Featured
-
-
Article
| Open Access Analysis of somatic mutations in whole blood from 200,618 individuals identifies pervasive positive selection and novel drivers of clonal hematopoiesis
Analysis of somatic mutations in whole blood from 200,618 individuals identifies pervasive positive selection and novel drivers of clonal hematopoiesis
Fitness-based analysis of 200,618 UK Biobank exomes and single-cell-derived hematopoietic clones identifies 17 genes under positive selection, including novel drivers of clonal hematopoiesis.
- Nicholas Bernstein
- , Michael Spencer Chapman
- & Jyoti Nangalia
-
Technical Report
| Open Access Direct transposition of native DNA for sensitive multimodal single-molecule sequencing
Direct transposition of native DNA for sensitive multimodal single-molecule sequencing
Two low-input tagmentation-based long-read sequencing methods, single-molecule real-time sequencing by tagmentation (SMRT-Tag), which identifies genetic variation and CpG methylation, and single-molecule adenine-methylated oligonucleosome sequencing assay by tagmentation (SAMOSA-Tag), which detects chromatin accessibility, are presented. Application of SAMOSA-Tag to prostate cancer patient-derived xenograft samples identifies metastasis-associated epigenomic alterations.
- Arjun S. Nanda
- , Ke Wu
- & Vijay Ramani
-
News & Views |
 Contraction or sequence variant of an intergenic repeat-Alu element leads to inherited thyroid disease
Contraction or sequence variant of an intergenic repeat-Alu element leads to inherited thyroid disease
Genomic and epigenomic techniques identify a new variation type causing Mendelian disease by altering the non-coding regulatory network in thyroid cells — solving a hidden cause linked for 20 years.
- Andrea Cortese
- , Elisa Vegezzi
- & Henry Houlden
-
Article
| Open Access Genomic analyses reveal the stepwise domestication and genetic mechanism of curd biogenesis in cauliflower
Genomic analyses reveal the stepwise domestication and genetic mechanism of curd biogenesis in cauliflower
A high-quality reference genome assembly of cauliflower C-8 (V2) and genomic analyses of 971 diverse accessions and their relatives reveal the stepwise domestication and the genetic mechanism of curd biogenesis.
- Rui Chen
- , Ke Chen
- & Deling Sun
-
Article |
 Genomic variation in weedy and cultivated broomcorn millet accessions uncovers the genetic architecture of agronomic traits
Genomic variation in weedy and cultivated broomcorn millet accessions uncovers the genetic architecture of agronomic traits
A comprehensive variation map constructed by deep sequencing 1,904 accessions of weedy and cultivated broomcorn millet sheds light on the genetic architecture of agronomic traits during domestication.
- Qiong Lu
- , Hainan Zhao
- & Weibin Song
-
Article |
 Cell subtype-specific effects of genetic variation in the Alzheimer’s disease brain
Cell subtype-specific effects of genetic variation in the Alzheimer’s disease brain
Single-nucleus RNA sequencing from the dorsolateral prefrontal cortex of 424 aging individuals, and mapping the effect of genetic variation on gene expression, identified a large number of cis-expression quantitative trait loci at the level of cell types and cell subtypes.
- Masashi Fujita
- , Zongmei Gao
- & Philip L. De Jager
-
Article
| Open Access Inherited polygenic effects on common hematological traits influence clonal selection on JAK2V617F and the development of myeloproliferative neoplasms
Inherited polygenic effects on common hematological traits influence clonal selection on JAK2V617F and the development of myeloproliferative neoplasms
Inherited polygenic scores for blood cell traits are associated with an increased risk of JAK2V617F clonal expansion and influence clinical phenotypes in individuals with myeloproliferative neoplasms.
- Jing Guo
- , Klaudia Walter
- & Nicole Soranzo
-
Letter
| Open Access Primate-specific ZNF808 is essential for pancreatic development in humans
Primate-specific ZNF808 is essential for pancreatic development in humans
Loss-of-function mutations in primate-specific ZNF808 cause pancreatic agenesis. Mechanistically, the loss of ZNF808 leads to the activation of the MER11 family of transposable elements in a regulatory capacity that ultimately induces a liver-specific program of gene expression during pancreatic differentiation.
