Abstract
Somatic mutations in cancer genes have been detected in clonal expansions across healthy human tissue, including in clonal hematopoiesis. However, because mutated and wild-type cells are admixed, we have limited ability to link genotypes with phenotypes. To overcome this limitation, we leveraged multi-modality single-cell sequencing, capturing genotype, transcriptomes and methylomes in progenitors from individuals with DNMT3A R882 mutated clonal hematopoiesis. DNMT3A mutations result in myeloid over lymphoid bias, and an expansion of immature myeloid progenitors primed toward megakaryocytic–erythroid fate, with dysregulated expression of lineage and leukemia stem cell markers. Mutated DNMT3A leads to preferential hypomethylation of polycomb repressive complex 2 targets and a specific CpG flanking motif. Notably, the hypomethylation motif is enriched in binding motifs of key hematopoietic transcription factors, serving as a potential mechanistic link between DNMT3A mutations and aberrant transcriptional phenotypes. Thus, single-cell multi-omics paves the road to defining the downstream consequences of mutations that drive clonal mosaicism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All of the processed counts matrix data and raw fastq files for murine experiments are available via Gene Expression Omnibus (GEO) under the accession number GSE158067. The raw fastq files for human samples are available via European Genome-Phenome Archive under the accession number EGAS00001006364.
Code availability
IronThrone v.2.2 pipeline is available at https://github.com/landau-lab/IronThrone-GoT
References
Martincorena, I. et al. Somatic mutant clones colonize the human esophagus with age. Science 362, 911–917 (2018).
Yizhak, K. et al. RNA sequence analysis reveals macroscopic somatic clonal expansion across normal tissues. Science 364, eaaw0726 (2019).
Yokoyama, A. et al. Age-related remodelling of oesophageal epithelia by mutated cancer drivers. Nature 565, 312–317 (2019).
Yoshida, K. et al. Tobacco smoking and somatic mutations in human bronchial epithelium. Nature 578, 266–272 (2020).
Martincorena, I. et al. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015).
Mustjoki, S. & Young, N. S. Somatic mutations in ‘benign’ disease. N. Engl. J. Med. 384, 2039–2052 (2021).
Shlush, L. I. et al. Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature 506, 328–333 (2014).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014).
Xie, M. et al. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat. Med. 20, 1472–1478 (2014).
Steensma, D. P. et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 126, 9–16 (2015).
Young, A. L., Challen, G. A., Birmann, B. M. & Druley, T. E. Clonal haematopoiesis harbouring AML-associated mutations is ubiquitous in healthy adults. Nat. Commun. 7, 12484 (2016).
Zink, F. et al. Clonal hematopoiesis, with and without candidate driver mutations, is common in the elderly. Blood 130, 742–752 (2017).
Champion, K. M., Gilbert, J. G. R., Asimakopoulos, F. A., Hinshelwood, S. & Green, A. R. Clonal haemopoiesis in normal elderly women: implications for the myeloproliferative disorders and myelodysplastic syndromes. Br. J. Haematol. 97, 920–926 (1997).
SanMiguel, J. M. et al. Cell-extrinsic stressors from the aging bone marrow (BM) microenvironment promote Dnmt3a-mutant clonal hematopoiesis. Blood 134 (Suppl), 5 (2019).
Terao, C. et al. Chromosomal alterations among age-related haematopoietic clones in Japan. Nature 584, 130–135 (2020).
Watson, C. J. et al. The evolutionary dynamics and fitness landscape of clonal hematopoiesis. Science 367, 1449–1454 (2020).
Teixeira, V. H. et al. Deciphering the genomic, epigenomic, and transcriptomic landscapes of pre-invasive lung cancer lesions. Nat. Med. 25, 517–525 (2019).
Steensma, D. P. & Ebert, B. L. Clonal hematopoiesis as a model for premalignant changes during aging. Exp. Hematol. 83, 48–56 (2020).
Desai, P. et al. Somatic mutations precede acute myeloid leukemia years before diagnosis. Nat. Med. 24, 1015–1023 (2018).
Papaemmanuil, E. et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood 122, 3616–3627 (2013).
Abelson, S. et al. Prediction of acute myeloid leukaemia risk in healthy individuals. Nature 559, 400–404 (2018).
Buscarlet, M. et al. DNMT3A and TET2 dominate clonal hematopoiesis and demonstrate benign phenotypes and different genetic predispositions. Blood 130, 753–762 (2017).
Young, A. L., Spencer Tong, R., Birmann, B. M. & Druley, T. E. Clonal hematopoiesis and risk of acute myeloid leukemia. Haematologica 104, 2410–2417 (2019).
Jeong, M. et al. Loss of Dnmt3a immortalizes hematopoietic stem cells in vivo. Cell Rep. 23, 1–10 (2018).
Ostrander, E. L. et al. Divergent effects of Dnmt3a and Tet2 mutations on hematopoietic progenitor cell fitness. Stem Cell Rep. 14, 551–560 (2020).
