Key Points
-
In the past 10 years, there has been great progress in the development of fluorescent proteins (FPs) to study protein movement and protein interactions. Green FP (GFP)-like proteins have been mutated to be monomers as useful tags for analysing protein movement.
-
FPs have been used to study bulk protein movement, primarily through photobleaching or photoactivation techniques, which allow the determination of protein diffusion and protein binding kinetics.
-
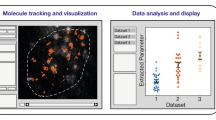
FPs have also been used to track single proteins within the cell. The technique may vary depending on properties of the protein; for example, whether it is soluble.
-
By studying movement, insights have been obtained on protein–protein interactions. Fluorescence cross-correlation spectroscopy (FCCS) has been used to detect the synchronous movement of two proteins fused to different FPs. Bimolecular fluorescence resonance energy transfer (FRET) and bimolecular fluorescence complementation (BiFC) have both been used to study protein proximity, an indicator of protein interactions.
Abstract
Proteins are always on the move, and this may occur through diffusion or active transport. The realization that the regulation of signal transduction is highly dynamic in space and time has stimulated intense interest in the movement of proteins. Over the past decade, numerous new technologies using fluorescent proteins have been developed, allowing us to observe the spatiotemporal dynamics of proteins in living cells. These technologies have greatly advanced our understanding of protein dynamics, including protein movement and protein interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Tsien, R. Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Miyawaki, A. Innovations in the imaging of brain functions using fluorescent proteins. Neuron 48, 189–199 (2005).
Shaner, N. C., Steinbach, P. A. & Tsien, R. Y. A guide to choosing fluorescent proteins. Nature Methods 2, 905–909 (2005).
Lin, M. Z., Miyawaki, A. & Tsien, R. Y. Fluorescent proteins illuminate cell biology (Poster). Nature Rev. Mol. Cell Biol. 10 (2010).
Shaner, N. C., Patterson, G. H. & Davidson, M. W. Advances in fluorescent protein technology. J. Cell Sci. 120, 4247–4260 (2007).
Day, R. N. & Davidson, M. W. The fluorescent protein palette: tools for cellular imaging. Chem. Soc. Rev. 38, 2887–2921 (2009).
Chudakov, D. M., Matz, M. V., Lukyanov, S. & Lukyanov, K. A. Fluorescent proteins and their applications in imaging living cells and tissues. Physiol. Rev. 90, 1103–1163 (2010).
Lippincott-Schwartz, J., Snapp, E. & Kenworthy, A. Studying protein dynamics in living cells. Nature Rev. Mol. Cell Biol. 2, 444–456 (2001).
Yokoe, H. & Meyer, T. Spatial dynamics of GFP-tagged proteins investigated by local fluorescence enhancement. Nature Biotech. 14, 1252–1256 (1996).
Brejc, K. et al. Structural basis for dual excitation and photoisomerization of the Aequorea victoria green fluorescent protein. Proc. Natl Acad. Sci. USA 94, 2306–2311 (1997).
Matz, M. V. et al. Fluorescent proteins from nonbioluminescent Anthozoa species. Nature Biotech. 17, 969–973 (1999).
Patterson, G. H. & Lippincott-Schwartz, J. A photoactivatable GFP for selective photolabeling of proteins and cells. Science 297, 1873–1877 (2002).
Ando, R., Hama, H., Yamamoto-Hino, M., Mizuno, H. & Miyawaki, A. An optical marker based on the UV-induced green-to-red photoconversion of a fluorescent protein. Proc. Natl Acad. Sci. USA 99, 12651–12656 (2002).
Phair, R. D. & Misteli, T. Kinetic modelling approaches to in vivo imaging. Nature Rev. Mol. Cell Biol. 2, 898–907 (2001).
Brandizzi, F., Fricker, M. & Hawes, C. A greener world: the revolution in plant bioimaging. Nature Rev. Mol. Cell Biol. 3, 520–530 (2002).