- Elisa De Franco
- , Nick D. L. Owens
- & Michael Imbeault
-
-
Article
| Open Access Single-molecule genome-wide mutation profiles of cell-free DNA for non-invasive detection of cancer
Single-molecule genome-wide mutation profiles of cell-free DNA for non-invasive detection of cancer
Somatic mutations are identified from circulating cell-free DNA using a single-molecule-based lowpass whole-genome sequencing method. The regional distribution of mutations can identify a tumor-specific mutational profile in patients with cancer and can be used to monitor patients through treatment.
- Daniel C. Bruhm
- , Dimitrios Mathios
- & Victor E. Velculescu
-
-
Brief Communication
| Open Access Imputation of low-coverage sequencing data from 150,119 UK Biobank genomes
Imputation of low-coverage sequencing data from 150,119 UK Biobank genomes
GLIMPSE2 is an improved method using sparse models for accurate, efficient and cost-effective genotype imputation in low-coverage whole-genome sequencing data.
- Simone Rubinacci
- , Robin J. Hofmeister
- & Olivier Delaneau
-
Article |
 Single-cell multi-omics of mitochondrial DNA disorders reveals dynamics of purifying selection across human immune cells
Single-cell multi-omics of mitochondrial DNA disorders reveals dynamics of purifying selection across human immune cells
Single-cell analyses of immune cells from patients with pathogenic, single large-scale mitochondrial DNA (mtDNA) deletions including Pearson syndrome describe heteroplasmy dynamics consistent with purifying selection, as well as T-cell state-specific regulatory mechanisms and metabolic vulnerabilities.
- Caleb A. Lareau
- , Sonia M. Dubois
- & Leif S. Ludwig
-
Technical Report
| Open Access Mapping interindividual dynamics of innate immune response at single-cell resolution
Mapping interindividual dynamics of innate immune response at single-cell resolution
GASPACHO is a statistical method that identifies nonlinear dynamic genetic effects using single-cell RNA-seq data. Analysis of an antiviral response in human fibroblasts identifies 1,275 expression QTLs, many of which colocalize with risk loci for autoimmune and infectious diseases.
- Natsuhiko Kumasaka
- , Raghd Rostom
- & Sarah A. Teichmann
-
-
Research Briefing |
 Disentangling extrachromosomal circular DNA heterogeneity in single cells with scEC&T-seq
Disentangling extrachromosomal circular DNA heterogeneity in single cells with scEC&T-seq
We introduce scEC&T-seq, a new single-cell sequencing method that enables parallel profiling of extrachromosomal circular DNA and mRNAs in single cells. Using scEC&T-seq, we characterized all types of circular DNA elements in single human cancer cells and profiled the intercellular heterogeneity and structural dynamics of cancer-specific extrachromosomal DNA.
-
Technical Report
| Open Access Single duplex DNA sequencing with CODEC detects mutations with high sensitivity
Single duplex DNA sequencing with CODEC detects mutations with high sensitivity
Concatenating Original Duplex for Error Correction (CODEC) is a method that concatenates both strands of each DNA duplex to enable highly sensitive mutation detection in a range of analytes with fewer reads and lower error rates than current methods.
- Jin H. Bae
- , Ruolin Liu
- & Viktor A. Adalsteinsson
-
-
Brief Communication |
 Adjusting for common variant polygenic scores improves yield in rare variant association analyses
Adjusting for common variant polygenic scores improves yield in rare variant association analyses
Adjusting for common variant polygenic scores improves yield in gene-based rare variant association tests for quantitative traits, particularly when using sparse mixed models or simple linear models as an alternative to dense mixed-model approaches.