Koya, J. et al. DNMT3A R882 mutants interact with polycomb proteins to block haematopoietic stem and leukaemic cell differentiation. Nat. Commun. 7, 10924 (2016).
Kim, S. J. et al. A DNMT3A mutation common in AML exhibits dominant-negative effects in murine ES cells. Blood 122, 4086–4089 (2013).
Russler-Germain, D. A. et al. The R882H DNMT3A mutation associated with AML dominantly inhibits wild-type DNMT3A by blocking its ability to form active tetramers. Cancer Cell 25, 442–454 (2014).
Nam, A. S. et al. Somatic mutations and cell identity linked by genotyping of transcriptomes. Nature 571, 355–360 (2019).
Gaiti, F. et al. Epigenetic evolution and lineage histories of chronic lymphocytic leukaemia. Nature 569, 576–580 (2019).
Mouhieddine, T. H. et al. Clonal hematopoiesis is associated with adverse outcomes in multiple myeloma patients undergoing transplant. Nat. Commun. 11, 2996 (2020).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in Unique Molecular Identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Tickle, T., Tirosh, I., Georgescu, C., Brown, M. & Haas, B. inferCNV of the Trinity CTAT Project (Broad Institute of MIT and Harvard, 2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e21 (2019).
Pellin, D. et al. A comprehensive single cell transcriptional landscape of human hematopoietic progenitors. Nat. Commun. 10, 2395 (2019).
Challen, G. A. et al. Dnmt3a and Dnmt3b have overlapping and distinct functions in hematopoietic stem cells. Cell Stem Cell 15, 350–364 (2014).
Challen, G. A. et al. Dnmt3a is essential for hematopoietic stem cell differentiation. Nat. Genet. 44, 23–31 (2011).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Guryanova, O. A. et al. DNMT3A mutations promote anthracycline resistance in acute myeloid leukemia via impaired nucleosome remodeling. Nat. Med. 22, 1488–1495 (2016).
Velten, L. et al. Human haematopoietic stem cell lineage commitment is a continuous process. Nat. Cell Biol. 19, 271–281 (2017).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Clay, D. et al. CD9 and megakaryocyte differentiation. Blood 97, 1982–1989 (2001).
Noetzli, L. J., French, S. L. & Machlus, K. R.New insights into the differentiation of megakaryocytes from hematopoietic progenitors. Arterioscler. Thromb. Vasc. Biol. 39, 1288–1300 (2019).
Slupsky, J. R. et al. The platelet antigens CD9, CD42 and integrin alphaIIbbetaIIIa can be topographically associated and transduce functionally similar signals. Eur. J. Biochem. 244, 168–175 (1997).
Dai, Y. J. et al. Conditional knockin of Dnmt3a R878H initiates acute myeloid leukemia with mTOR pathway involvement. Proc. Natl Acad. Sci. USA 114, 5237–5242 (2017).
Thol, F. et al. Incidence and prognostic influence of DNMT3A mutations in acute myeloid leukemia. J. Clin. Oncol. 29, 2889–2896 (2011).
Xu, J. et al. DNMT3A Arg882 mutation drives chronic myelomonocytic leukemia through disturbing gene expression/DNA methylation in hematopoietic cells. Proc. Natl Acad. Sci. USA 111, 2620–2625 (2014).
Saito, Y. et al. Identification of therapeutic targets for quiescent, chemotherapy-resistant human leukemia stem cells. Sci. Transl. Med. 2, 17ra9 (2010).
Tsuboi, N., Asano, K., Lauterbach, M. & Mayadas, T. N. Human neutrophil Fcgamma receptors initiate and play specialized nonredundant roles in antibody-mediated inflammatory diseases. Immunity 28, 833–846 (2008).
Lanier, L. L., Corliss, B. C., Wu, J., Leong, C. & Phillips, J. H. Immunoreceptor DAP12 bearing a tyrosine-based activation motif is involved in activating NK cells. Nature 391, 703–707 (1998).
Bouchon, A., Hernández-Munain, C., Cella, M. & Colonna, M. A DAP12-mediated pathway regulates expression of CC chemokine receptor 7 and maturation of human dendritic cells. J. Exp. Med. 194, 1111–1122 (2001).
Karulf, M., Kelly, A., Weinberg, A. D. & Gold, J. A. OX40 ligand regulates inflammation and mortality in the innate immune response to sepsis. J. Immunol. 185, 4856–4862 (2010).
Leoni, C. et al. Dnmt3a restrains mast cell inflammatory responses. Proc. Natl Acad. Sci. USA 114, E1490–E1499 (2017).
Fuster, J. J. et al. Clonal hematopoiesis associated with TET2 deficiency accelerates atherosclerosis development in mice. Science 355, 842–847 (2017).
Jaiswal, S. & Libby, P. Clonal haematopoiesis: connecting ageing and inflammation in cardiovascular disease. Nat. Rev. Cardiol. 17, 137–144 (2019).