Zhang, J., Campbell, R. E., Ting, A. Y. & Tsien, R. Y. Creating new fluorescent probes for cell biology. Nature Rev. Mol. Cell Biol. 3, 906–918 (2002).
Lukyanov, K. A., Chudakov, D. M., Lukyanov, S. & Verkhusha, V. V. Photoactivatable fluorescent proteins. Nature Rev. Mol. Cell Biol. 6, 885–891 (2005).
Kerppola, T. K. Visualization of molecular interactions by fluorescence complementation. Nature Rev. Mol. Cell Biol. 7, 449–456 (2006).
Pepperkok, R. & Ellenberg, J. High-throughput fluorescence microscopy for systems biology. Nature Rev. Mol. Cell Biol. 7, 690–696 (2006).
Fernández-Suárez, M. & Ting, A. Y. Fluorescent probes for super-resolution imaging in living cells. Nature Rev. Mol. Cell Biol. 9, 929–943 (2008).
Shav-Tal, Y., Singer, R. H. & Darzacq, X. Imaging gene expression in single living cells. Nature Rev. Mol. Cell Biol. 5, 855–861 (2004).
Dehmelt, L. & Bastiaens, P. I. Spatial organization of intracellular communication: insights from imaging. Nature Rev. Mol. Cell Biol. 11, 440–452 (2010).
Yang, M., Jiang, P. & Hoffman, R. M. Whole-body subcellular multicolor imaging of tumor-host interaction and drug response in real time. Cancer Res. 67, 5195–5200 (2007).
Nowotschin, S., Eakin, G. S. & Hadjantonakis, A. K. Live-imaging fluorescent proteins in mouse embryos: multi-dimensional, multi-spectral perspectives. Trends Biotechnol. 27, 266–276 (2009).
Chudakov, D. M., Lukyanov, S. & Lukyanov, K. A. Fluorescent proteins as a toolkit for in vivo imaging. Trends Biotechnol. 23, 605–613 (2005).
Labas, Y. A. et al. Diversity and evolution of the green fluorescent protein family. Proc. Natl Acad. Sci. USA 99, 4256–4261 (2002).
Matz, M. V., Lukyanov, K. A. & Lukyanov, S. A. Family of the green fluorescent protein: journey to the end of the rainbow. Bioessays 24, 953–959 (2002).
Shagin, D. A. et al. GFP-like proteins as ubiquitous metazoan superfamily: evolution of functional features and structural complexity. Mol. Biol. Evol. 21, 841–850 (2004).
Zacharias, D. A., Violin, J. D., Newton, A. C. & Tsien, R. Y. Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells. Science 296, 913–916 (2002).
Campbell, R. E. et al. A monomeric red fluorescent protein. Proc. Natl Acad. Sci. USA 99, 7877–7882 (2002).
Wall, M. A., Socolich, M. & Ranganathan, R. The structural basis for red fluorescence in the tetrameric GFP homolog DsRed. Nature Struct. Biol. 7, 1133–1138 (2000).
Yarbrough, D., Wachter, R. M., Kallio, K., Matz, M. V. & Remington, S. J. Refined crystal structure of DsRed, a red fluorescent protein from coral, at 2.0-Å resolution. Proc. Natl Acad. Sci. USA 98, 462–467 (2001).
Shaner, N. C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nature Biotech. 22, 1567–1572 (2004).
Wiedenmann, J. et al. EosFP, a fluorescent marker protein with UV-inducible green-to-red fluorescence conversion Proc. Natl Acad. Sci. USA 101, 15905–15910 (2004).
Yanushevich, Y. G. et al. A strategy for the generation of non-aggregating mutants of Anthozoa fluorescent proteins. FEBS Lett. 511, 11–14 (2002).
Katayama, H., Yamamoto, A., Yoshimori, T., Mizushima, N. & Miyawaki, A. GFP-like proteins stably accumulate in lysosomes. Cell Struct. Funct. 33, 1–12 (2008).