- Sean J. Jurgens
- , James P. Pirruccello
- & Patrick T. Ellinor
-
-
-
Research Briefing |
 sortChIC enables detailed chromatin analysis of rare cell types
sortChIC enables detailed chromatin analysis of rare cell types
Current methods of chromatin analysis focus mainly on the most abundant cell types in a sample. We present a workflow that combines enrichment of rare cell types with high-resolution mapping of histone modifications, which enables us to study chromatin dynamics in rare stem and progenitor cell populations.
-
Article
| Open Access Notch1 mutations drive clonal expansion in normal esophageal epithelium but impair tumor growth
Notch1 mutations drive clonal expansion in normal esophageal epithelium but impair tumor growth
Notch1 mutations have opposing effects on clonal growth in normal and tumor cells of the mouse esophagus. In a mouse model of squamous esophageal tumorigenesis, Notch1 blockade reduced premalignant tumor growth, suggesting that it might be an effective prevention strategy for the disease.
- Emilie Abby
- , Stefan C. Dentro
- & Philip H. Jones
-
Article |
 De novo genome assembly and analyses of 12 founder inbred lines provide insights into maize heterosis
De novo genome assembly and analyses of 12 founder inbred lines provide insights into maize heterosis
De novo genome assembly and analyses of 12 maize FILs provide insights into genomic and phenotypic differentiation of various heterotic groups and the molecular basis of heterosis in maize.
- Baobao Wang
- , Mei Hou
- & Haiyang Wang
-
-
Technical Report |
 Powerful, scalable and resource-efficient meta-analysis of rare variant associations in large whole genome sequencing studies
Powerful, scalable and resource-efficient meta-analysis of rare variant associations in large whole genome sequencing studies
MetaSTAAR enables functionally informed rare variant association analysis in biobank-scale cohorts using an efficient, sparse matrix approach for summary statistic storage.
- Xihao Li
- , Corbin Quick
- & Xihong Lin
-
Article |
 Dietary stress remodels the genetic architecture of lifespan variation in outbred Drosophila
Dietary stress remodels the genetic architecture of lifespan variation in outbred Drosophila
An analysis of the effects of dietary stress in outbred Drosophila shows that lifespan has a polygenic architecture and is subject to environmental influence, suggesting that this context dependency is important for complex trait variation and evolution.
- Luisa F. Pallares
- , Amanda J. Lea
- & Julien F. Ayroles
-
Technical Report
| Open Access Single-cell sortChIC identifies hierarchical chromatin dynamics during hematopoiesis
Single-cell sortChIC identifies hierarchical chromatin dynamics during hematopoiesis
Sort-assisted single-cell chromatin immunocleavage (sortChIC) combines single-cell histone modification profiling with fluorescence-activated cell sorting (FACS), enabling the study of rare cell populations. H3K4me1/H3K4me3, H3K9me3 and H3K27me3 profiling of blood suggest a model of lineage-shared repressive and cell type-specific active chromatin.
- Peter Zeller
- , Jake Yeung
- & Alexander van Oudenaarden
-
Research Briefing |
 Beating the odds: molecular characteristics of long-term survivors of ovarian cancer
Beating the odds: molecular characteristics of long-term survivors of ovarian cancer
High-grade serous ovarian cancer, the most common form of the disease, is often fatal. This study investigated the genomic and immune characteristics of tumors from women who survived more than 10 years after their initial diagnosis, and compared them with short-term and moderate-term survivors.
-
Article |
 The genomic and immune landscape of long-term survivors of high-grade serous ovarian cancer
The genomic and immune landscape of long-term survivors of high-grade serous ovarian cancer
A genomic and transcriptomic analysis identifies molecular features associated with long-term survival in ovarian cancer. Exceptional survival was heterogeneous across the cohort, suggesting that it is likely the function of multiple cell-intrinsic and microenvironmental factors working in combination.