Jaiswal, S. et al. Clonal hematopoiesis and risk of atherosclerotic cardiovascular disease. N. Engl. J. Med. 377, 111–121 (2017).
Vester, S. K. et al. Nucleolin acts as the receptor for C1QTNF4 and supports C1QTNF4-mediated innate immunity modulation. J. Biol. Chem. 296, 100513 (2021).
Li, Q. et al. Identification of C1qTNF-related protein 4 as a potential cytokine that stimulates the STAT3 and NF-κB pathways and promotes cell survival in human cancer cells. Cancer Lett. 308, 203–214 (2011).
Ramalingam, P. et al. Chronic activation of endothelial MAPK disrupts hematopoiesis via NFKB dependent inflammatory stress reversible by SCGF. Nat. Commun. 11, 666 (2020).
Shen, B. et al. Integrin alpha11 is an Osteolectin receptor and is required for the maintenance of adult skeletal bone mass. eLife 8, e42274 (2019).
Ito, C. et al. Serum stem cell growth factor for monitoring hematopoietic recovery following stem cell transplantation. Bone Marrow Transpl. 32, 391–398 (2003).
Wingender, E., Dietze, P., Karas, H. & Knüppel, R. TRANSFAC: a database on transcription factors and their DNA binding sites. Nucleic Acids Res. 24, 238–241 (1996).
Ben-Porath, I. et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat. Genet. 40, 499–507 (2008).
Coller, H. A. et al. Expression analysis with oligonucleotide microarrays reveals that MYC regulates genes involved in growth, cell cycle, signaling, and adhesion. Proc. Natl Acad. Sci. USA 97, 3260–3265 (2000).
Delgado, M. D. & León, J. Myc roles in hematopoiesis and leukemia. Genes Cancer 1, 605–616 (2010).
Guo, Y. et al. c-Myc-mediated control of cell fate in megakaryocyte–erythrocyte progenitors. Blood 114, 2097–2106 (2009).
Mayers, S. et al. Establishment of an erythroid progenitor cell line capable of enucleation achieved with an inducible c-Myc vector. BMC Biotech. 19, 21 (2019).
Vaisvila, R. et al. Enzymatic methyl sequencing detects DNA methylation at single-base resolution from picograms of DNA. Genome Res. 31, 1280–1289 (2021).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Spencer, D. H. et al. CpG island hypermethylation mediated by DNMT3A is a consequence of AML progression. Cell 168, 801–816 e13 (2017).
Glass, J. L. et al. Epigenetic identity in AML depends on disruption of nonpromoter regulatory elements and is affected by antagonistic effects of mutations in epigenetic modifiers. Cancer Discov. 7, 868–883 (2017).
Vaisvila, R. et al. Enzymatic methyl sequencing detects DNA methylation at single-base resolution from picograms of DNA. Genome Res. 31, 1280–1289 (2021).
Wang, J. et al. Double restriction-enzyme digestion improves the coverage and accuracy of genome-wide CpG methylation profiling by reduced representation bisulfite sequencing. BMC Genomics 14, 11 (2013).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Davis, C. A. et al. The encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res. 46, D794–D801 (2018).
Adelman, E. R. et al. Aging human hematopoietic stem cells manifest profound epigenetic reprogramming of enhancers that may predispose to leukemia. Cancer Discov. 9, 1080–1101 (2019).
Jeong, M. et al. Large conserved domains of low DNA methylation maintained by Dnmt3a. Nat. Genet. 46, 17–23 (2014).
Zhang, X. et al. Large DNA methylation nadirs anchor chromatin loops maintaining hematopoietic stem cell identity. Mol. Cell 78, 506–521 e6 (2020).
Hawkins, R. D. et al. Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell 6, 479–491 (2010).
Margueron, R. & Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 469, 343–349 (2011).
Mohn, F. et al. Lineage-specific polycomb targets and de novo DNA methylation define restriction and potential of neuronal progenitors. Mol. Cell 30, 755–766 (2008).
Xie, H. et al. Polycomb repressive complex 2 regulates normal hematopoietic stem cell function in a developmental-stage-specific manner. Cell Stem Cell 14, 68–80 (2014).
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Emperle, M. et al. Mutations of R882 change flanking sequence preferences of the DNA methyltransferase DNMT3A and cellular methylation patterns. Nucleic Acids Res. 47, 11355–11367 (2019).
Anteneh, H., Fang, J. & Song, J. Structural basis for impairment of DNA methylation by the DNMT3A R882H mutation. Nat. Commun. 11, 2294 (2020).
Yin, Y. et al. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science 356, eaaj2239 (2017).
Prendergast, G. C. & Ziff, E. B. Methylation-sensitive sequence-specific DNA binding by the c-Myc basic region. Science 251, 186–189 (1991).
Tate, P. H. & Bird, A. P. Effects of DNA methylation on DNA-binding proteins and gene expression. Curr. Opin. Genet Dev. 3, 226–231 (1993).