Tsutsui, H., Karasawa, S., Shimizu, H., Nukina, N. & Miyawaki, A. Semi-rational engineering of a coral fluorescent protein into an efficient highlighter. EMBO Rep. 6, 233–238 (2005).
Shimozono, S., Tsutsui, H. & Miyawaki, A. Diffusion of large molecules into assembling nuclei revealed using an optical highlighting techniques. Biophys. J. 97, 1288–1294 (2009).
Lauf, U., Lopez, P. & Falk, M. M. Expression of fluorescently tagged connexins, a novel approach to rescue function of oligomeric DsRed-tagged proteins. FEBS Lett. 498, 11–15 (2001).
Bulina, M. E., Verkhusha, V. V., Staroverov, D. B., Chudakov, D. M. & Lukyanov, K. A. Hetero-oligomeric tagging diminishes non-specific aggregation of target proteins fused with Anthozoa fluorescent proteins. Biochem. J. 371, 109–114 (2003).
Axelrod, D., Koppel, D. E., Schlessinger, J., Elson, E. & Webb, W. W. Mobility measurement by analysis of fluorescence photobleaching recovery kinetics. Biophys. J. 16, 1055–1069 (1976).
Jacobson, K. A. et al. Cellular determinants of the lateral mobility of neural cell adhesion molecules. Biochim. Biophys. Acta 1330, 138–144 (1997).
Ellenberg, J. et al. Nuclear membrane dynamics and reassembly in living cells: targeting of an inner nuclear membrane protein in interphase and mitosis. J. Cell Biol. 138, 1193–1206 (1997).
Sprague, B. L. & McNally, J. G. FRAP analysis of binding: proper and fitting. Trends Cell Biol. 15, 84–91 (2005).
Rabut, G. & Ellenberg, J. in Live Cell Imaging (eds Goldman, R. D. & Spector, D. L.) 101–127 (Cold Spring Harbor Laboratory Press, New York, 2005).
Cole, N. B. et al. Diffusional mobility of Golgi proteins in membranes of living cells. Science 273, 797–801 (1996).
Kohler, R. H., Cao, J., Zipfel, W. R., Webb, W. W. & Hanson, M. R. Exchange of protein molecules through connections between higher plant plastids. Science 276, 2039–2042 (1997).
Zaal, K. J. et al. Golgi membranes are absorbed into and remerge from the ER during mitosis. Cell 99, 589–601 (1999).
Kruhlak, M. J. et al. Reduced mobility of the alternate splicing factor (ASF) through the nucleoplasm and steady state speckle compartments. J. Cell Biol. 150, 41–51 (2000).
Swaminathan, R., Hoang, C. P. & Verkman, A. S. Photobleaching recovery and anisotropy decay of green fluorescent protein GFP-S65T in solution and cells: cytoplasmic viscosity probed by green fluorescent protein translational and rotational diffusion. Biophys. J. 72, 1900–1907 (1997).
Partikian, A., Olveczky, B., Swaminathan, R., Li, Y. & Verkman, A. S. Rapid diffusion of green fluorescent protein in the mitochondrial matrix. J. Cell Biol. 140, 821–829 (1998).
Dayel, M. J., Hom, E. F. Y. & Verkman, A. S. Diffusion of green fluorescent protein in the aqueous-phase lumen of endoplasmic reticulum. Biophys. J. 76, 2843–2851 (1999).
Lippincott-Schwartz, J. & Patterson, G. H. Photoactivatable fluorescent proteins for diffraction-limited and super-resolution imaging. Trends Cell Biol. 19, 555–565 (2009).
Sample, V., Newman, R. H. & Zhang, J. The structure and function of fluorescent proteins. Chem. Soc. Rev. 38, 2582–2864 (2009).