- Dale W. Garsed
- , Ahwan Pandey
- & David D. L. Bowtell
-
Comment |
GATTACA is still pertinent 25 years later
It has been 25 years since the release of GATTACA, a film that tells the story of a credible near future in which society’s inequalities, formerly associated with race and class, have been replaced with new prejudices based on genetic determinism. Here we compare GATTACA’s fictional technologies with reality’s state of the art, assessing the legal protections afforded in today’s society against GATTACA’s dystopian future in which personal freedom and privacy rights are substantially curtailed by genomic innovations. We further discuss how GATTACA’s prescient forewarnings are still relevant today in light of the current trajectory of genomic science and technology.
- Dov Greenbaum
- & Mark Gerstein
-
-
-
Letter |
 A combined polygenic score of 21,293 rare and 22 common variants improves diabetes diagnosis based on hemoglobin A1C levels
A combined polygenic score of 21,293 rare and 22 common variants improves diabetes diagnosis based on hemoglobin A1C levels
This study presents a method for constructing complex trait polygenic scores from rare variants and shows that a polygenic score combining common and rare variants improves the accuracy of diagnosis for type 2 diabetes based on hemoglobin A1C levels.
- Peter Dornbos
- , Ryan Koesterer
- & Jason Flannick
-
Technical Report |
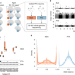 Single-cell genome sequencing of human neurons identifies somatic point mutation and indel enrichment in regulatory elements
Single-cell genome sequencing of human neurons identifies somatic point mutation and indel enrichment in regulatory elements
Single-cell DNA sequencing data are generated from human neurons using primary template-directed amplification and analyzed using SCAN2, an improved genotyping tool. Indels are enriched in neuronal regulatory elements and may be deleterious.
- Lovelace J. Luquette
- , Michael B. Miller
- & Peter J. Park
-
Research Briefing |
 Large-scale genomic analysis of human iPSCs identifies recurrent somatic driver mutations
Large-scale genomic analysis of human iPSCs identifies recurrent somatic driver mutations
The genetic landscape of human induced pluripotent stem cells (iPSCs) is strongly influenced by the somatic cells of origin, and mutational signatures directly reflect pre-reprogramming and post-reprogramming mutagenic processes. BCOR mutations are recurrent and have functional consequences for the differentiation capacity of iPSCs.
-
Article
| Open Access Integrating de novo and inherited variants in 42,607 autism cases identifies mutations in new moderate-risk genes
Integrating de novo and inherited variants in 42,607 autism cases identifies mutations in new moderate-risk genes
An integrated analysis of de novo and inherited coding variants in 42,607 individuals with autism spectrum disorder identifies 60 risk genes of which five have not previously been associated with neurodevelopmental disorders.
- Xueya Zhou
- , Pamela Feliciano
- & Wendy K. Chung
-
Article
| Open Access Genomic analysis defines clonal relationships of ductal carcinoma in situ and recurrent invasive breast cancer
Genomic analysis defines clonal relationships of ductal carcinoma in situ and recurrent invasive breast cancer
A genomic analysis of ductal carcinoma in situ (DCIS) samples with matched ipsilateral invasive breast cancer recurring later shows that around 18% of tumors were unrelated to the DCIS, and had distinct clonal origins.
- Esther H. Lips
- , Tapsi Kumar
- & Elinor J. Sawyer
-
Comment |
 Promoting the genomic revolution in Africa through the Nigerian 100K Genome Project
Promoting the genomic revolution in Africa through the Nigerian 100K Genome Project
To leverage the genetic diversity in Nigeria, we established the Non-Communicable Diseases Genetic Heritage Study (NCD-GHS) consortium to help produce a comprehensive catalog of human genetic variation in Nigeria and assess the burden and etiological characteristics of non-communicable diseases in 100,000 adults in Nigeria.
- Segun Fatumo
- , Aminu Yakubu
- & Abasi Ene-Obong
-
Article |
 Single-cell analysis of somatic mutations in human bronchial epithelial cells in relation to aging and smoking
Single-cell analysis of somatic mutations in human bronchial epithelial cells in relation to aging and smoking
Single-cell whole-genome sequencing of proximal bronchial basal cells shows that somatic mutations accumulate with age and at a higher level in smokers compared to never-smokers. Mutation frequencies increased with smoking dose but then plateaued, suggesting intrinsic mechanisms to limit mutation burden.