Grau, J., Schmidt, F. & Schulz, M.H. Widespread effects of DNA methylation and intra-motif dependencies revealed by novel transcription factor binding models. Preprint at bioRxiv https://doi.org/10.1101/2020.10.21.348193 (2020).
Takubo, K. et al. Regulation of the HIF-1alpha level is essential for hematopoietic stem cells. Cell Stem Cell 7, 391–402 (2010).
Krock, B. L. et al. The aryl hydrocarbon receptor nuclear translocator is an essential regulator of murine hematopoietic stem cell viability. Blood 125, 3263–3272 (2015).
Mariani, C. J. et al. TET1-mediated hydroxymethylation facilitates hypoxic gene induction in neuroblastoma. Cell Rep. 7, 1343–1352 (2014).
Li, Y. et al. Setd1a and NURF mediate chromatin dynamics and gene regulation during erythroid lineage commitment and differentiation. Nucleic Acids Res. 44, 7173–7188 (2016).
Huang, S., Li, X., Yusufzai, T. M., Qiu, Y. & Felsenfeld, G. USF1 recruits histone modification complexes and is critical for maintenance of a chromatin barrier. Mol. Cell. Biol. 27, 7991–8002 (2007).
Arnaud, L. et al. A dominant mutation in the gene encoding the erythroid transcription factor KLF1 causes a congenital dyserythropoietic anemia. Am. J. Hum. Genet. 87, 721–727 (2010).
Grosselin, K. et al. High-throughput single-cell ChIP-seq identifies heterogeneity of chromatin states in breast cancer. Nat. Genet. 51, 1060–1066 (2019).
Rotem, A. et al. Single-cell ChIP-seq reveals cell subpopulations defined by chromatin state. Nat. Biotechnol. 33, 1165–1172 (2015).
Wang, Q. et al. CoBATCH for high-throughput single-cell epigenomic profiling. Mol. Cell 76, 206–216 e7 (2019).
Fu, L. et al. Predicting transcription factor binding in single cells through deep learning. Sci. Adv. 6, eaba9031 (2020).
Ugarte, F. et al. Progressive chromatin condensation and H3K9 methylation regulate the differentiation of embryonic and hematopoietic stem cells. Stem Cell Rep. 5, 728–740 (2015).
Martin, E. W. et al. Chromatin accessibility maps provide evidence of multilineage gene priming in hematopoietic stem cells. Epigenetics Chromatin 14, 2 (2021).
Granja, J. M. et al. Single-cell multiomic analysis identifies regulatory programs in mixed-phenotype acute leukemia. Nat. Biotechnol. 37, 1458–1465 (2019).
Setty, M. et al. Characterization of cell fate probabilities in single-cell data with Palantir. Nat. Biotechnol. 37, 451–460 (2019).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Zavidij, O. et al. Single-cell RNA sequencing reveals compromised immune microenvironment in precursor stages of multiple myeloma. Nat. Cancer 1, 493–506 (2020).
Izzo, F. et al. DNA methylation disruption reshapes the hematopoietic differentiation landscape. Nat. Genet. 52, 378–387 (2020).
Lessard, J. & Sauvageau, G. Bmi-1 determines the proliferative capacity of normal and leukaemic stem cells. Nature 423, 255–260 (2003).
Liu, Y. et al. Convergence of oncogenic cooperation at single-cell and single-gene levels drives leukemic transformation. Nat. Commun. 12, 6323 (2021).
Lu, R. et al. A model system for studying the DNMT3A hotspot mutation (DNMT3A(R882)) demonstrates a causal relationship between its dominant-negative effect and leukemogenesis. Cancer Res. 79, 3583–3594 (2019).
van Galen, P. et al. The unfolded protein response governs integrity of the haematopoietic stem-cell pool during stress. Nature 510, 268–272 (2014).
Rodriguez-Meira, A. et al. Unravelling intratumoral heterogeneity through high-sensitivity single-cell mutational analysis and parallel RNA sequencing. Mol. Cell 73, 1292–1305 e8 (2019).
Petti, A. A. et al. A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing. Nat. Commun. 10, 3660 (2019).
Ludwig, L. S. et al. Lineage tracing in humans enabled by mitochondrial mutations and single-cell genomics. Cell 176, 1325–1339 e22 (2019).
Nam, A. S., Chaligne, R. & Landau, D. A. Integrating genetic and non-genetic determinants of cancer evolution by single-cell multi-omics. Nat. Rev. Genet. 22, 3–18 (2021).