Verkhusha, V. V. & Sorkin, A. Conversion of the monomeric red fluorescent protein into a photoactivatable probe. Chem. Biol. 12, 279–285 (2005).
Subach, F. V. et al. Photoactivatable mCherry for high-resolution two-color fluorescence microscopy. Nature Methods 6, 153–159 (2009).
Subach, F. V., Patterson, G. H., Renz, M., Lippincott-Schwartz, J. & Verkhusha, V. V. Bright monomeric photoactivatable red fluorescent protein for two-color super-resolution sptPALM of live cells. J. Am. Chem. Soc. 132, 6481–6491 (2010).
Matsuda, T., Miyawaki, A. & Nagai, T. Direct measurement of protein dynamics inside cells using a rationally designed photoconvertible protein. Nature Methods 5, 339–345 (2008).
McKinney, S. A., Murphy, C. S., Hazelwood, K. L., Davidson, M. W. & Looger, L. L. A bright and photostable photoconvertible fluorescent protein. Nature Methods 6, 131–133 (2009).
Habuchi, S., Tsutsui, H., Kochaniak, A. B., Miyawaki, A. & van Oijen, A. M. mKikGR, a monomeric photoswitchable fluorescent protein. PloS ONE 3, e3944 (2008).
Gurskaya, N. G. et al. Engineering of a monomeric green-to-red photoactivatable fluorescent protein induced by blue light. Nature Biotech. 24, 461–465 (2006).
Fuchs, J. et al. A photoactivatable marker protein for pulse-chase imaging with superresolution. Nature Methods 7, 627–630 (2010).
Chudakov, D. M. et al. Photoswitchable cyan fluorescent protein for protein tracking. Nature Biotech. 22, 1435–1439 (2004).
Subach, O. M. et al. A photoswitchable orange-to-far-red fluorescent protein, PSmOrange. Nature Methods 8, 771–777 (2011).
Bergeland, T., Haugen, L., Landsverk, O. J., Stenmark, H. & Bakke, O. Cell-cycle-dependent binding kinetics for the early endosomal tethering factor EEA1. EMBO Rep. 9, 171–178 (2008).
Plachta, N., Bollenbach, T., Pease, S., Fraser, S. E. & Pantazis, P. Oct4 kinetics predict cell lineage patterning in the early mammalian embryo. Nature Cell Biol. 13, 117–123 (2011).
Svoboda, K., Tank, D. W. & Denk, W. Direct measurement of coupling between dendritic spines and shafts. Science 272, 716–719 (1996).
Helmchen, F. & Denk, W. Deep tissue two-photon microscopy. Nature Methods 2, 932–940 (2005).
Brown, E. B., Wu, E. S., Zipfel, W. & Webb, W. W. Measurement of molecular diffusion in solution by multiphoton fluorescence photobleaching recovery. Biophys. J. 77, 2837–2849 (1999).
Star, E. N., Kwiatkowski, D. J. & Murthy, V. N. Rapid turnover of actin in dendritic spines and its regulation by activity. Nature Neurosci. 5, 239–246 (2002).
Honkura, N., Matsuzaki, M., Noguchi, J., Ellis-Davies, G. C. & Kasai, H. The subspine organization of actin fibers regulates the structure and plasticity of dendritic spines. Neuron 57, 719–729 (2008).
Gray, N. W., Weimer, R. M., Bureau, I. & Svoboda, K. Rapid redistribution of synaptic PSD-95 in the neocortex in vivo. PLoS Biol. 4, e370 (2006).
Sturgill, J. F., Steiner, P., Czervionke, B. L. & Sabatini, B. L. Distinct domains within PSD-95 mediate synaptic incorporation, stabilization, and activity-dependent trafficking. J. Neurosci. 29, 12845–12854 (2009).
Lukyanov, K. A. et al. Natural animal coloration can be determined by a nonfluorescent green fluorescent protein homolog. J. Biol. Chem. 275, 25879–25882 (2000).
