells after birth. Dev. Cell 57, 2745–2760.e6 (2022).
Nasser, J. et al. Genome-wide enhancer maps link risk variants to disease genes. Nature 593, 238–243 (2021).
Chen, A. F. et al. NEAT-seq: simultaneous profiling of intra-nuclear proteins, chromatin accessibility and gene expression in single cells. Nat. Methods 19, 547–553 (2022).
Plongthongkum, N., Diep, D., Chen, S., Lake, B. B. & Zhang, K. Scalable dual-omics profiling with single-nucleus chromatin accessibility and mRNA expression sequencing 2 (SNARE-seq2). Nat. Protoc. 16, 4992–5029 (2021).
Tan, L., Xing, D., Daley, N. & Xie, X. S. Three-dimensional genome structures of single sensory neurons in mouse visual and olfactory systems. Nat. Struct. Mol. Biol. 26, 297–307 (2019).
Mulqueen, R. M. et al. High-content single-cell combinatorial indexing. Nat. Biotechnol. 39, 1574–1580 (2021).
Collombet, S. et al. Parental-to-embryo switch of chromosome organization in early embryogenesis. Nature 580, 142–146 (2020).
Gassler, J. et al. A mechanism of cohesin-dependent loop extrusion organizes zygotic genome architecture. EMBO J. 36, 3600–3618 (2017).
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 361, 1380–1385 (2018).
Chen, S., Lake, B. B. & Zhang, K. High-throughput sequencing of the transcriptome and chromatin accessibility in the same cell. Nat. Biotechnol. 37, 1452–1457 (2019).
Zhu, C. et al. An ultra high-throughput method for single-cell joint analysis of open chromatin and transcriptome. Nat. Struct. Mol. Biol. 26, 1063–1070 (2019).
Mimitou, E. P. et al. Multiplexed detection of proteins, transcriptomes, clonotypes and CRISPR perturbations in single cells. Nat. Methods 16, 409–412 (2019).
Xu, W. et al. ISSAAC-seq enables sensitive and flexible multimodal profiling of chromatin accessibility and gene expression in single cells. Nat. Methods 19, 1243–1249 (2022).
Xiong, H., Luo, Y., Wang, Q., Yu, X. & He, A. Single-cell joint detection of chromatin occupancy and transcriptome enables higher-dimensional epigenomic reconstructions. Nat. Methods 18, 652–660 (2021).
Yao, Z. et al. A taxonomy of transcriptomic cell types across the isocortex and hippocampal formation. Cell 184, 3222–3241.e26 (2021).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Goloborodko, A., Abdennur, N. & Venev, S. hbbrandao, gfudenberg. mirnylab/pairtools: v0.3.0. Zenodo https://doi.org/10.5281/zenodo.2649383 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Zhou T. GAGE-seq analysis workflow. Zenodo https://doi.org/10.5281/zenodo.10888453 (2024).
Acknowledgements
We thank Y. Zhang for assistance with the figures. This work was primarily supported by the National Institutes of Health (NIH) grant no. R01HG012303 (J.M. and Z.D.), with additional funding, in part, provided by NIH Common Fund 4D Nucleome Program grant nos UM1HG011593 (J.M.) and UM1HG011586 (Z.D.), NIH Common Fund Cellular Senescence Network Program grant no. UG3CA268202 (J.M.) and NIH grant nos R01HG007352 (J.M.) and R61DA047010 (Z.D.). Z.D. was additionally supported by EvansMDS Discovery Research Grant 2019. J.M. received additional support from a Guggenheim Fellowship from the John Simon Guggenheim Memorial Foundation, a Google Research Collabs Award and a Single-Cell Biology Data Insights award from the Chan Zuckerberg Initiative. R.Z. was supported by the Eric and Wendy Schmidt Center at the Broad Institute. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Z.D. and J.M. conceived and oversaw the project. Z.D. conceived and developed the GAGE-seq protocol with critical contributions from T.Z. and J.M. T.Z. developed the computational workflow and performed all the data analysis with assistance from R.Z., under the supervision of Z.D. and J.M. D.J. provided mice and dissected the mouse brain tissues, under the supervision of L.X. R.T.D. and A.D.M. prepared human PBMCs for method optimization, under the supervision of J.L.A. D.G. performed experiments under the supervision of Z.D. T.Z., Z.D. and J.M. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
Z.D. is listed as the inventor on a provisional patent application that covered the GAGE-seq experimental protocol filed by the University of Washington. The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Fulai Jin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Results and Figs. 1–44.
Supplementary Tables
Supplementary Table 1. Sequences of the GAGE-seq primers. Table 2. Metadata of the single cells detected in the K562-NIH3T3 library. Table 3. Metadata of the single cells detected in the K562-GM12878 library. Table 4. Metadata of the single cells detected in the MDS-L library. Table 5. Metadata of the single cells detected in the mBCortex libraries. Table 6. Metadata of the single cells detected in the hBMCD libraries.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, T., Zhang, R., Jia, D. et al. GAGE-seq concurrently profiles multiscale 3D genome organization and gene expression in single cells. Nat Genet (2024). https://doi.org/10.1038/s41588-024-01745-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41588-024-01745-3