ENCODE Project Consortium.An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Saxonov, S., Berg, P. & Brutlag, D. L. A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proc. Natl Acad. Sci. USA 103, 1412–1417 (2006).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Gothert, J. R. et al. In vivo fate-tracing studies using the Scl stem cell enhancer: embryonic hematopoietic stem cells significantly contribute to adult hematopoiesis. Blood 105, 2724–2732 (2005).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Tange, O. GNU Parallel 20200622. Computer Program. Zenodo https://doi.org/10.5281/zenodo.3956817 (2020).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Acknowledgements
The work was enabled by the Weill Cornell Epigenomics Core and Flow Cytometry Core. We thank A. Melnick (Weill Cornell Medicine) for a critical review of the manuscript. A.S.N. is supported by the Burroughs Wellcome Fund Career Award for Medical Scientists, the National Institutes of Health Director’s Early Independence Award (DP5 OD029619) and the Starr Cancer Consortium. N.D. is supported by a F30 Predoctoral Fellowship from the National Heart, Lung, and Blood Institute (NHLBI) of the National Institutes of Health (F30HL156496). N.D. and R.M.M. are supported by a Medical Scientist Training Program grant from the National Institute of General Medical Sciences of the National Institutes of Health under award number T32GM007739 to the Weill Cornell/Rockefeller/Sloan Kettering Tri-Institutional MD-PhD Program. F.I. is supported by the American Society of Hematology Fellow-to-Faculty Scholar Award. R.C. is supported by Lymphoma Research Foundation and Marie Skłodowska-Curie fellowships. I.G. is supported by a grant from the Dr Miriam and Sheldon G. Adelson Medical Research Foundation, and a Stand Up To Cancer Dream Team Research Grant (SU2C-AACR-DT-28-18). D.A.L. is supported by the Burroughs Wellcome Fund Career Award for Medical Scientists, Valle Scholar Award, the National Institutes of Health Director’s New Innovator Award (DP2-CA239065), the Chan Zuckerberg Initiative Award, the Leukemia Lymphoma Society Career Development Program Award and the Mark Foundation Emerging Leader Award. This work was also supported by the NHLBI (R01HL145283) and the National Human Genome Research Institute, Center of Excellence in Genomic Science (RM1HG011014).
Author information
Authors and Affiliations
Contributions
A.S.N., N.D., F.I., R.M., R.C., T.H.M., I.G. and D.A.L. conceived the project, designed the study and interpreted results. I.G. and L.A.G. provided clinical samples. R.C., R.M.M., J.S., S.B. and M.W. performed the experiments. A.S.N., N.D., F.I. and R.M. performed the analyses. A.S.N., N.D., F.I., R.M., R.M.M., R.C., I.G. and D.A.L. wrote the manuscript. A.S.N., N.D., F.I., R.M., T.H.M., J.S., S.B., F.G., S.T., L.A.G., R.C., R.L., O.A.-W., I.G. and D.A.L. helped interpret results. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
O.A.-W. has served as a consultant for H3B Biomedicine, Foundation Medicine Inc., Merck Pfizer, and Janssen, and is on the scientific advisory board of Envisagenics Inc. and AIChemy; O.A.-W. has received prior research funding from H3B Biomedicine and LOXO Oncology unrelated to the current manuscript. I.G. serves on the advisory board of Bristol Myers Squibb, Takeda, Janssen, Sanofi and GlaxoSmithKline. D.A.L. has served as a consultant for Abbvie, AstraZeneca and Illumina, and is on the scientific advisory board of Mission Bio, Alethiomics, Pangea and C2i Genomics; D.A.L. has received prior research funding from BMS, 10x Genomics, Ultima Genomics, Abbvie and Illumina unrelated to the current manuscript.
Peer review
Peer review information
Nature Genetics thanks Michael Rehli, Vijay Sankaran and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 GoT captures genotyping information of thousands of CD34+ cells in scRNA-seq.
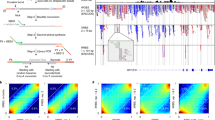
a, Summary of Genotyping of Transcriptomes (GoT) data from clonal hematopoiesis (CH) patient samples with DNMT3A R882 mutations. b, Number of genes per cell (left) and number of unique molecular identifiers (UMIs) per cell (right) from CD34+ sorted hematopoietic progenitors by patient sample after quality control (QC) filters (CH01 n = 6133, CH02 n = 5372, CH03 n = 7379, CH04 n = 8440). c, DNMT3A R882 mutant fraction of single cells determined by GoT versus DNMT3A R882 mutation variant allele frequencies (VAF) in bulk sequencing of matched unsorted stem cell product31. d, Fraction of cells by number of DNMT3A UMIs in standard 10x Genomics data without genotyping information (left), DNMT3A UMIs with R882 locus coverage in standard 10x data (middle), and DNMT3A UMIs with R882 locus coverage in GoT amplicon library (right). e, Species-mixing experiment data in which mouse cells (Ba/F3) with a human mutant CALR transgene were mixed with human cells (UT-7) with a human wildtype CALR transgene29. Mouse and human genome alignment of 10x data with genotyping data from GoT pre (top) and post (bottom) implementation of UMI consensus assembly based on Levenshtein distance (methods). f, Number of duplicate reads supporting cell barcode-UMI pair in the GoT library that is identified in the 10x gene expression (GEX) library as a DNMT3A gene (left), no gene (middle), or a non−DNMT3A gene (right). P-values from two-sided Wilcoxon rank sum test. g, Heatmap of relative expression of genes ordered by chromosome/chromosomal position following copy number variation analysis using the InferCNV package33. Cells (y-axis) are stratified by patient and DNMT3A R882 genotype status. h, Heatmap of relative expression of Y-chromosome genes following copy number variation analysis and cell stratification as in g. WT, wildtype, MUT, mutant.