Chudakov, D. M. et al. Kindling fluorescent proteins for precise in vivo photolabeling. Nature Biotech. 21, 191–194 (2003).
Ando, R., Mizuno, H. & Miyawaki, A. Regulated fast nucleocytoplasmic shuttling observed by reversible protein highlighting. Science 306, 1370–1373 (2004).
Ando, R., Flors, C., Mizuno, H., Hofkens, J. & Miyawaki, A. Highlighted generation of fluorescence signals using simultaneous two-color irradiation on Dronpa mutants. Biophys. J. 92, L97–L99 (2007).
Brakemann, T. et al. Molecular basis of the light-driven switching of the photochromic fluorescent protein Padron. J. Biol. Chem. 285, 14603–14609 (2010).
Henderson, J. N., Ai, H. W., Campbell, R. E. & Remington, S. J. Structural basis for reversible photobleaching of a green fluorescent protein homologue. Proc. Natl Acad. Sci. USA 104, 6672–6677 (2007).
Adam, V. et al. Structural characterization of IrisFP, an optical highlighter undergoing multiple photo-induced transformations. Proc. Natl Acad. Sci. USA 105, 18343–18348 (2008).
Stiel, A. C. et al. Generation of monomeric reversibly switchable red fluorescent proteins for far-field fluorescence nanoscopy. Biophys. J. 95, 2989–2997 (2008).
Subach, F. V. et al. Red fluorescent protein with reversibly photoswitchable absorbance for photochromic FRET. Chem. Biol. 17, 745–755 (2010).
Sawano, A., Takayama, S., Matsuda, M. & Miyawaki, A. Lateral propagation of EGF signaling after local stimulation is dependent on receptor density. Dev. Cell 3, 245–257 (2002).
Miyawaki, A. Visualization of the spatial and temporal dynamics of intracellular signaling. Dev. Cell 4, 295–305 (2003).
Sheetz, M. P., Turney, S., Qian, H. & Elson, E. L. Nanometre-level analysis demonstrates that lipid flow does not drive membrane glycoprotein movements. Nature 340, 284–288 (1989).
Schmidt, T., Schütz, G. J., Baumgartner, W., Gruber, H. J. & Schindler, H. Imaging of single molecule diffusion. Proc. Natl Acad. Sci. USA 93, 2926–2929 (1996).
Kusumi, A., Shirai, Y. M., Koyama-Honda, I., Suzuki, K. G. & Fujiwara, T. K. Hierarchical organization of the plasma membrane: investigations by single-molecule tracking vs. fluorescence correlation spectroscopy. FEBS Lett. 584, 1814–1823 (2010).
Tokunaga, M., Kitamura, K., Saito, K., Iwane, A. H. & Yanagida, T. Single molecule imaging of fluorophores and enzymatic reactions achieved by objective-type total internal reflection fluorescence microscopy. Biochem. Biophys. Res. Commun. 235, 47–53 (1997).
Mashanov, G. I., Tacon, D., Knight, A. E., Peckham, M. & Molloy, J. E. Visualizing single molecules inside living cells using total internal reflection fluorescence microscopy. Methods 29, 142–152 (2003).
Schneckenburger, H. Total internal reflection fluorescence microscopy: technical innovations and novel applications. Curr. Opin. Biotechnol. 16, 13–18 (2005).
Axelrod, D. Total internal reflection fluorescence microscopy. Methods Cell Biol. 89, 169–221 (2008).
Toomre, D. & Bewersdorf, J. A new wave of cellular imaging. Annu. Rev. Cell Dev. Biol. 26, 285–314 (2010).
Toomre, D., Steyer, J. A., Keller, P., Almers, W. & Simons, K. Fusion of constitutive membrane traffic with the cell surface observed by evanescent wave microscopy. J. Cell Biol. 149, 33–40 (2000).
Merrifield, C. J., Feldman, M. E., Wan, L. & Almers, W. Imaging actin and dynamin recruitment during invagination of single clathrin-coated pits. Nature Cell Biol. 4, 691–698 (2002).