- Zhenqiu Huang
- , Shixiang Sun
- & Jan Vijg
-
Article |
 Exome sequencing in bipolar disorder identifies AKAP11 as a risk gene shared with schizophrenia
Exome sequencing in bipolar disorder identifies AKAP11 as a risk gene shared with schizophrenia
Exome sequencing analysis of 13,933 individuals with bipolar disorder finds enrichment of ultra-rare protein-truncating variants in constrained genes. Combined analysis with schizophrenia exome data identifies AKAP11 as a risk gene for both disorders.
- Duncan S. Palmer
- , Daniel P. Howrigan
- & Benjamin M. Neale
-
News & Views |
 Discovering missing heritability in whole-genome sequencing data
Discovering missing heritability in whole-genome sequencing data
The gap between heritability estimates from twin studies and those from genotyping array data has puzzled researchers for over a decade. New research suggests that much of the ‘missing’ heritability is due to rare variants that can only be captured by whole-genome sequencing (WGS) data.
- Alexander I. Young
-
Article |
 Analysis of rare genetic variation underlying cardiometabolic diseases and traits among 200,000 individuals in the UK Biobank
Analysis of rare genetic variation underlying cardiometabolic diseases and traits among 200,000 individuals in the UK Biobank
Analysis of whole-exome sequencing data from over 200,000 individuals in the UK Biobank provides new insights into the contribution of rare variants to cardiometabolic diseases and traits.
- Sean J. Jurgens
- , Seung Hoan Choi
- & Patrick T. Ellinor
-
Article |
 Chromothripsis followed by circular recombination drives oncogene amplification in human cancer
Chromothripsis followed by circular recombination drives oncogene amplification in human cancer
Seismic amplifications arise from several cycles of circular recombination of circular extrachromosomal DNA formed as a result of chromothripsis. The process provides a mechanism for oncogene amplification in a number of different human tumor types.
- Carolina Rosswog
- , Christoph Bartenhagen
- & Matthias Fischer
-
-
Article |
 Somatic mutations of GNA11 and GNAQ in CTNNB1-mutant aldosterone-producing adenomas presenting in puberty, pregnancy or menopause
Somatic mutations of GNA11 and GNAQ in CTNNB1-mutant aldosterone-producing adenomas presenting in puberty, pregnancy or menopause
Sequence analysis identifies gain-of-function somatic mutations in GNA11 or GNAQ in CTNNB1-mutant aldosterone-producing adenomas. Most patients with these mutations presented during puberty, pregnancy or menopause, with elevated LHCGR expression.
- Junhua Zhou
- , Elena A. B. Azizan
- & Morris J. Brown
-
Article
| Open Access High-quality genome assembly and resequencing of modern cotton cultivars provide resources for crop improvement
High-quality genome assembly and resequencing of modern cotton cultivars provide resources for crop improvement
High-quality genomes of two cultivated tetraploid cottons Gossypium hirsutum cv. NDM8 and Gossypium barbadense acc. Pima90 and resequencing of 1,081 G. hirsutum accessions provide insights into the role of structural variations.
- Zhiying Ma
- , Yan Zhang
- & Xingfen Wang
-
-
Comment |
 A call for direct sequencing of full-length RNAs to identify all modifications
A call for direct sequencing of full-length RNAs to identify all modifications
For most organisms, DNA sequences are available, but the complete RNA sequences are not. Here, we call for technologies to sequence full-length RNAs with all their modifications.
- Juan D. Alfonzo
- , Jessica A. Brown
- & Robert L. Ross

 GAGE-seq concurrently profiles multiscale 3D genome organization and gene expression in single cells
GAGE-seq concurrently profiles multiscale 3D genome organization and gene expression in single cells