Extended Data Fig. 2 Integration of DNMT3A R882 mutation and assignment of progenitor subsets in clonal hematopoiesis patient samples.
a, Uniform manifold approximation and projection (UMAP) of CD34+ progenitor cells from samples CH01-CH04 after integration using the Seurat package (methods). b, Heatmap of top 10 differentially expressed genes for progenitor subsets. c, Lineage-specific genes (left) and modules from Velten et al.41 (right, Supplementary Table 2) are scored and projected onto the UMAP representation of CD34+ cells. d, UMAP of CD34+ cells overlaid with cluster assignments, split by patient sample. HSPC, hematopoietic stem progenitor cells; IMP, immature myeloid progenitors; IMP-ME, megakaryocytic-erythroid biased IMP; IMP-GM, granulo-monocytic biased IMP; LMPP, lympho-myeloid primed progenitors; CLP, common lymphoid progenitor; MEP, megakaryocytic-erythroid progenitors; E/B/M, eosinophil, basophil, and mast cell progenitors; EP, erythroid progenitor; MkP, megakaryocytic progenitor; NP, neutrophil progenitor.
Extended Data Fig. 3 Classification of IMPs showing lineage biases and pseudotime analysis between mutated and wildtype cells.
a, UMAP of CD34+ cells, overlaid with cluster assignment of the IMP subsets. b, Neutrophil and Megakaryocytic-Erythroid lineage specific gene module scores from Velten et al.41 compared across the three IMP clusters. P-value was calculated from two-sided Wilcoxon rank sum test. c, UMAP of CD34+ cells overlaid with mutation status for wildtype (WT), DNMT3A R882 mutant (MUT), or unassigned (NA), split by genotype for all samples (top) and by patient sample (bottom). d, UMAP with projected pseudotime values (top left). Pseudotime comparison between WT and MUT cells for all samples (top right) and for individual samples (bottom) as estimated by Monocle39. P-value was calculated from likelihood ratio test of linear mixed model (LMM) with/without mutation status for aggregate analysis (methods, top) and two-sided Wilcoxon rank sum test for individual samples (bottom).
Extended Data Fig. 4 Comparison between mutated and wildtype progenitor cells of cell cycle module expression and RNA velocity derived transition probabilities.
a, Cell cycle module score represents the union of S-phase and G2M-phase gene-module expression (Supplementary Table 2)42. P-value was calculated from likelihood ratio test of linear mixed model with/without mutation status (methods). Analysis was performed for clusters with at least 200 genotyped cells across all patient samples. b, Single cell mean IMP → IMP-ME and c, IMP → IMP-GM transition probabilities, as measured via RNA velocity34, between wildtype or DNMT3A R882 mutant IMPs for each sample. P-values from two-sided Wilcoxon rank-sum test.
Extended Data Fig. 5 Comparison of differential expression analysis between permutation test and linear mixed model and MYC gene expression.
a, P-values from permutation test and linear mixed model (methods) are plotted per gene. Correlation coefficient R calculated using Pearson’s Correlation. P-values derived from two-sided Student’s t-distribution. Shading denotes 95% confidence interval. b, Normalized MYC gene expression between mutated and wildtype cells in MEP and EP. P-value was calculated from likelihood ratio test of linear mixed model with/without mutation status (methods).
Extended Data Fig. 6 Multi-omics single cell methylome, transcriptomic, and somatic genotyping reveals hypomethylation of PRC2 targets in DNMT3A R882 CH.
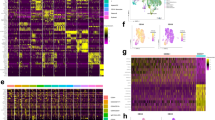
a, UMAP dimensionality reduction (n = 528 cells) based on scRNA-seq data (Smart-seq2) after integration and batch correction of six plates (methods). b, UMAP dimensionality reduction showing cluster gene markers for the transcriptome data. c, Number of CpG sites captured per cell after quality filtering (methods). The metrics for each sample according to enzymatic digestion with Msp1 (Single) or Msp1 plus HaeIII (Double) are shown. d, Average single cell methylation at all regions (global, double digest), promoters, introns or exons. P-values from likelihood ratio test of LMM with/without mutation status (methods). e, Average single cell methylation at CpH (that is CpA, CpC or CpT) sites. P-values from likelihood ratio test of LMM with/without mutation status (methods). f, Average single cell methylation at 269 hypomethylated promoters identified with differentially methylated region (DMR) analysis (shown in Fig. 4e, promoters with P-value < 0.05 and at least −5% methylation change) in CH02 and CH04. P-values from two-sided Wilcoxon rank sum test. g, Average single cell methylation at SUZ12 (top panel) and EZH2 (bottom panel) ENCODE chromatin immunoprecipitation sequencing (ChIP-seq) peaks intersected with bivalently H3K27me3, H3K4me3-marked regions in CD34+ cells for CH02 and CH04. P-values from likelihood ratio test of LMM with/without mutation status. h, Normalized expression of PRC2 target genes with preferentially hypomethylated transcription start site (TSS) (from Fig. 4e) in GoT data of WT versus MUT cells by progenitor subtype. P-values from likelihood ratio test of LMM with/without mutation status. i, Comparison of average methylation values for TSS ± 1 kb regions in normal HSPCs79 and DNMT3A WT (n = 6) versus DNMT3A R882, NPM1 mutated acute myeloid leukemia (AML; n = 7) samples in regions without (left) or with (right) PRC2 ChIP-seq peaks, controlling for CpG content. P-values from two-sided Wilcoxon rank sum test. j, Comparison of average methylation values for promoter regions in WT (n = 122) versus DNMT3A R882 mutated AML (n = 9) samples from TCGA in regions without (left) or with (right) PRC2 ChIP-seq peaks, controlling for CpG content. P-values from two-sided Wilcoxon rank sum test.