Tokunaga, M., Imamoto, N. & Sakata-Spgawa, K. Highly inclined thin illumination enables clear single-molecule imaging in cells. Nature Methods 5, 159–161 (2008).
Shaner, N. C. et al. Improving the photostability of bright monomeric orange and red fluorescent proteins. Nature Methods 5, 545–551 (2008).
Merzlyak, E. M. et al. Bright monomeric red fluorescent protein with an extended fluorescence lifetime. Nature Methods 4, 555–557 (2007).
Chen, H., Farkas, E. R. & Watt, W. W. In vivo applications of fluorescence correlation spectroscopy. Methods Cell Biol. 89, 3–35 (2008).
Kohl, T. & Schwille, P. Fluorescence correlation spectroscopy with autofluorescent proteins. Adv. Biochem. Eng. Biotechnol. 95, 107–142 (2005).
Kim, S. A., Heinze, K. G. & Schwille, P. Fluorescence correlation spectroscopy in living cells. Nature Methods 4, 963–973 (2007).
Haustein, E. & Schwille, P. Fluorescence correlation spectroscopy: novel variations of an established technique. Annu. Rev. Biophys. Biomol. Struct. 36, 151–169 (2007).
Adams, M. C. et al. Signal analysis of total internal reflection fluorescent speckle microscopy (TIR-FSM) and wide-field epi-fluorescence FSM of the actin cytoskeleton and focal adhesions in living cells. J. Microsc. 216, 138–152 (2004).
Danuser, G. & Waterman-Storer, C. M. Quantitative fluorescent speckle microscopy of cytoskeleton dynamics. Annu. Rev. Biophys. Biomol. Struct. 35, 361–387 (2006).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S. T., Girirajan, T. P. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nature Methods 3, 793–795 (2006).
Manley, S. et al. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nature Methods 5, 155–157 (2008).
Cardarelli, F. & Gratton, E. In vivo imaging of single-molecule translocation through nuclear pore complexes by pair correlation functions. PLoS ONE 5, e10475 (2010).
Petersen, N. O., Höddelius, P. L., Wiseman, P. W., Seger, O. & Magnusson, K. E. Quantitation of membrane receptor distributions by image correlation spectroscopy: concept and application. Biophys. J. 65, 1135–1146 (1993).
Brown, C. M. et al. Raster image correlation spectroscopy (RICS) for measuring fast protein dynamics and concentrations with a commercial laser scanning confocal microscope. J. Microsc. 229, 78–91 (2008).
Capoulade, J., Wachsmuth, M., Hufnagel, L. & Knop, M. Quantitative fluorescence imaging of protein diffusion and interaction in living cells. Nature Biotech. 7 Aug 2011 (doi:10.1038/nbt.1928).
Kohl, T., Haustein, E. & Schwille, P. Determining protease activity in vivo by fluorescence cross-correlation analysis. Biophys. J. 89, 2770–2782 (2005).
Kogure, T. et al. A fluorescent variant of a protein from the stony coral Montipora facilitates dual-color single-laser fluorescence cross-correlation spectroscopy. Nature Biotech. 24, 577–581 (2006).
Piatkevich, K. D. et al. Monomeric red fluorescent proteins with a large Stokes shift. Proc. Natl Acd. Sci. USA 107, 5369–5374 (2010).
Miyawaki, A. & Tsien, R. Y. Monitoring protein conformations and interactions by fluorescence resonance energy transfer between mutants of green fluorescent protein. Methods Enzymol. 327, 472–500 (2000).
Jares-Erijman, E. A. & Jovin, T. M. FRET imaging. Nature Biotech. 21, 1387–1395 (2003).
Jares-Erijman, E. A. & Jovin, T. M. Imaging molecular interactions in living cells by FRET microscopy. Curr. Opin. Chem. Biol. 10, 409–416 (2006).