Extended Data Fig. 7 Motif enrichment at hypomethylated CpGs and hypomethylated motif enrichment in regions around differentially expressed genes.
a, Base frequency odds ratio of hypo- versus hyper-methylated CpG flanking sequences at positions N-2, N-1, N + 1, and N + 2. The odds ratios were derived from base frequencies of flanking positions of the CpG sites hypo- or hyper-methylated in mutant versus wildtype cells above the thresholds shown in the x axis for minimum absolute CpG methylation difference (Pearson correlation, P-values derived from two-sided F-test, shading denotes 95% confidence interval). b, Reported motif logos derived from Emperle et al.87 for either hypomethylated (disfavored) or hypermethylated (favored) sites for DNMT3A R882 compared with its wildtype counterpart (left). c, Similarity scores between the reported and our de novo DNMT3A R882 hypo- and hypermethylated motifs as measured by correlation coefficients of the position weight matrices for the respective motifs excluding the CpG dinucleotide. d, Heatmap of expression of transcription factors with binding motif similarity >0.5 compared with hypomethylated motif of DNMT3A R882 (that do not meet the overall expression threshold, Fig. 5a,b). e, Frequencies of DNMT3A R882 hypomethylated motif within 30 kb of TSS of the differentially expressed genes between MUT and WT cells in progenitor subsets. P-values were calculated by two-sided Wilcoxon rank sum test. f, Frequencies of DNMT3A R882 hypomethylated motif within 10 kb, 30 kb or 50 kb of TSS of the differentially expressed genes between MUT and WT cells in HSPCs and EPs. P-values were calculated by two-sided Wilcoxon rank sum test. g, Ratio of frequencies of DNMT3A R882 hypomethylated motif to those of the control shuffled motif with CpG (Fig. 5e) within 10 kb of TSS of the differentially expressed genes between MUT and WT cells in HSPCs and EPs. P-values were calculated by two-sided Wilcoxon rank sum test. h, Average per-gene incidence of DNMT3A R882 hypomethylated motif within 50 kb of TSS by distance from TSS for differentially expressed genes between MUT and WT cells in HSPCs (top) and EPs (bottom).
Extended Data Fig. 8 Single nucleus ATAC-seq of Dnmt3a R878H Lin-, c-Kit + progenitors reveals enhanced accessibility of R882 hypomethylated motif and TF motifs with high similarity scores to the hypomethylated motif.
a, Distribution of fragment size in single-nucleus assay for transposase-accessible chromatin using sequencing (snATAC-seq) data of Dnmt3a R878H and wildtype Lin-, c-Kit+ progenitors (n = 3 in each cohort). b, TSS enrichment of accessible fragments as a function of unique fragments per cell. c, UMAP of integrated datasets Dnmt3a R878H and wildtype Lin-, c-Kit+ progenitors, displayed per sample (n = 3 in each cohort). d, Heatmap of gene accessibility scores for differentially accessible progenitor identity marker genes across progenitor subsets. e, Scatterplot of similarity scores of mouse transcription factor (TF) motifs versus human TF motifs to the R882-hypomethylated motif (Pearson’s correlation, P-value derived from two-sided F-test). f, Binding motifs of mouse and human TFs with high similarity score to the R882-hypomethylated motif and expression in HSPCs (Fig. 5b, HOCOMOCO v11). g, Family-wise error rate (FWER) adjusted P-values calculated by two-sided Wilcoxon rank sum test for accessibility changes between wildtype and Dnmt3a R878H cells by progenitor identities for hypo-methylated motif and shuffled motifs controls (with and without CpG), as well as motif accessibility deviation of the TFs identified Fig. 5b (related to Fig. 5f). h, Accessibility of PRC2 targets between wildtype and Dnmt3a R878H and wildtype Lin-, c-Kit+ progenitor subsets. P-values from likelihood ratio test of LMM with/without mutation status.