Piston, D. W. & Kremers, G. J. Fluorescent protein FRET: the good, the bad and ugly. Trends Biochem. Sci. 32, 407–414 (2007).
Frommer, W. B., Davidson, M. W. & Campbell, R. E. Genetically encoded biosensors based on engineered fluorescent proteins. Chem. Soc. Rev. 38, 2833–2841 (2009).
Miyawaki, A. Development of probes for cellular functions using fluorescent proteins and fluorescence resonance energy transfer. Annu. Rev. Biochem. 80, 357–373 (2011).
Gordon, G. W., Berry, G., Liang, X. H., Levine, B. & Herman, B. Quantitative fluorescence energy transfer measurements using fluorescence microscopy. Biophys. J. 74, 2702–2713 (1998).
Erickson, M. G., Liang, H., Mori, M. X. & Yue, D. T. FRET two-hybrid mapping reveals function and location of L-type Ca2+ channel CaM preassociation. Neuron 39, 97–107 (2003).
Berney, C. & Danuser, G. FRET or no FRET: a quantitative comparison. Biophys. J. 84, 3992–4010 (2003).
Yasuda, R. et al. Supersensitive Ras activation in dendrites and spines revealed by two-photon fluorescence lifetime imaging. Nature Neurosci. 9, 283–291 (2006).
Murakoshi, H., Wang, H. & Yasuda, R. Local, persistent activation of Rho GTPases during plasticity of single dendritic spines. Nature 472, 100–104 (2011).
Yasuda, R. & Murakoshi, H. The mechanisms underlying the spatial spreading of signaling activity. Curr. Opin. Neurobiol. 21, 313–321 (2011).
Kerppola, T. K. Visualization of molecular interactions using bimolecular fluorescence complementation analysis: characteristics of protein fragment complementation. Chem. Soc. Rev. 38, 2876–2886 (2009).
Shroff, H. et al. Dual-color superresolution imaging of genetically expressed probes within individual adhesion complexes. Proc. Natl Acad. Sci. USA 104, 20308–20313 (2007).
Grecco, H. E., Schmick, M. & Bastiaens, P. I. Signaling from the living plasma membrane. Cell 144, 897–909 (2011).
Sasaki, E. et al. Generation of transgenic non-human primates with germline transmission. Nature 459, 523–527 (2009).
Perentes, J. Y. et al. In vivo imaging of extracellular matrix remodeling by tumor-associated fibroblasts. Nature Methods. 6, 143–145 (2009).
Harvey, S. A. & Smith, J. C. Visualisation and quantification of morphogen gradient formation in the zebrafish. PLoS Biol. 7, e1000101 (2009).
Ries, J., Yu, S. R., Burkhardt, M., Brand, M. & Schwille, P. Modular scanning FCS quantifies receptor–ligand interactions in living multicellular organisms. Nature Methods 6, 643–645 (2009).
Burnette, D. T. et al. A role for actin arcs in the leading-edge advance of migrating cells. Nature Cell Biol. 13, 371–381 (2011).
van Thor, J. J., Gensch, T., Hellingwerf, K. J. & Johnson, L. N. Phototransformation of green fluorescent protein with UV and visible light leads to decarboxylation of glutamate 222. Nature Struct. Biol. 9, 37–41 (2002).
Mizuno, H. et al. Photo-induced peptide cleavage in the green-to-red conversion of a fluorescent protein. Mol. Cell 12, 1051–1058 (2003).
Periasamy, N. & Verkman, A. S. Analysis of fluorophore diffusion by continuous distributions of diffusion coefficients: application to photobleaching measurements of multicomponent and anomalous diffusion. Biophys. J. 75, 557–567 (1998).
Malchus, N. & Weiss, M. Anomalous diffusion reports on the interaction of misfolded proteins with the quality control machinery in the endoplasmic reticulum. Biophys. J. 99, 1321–1328 (2010).