Extended Data Fig. 9 Integration of CH05 and control bone marrow CD34+ scRNA-seq data, assignment of progenitor subsets, and differentiation skews of CH cells.
a, UMAP of CD34+ progenitor cells from samples CH05 and control bone marrow samples BM01-05 after integration using the Seurat package (methods). b, Number of genes per cell (top) and number of UMIs per cell (bottom) from CD34+ hematopoietic progenitors by patient sample after QC filters and down-sampling to equivalent geometric means of UMIs per patient (BM1 n = 6,683, BM2 n = 7,254, BM3 n = 16,759, BM4 n = 2,200, BM5 n = 5,690, CH05 n = 5,952). c, Heatmap of top 10 differentially expressed genes for progenitor subsets. d, UMAP representation of CD34+ cells showing cell marker gene expressions. e, Modules from Velten et al.41 (Supplementary Table 2) are scored and projected onto the UMAP representation of CD34+ cells. f, Per-patient comparison of megakaryocytic-erythroid module scores in control bone marrow versus CH05 IMPs (Supplementary Table 2). Cell number downsampled to the same number (n = 132 cells per sample). P-values were calculated from likelihood ratio test of LMM with/without CH status. g, Per-patient comparison of granulocytic-monocytic module scores in control versus CH IMPs (Supplementary Table 2). Cell number downsampled to the same number (n = 132 cells per sample). P-values were calculated from likelihood ratio test of LMM with/without CH status. h, Fraction of IMP-ME cells out of all biased IMP (IMP-ME + IMP-GM) cells in control versus CH populations. P-value was calculated from one-sample t-test. Bar represents mean values ± SEM.
Extended Data Fig. 10 Bone marrow clonal hematopoiesis patient sample confirms results from CH01-CH04 and Dnmt3a R878H mouse model.
a, Per-patient comparison of module scores for differentially down- or up-regulated genes in mutant DNMT3A HSPCs (identified in GoT data, Fig. 3a,c) in control versus CH HSPCs. P-values were calculated from likelihood ratio test of LMM with/without CH status. b, Per-patient comparison of module scores for differentially down- or up-regulated genes in mutant DNMT3A EPs (identified in GoT data, Fig. 3a,c) in control versus CH EPs. P-values were calculated from likelihood ratio test of LMM with/without CH status. c, Module scores for genes upregulated in at least 2 cell types (identified in GoT data, Fig. 3b) in control versus CH cells of major cell types. P-values from likelihood ratio test of LMM with/without CH status. d, Fraction of control BM or CH05 cells in EP1 versus EP2 cell clusters. e, UMAP of CH05 cells (clustered independently of the control BM samples) with progenitor cell assignments. f, UMAP of CH05 cells with genotyping data for WT (n = 397 cells) and DNMT3A R882 mutant (MUT; n = 290 cells). g, Normalized expression of differentially upregulated genes in at least 2 cell types, highlighted in Fig. 3b in wildtype versus mutated cells in CH05. P-values from two-sided Wilcoxon rank sum test. h, UMAP of CH05 cells with protein expression (CITE-seq) and gene expression for CD38 and CD9. i, UMAP of CH05 cells highlighting HSPCs, IMP-ME, and MkPs (left) included in the comparison of CD9 expression in wildtype versus mutated cells (right). P-values from likelihood ratio test of LMM with/without mutation status, and cell type as random effect. j, Distribution of fragment size in snATAC-seq data of patient CH05 with DNMT3A R882 CH. k, TSS enrichment of accessible fragments as a function of unique fragments per cell. l, Heatmap of the gene accessibility scores for cluster marker genes (false discovery rate (FDR) < 0.01 and Log2(Fold-Change) (Log2FC) > 1) by cell cluster. m, Pseudotime trajectories for either erythroid (left, n = 1,843 cells) or lymphoid (right, n = 1,740 cells) differentiation. n, Difference between hypomethylated and shuffled motif accessibility z-scores across either erythroid (n = 1,843 cells) or lymphoid (n = 1,740 cells) pseudotime trajectory quartiles. P-values were calculated by two-sided Wilcoxon rank sum test.
Supplementary information
Supplementary Information
Supplementary Methods.
Supplementary Tables
Supplementary Tables 1−13.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nam, A.S., Dusaj, N., Izzo, F. et al. Single-cell multi-omics of human clonal hematopoiesis reveals that DNMT3A R882 mutations perturb early progenitor states through selective hypomethylation. Nat Genet 54, 1514–1526 (2022). https://doi.org/10.1038/s41588-022-01179-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01179-9
This article is cited by
-
Cell of origin epigenetic priming determines susceptibility to Tet2 mutation
Nature Communications (2024)
-
Structure-guided functional suppression of AML-associated DNMT3A hotspot mutations
Nature Communications (2024)
-
Tissue mosaicism following stem cell aging: blood as an exemplar
Nature Aging (2024)
-
Mitochondrial genetics through the lens of single-cell multi-omics
Nature Genetics (2024)
-
Clonal hematopoiesis and its impact on the aging osteo-hematopoietic niche
Leukemia (2024)