Bancaud, A. et al. Molecular crowding affects diffusion and binding of nuclear proteins in heterochromatin and reveals the fractal organization of chromatin. EMBO J. 28, 3785–3798 (2009).
Marks, K. M. & Nolan, G. P. Chemical labeling strategies for cell biology. Nature Methods 3, 591–596 (2006).
Manley, S. Gillete, J. & Lippincott-Scwartz, J. Single-particle tracking photoactivated localization microscopy for mapping single-molecule dynamics. Methods Enzymol. 475, 109–120 (2010).
Acknowledgements
I thank S. Shimozono and R. Ando for the images in figures 2d and 2h; H. Sakurai, V. Lakshmanam and T. Fukano for general assistance; and S. Trowbridge, D. Mou, V. Verkhusha and S. Manley for valuable comments.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Supplementary information
41580_2011_BFnrm3199_MOESM6_ESM.pdf
Supplementary information S2 (figure) | Molecular mechanisms for the irreversible photoactivation of PA-GFP and photoconversion of Kaede. (PDF 317 kb)
Glossary
- Site-directed mutagenesis
-
An in vitro mutagenesis procedure that is often carried out using a polymerase chain reaction in which specific mutations are introduced into a DNA molecule.
- Hydrodynamic radius
-
The effective size of the molecule as detected by its diffusion.
- Fractal model
-
A model in which diffusing molecules and complexes encounter the same obstructions regardless of their size.
- Dendritic spines
-
Small membranous protrusions from dendritic shafts that usually receive excitatory input.
- Pyramidal neurons
-
The predominant type of neuron in the neocortex. They are named after their triangular cell bodies.
- Barrel cortex
-
The dark-staining regions of layer 4 of the somatosensory cortex, where somatosensory inputs from the contralateral side of the body come in from the thalamus.
- Axial dimension
-
The dimension along the optical axis of an objective lens.
- Highly inclined and laminated optical sheet
-
The illumination light for single-molecule imaging inside cells. The light is generated by positioning the incident beam to propagate near the objective edge.
- Multiplexing
-
A method that allows the simultaneous imaging of multiple events in a single cell.
- Diffraction limit
-
Although an ideal optical system would image an object point perfectly as a point, diffraction occurs owing to the wave-like nature of radiation, and the result is that the image of a point is a blur. Thus, the ability of an imaging system to resolve detail is ultimately limited by diffraction.
- Stochastic optical reconstruction microscopy
-
A high-resolution fluorescence microscopy method based on high-accuracy localization of photoswitchable fluorophores.
- Pair correlation function
-
A function that shows the probability of finding the centre of a particle that is located a given distance from the centre of another particle.
- Euchromatin
-
A form of chromatin that is lightly packed and often transcriptionally active during interphase.
- Heterochromatin
-
A condensed form of chromatin in which the degree of compaction is similar to that of mitotic chromosomes. It is usually found around the centromere.
- Stokes shifts
-
The energy difference between the emitted photon and the absorbed photon. The Stokes shift is usually expressed as the difference between positions of the band maxima of the absorption and emission spectra.
Rights and permissions
About this article
Cite this article
Miyawaki, A. Proteins on the move: insights gained from fluorescent protein technologies. Nat Rev Mol Cell Biol 12, 656–668 (2011). https://doi.org/10.1038/nrm3199
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3199
This article is cited by
-
An interplay between cellular growth and atypical fusion defines morphogenesis of a modular glial niche in Drosophila
Nature Communications (2022)
-
DNA tags used to image sugar-bearing proteins on cells
Nature (2018)
-
Generation of photonic entanglement in green fluorescent proteins
Nature Communications (2017)
-
Protein expression guided chemical profiling of living cells by the simultaneous observation of Raman scattering and anti-Stokes fluorescence emission
Scientific Reports (2017)
-
PKCα diffusion and translocation are independent of an intact cytoskeleton
Scientific Reports (2017)