Abstract
Understanding the molecular mechanisms of aging is crucial for enhancing healthy longevity. We conducted untargeted lipidomics across 13 biological samples from mice at various life stages (2, 12, 19 and 24 months) to explore the potential link between aging and lipid metabolism, considering sex (male or female) and microbiome (specific pathogen-free or germ-free) dependencies. By analyzing 2,704 molecules from 109 lipid subclasses, we characterized common and tissue-specific lipidome alterations associated with aging. For example, the levels of bis(monoacylglycero)phosphate containing polyunsaturated fatty acids increased in various organs during aging, whereas the levels of other phospholipids containing saturated and monounsaturated fatty acids decreased. In addition, we discovered age-dependent sulfonolipid accumulation, absent in germ-free mice, correlating with Alistipes abundance determined by 16S ribosomal RNA gene amplicon sequencing. In the male kidney, glycolipids such as galactosylceramides, galabiosylceramides (Gal2Cer), trihexosylceramides (Hex3Cer), and mono- and digalactosyldiacylglycerols were detected, with two lipid classes—Gal2Cer and Hex3Cer—being significantly enriched in aged mice. Integrated analysis of the kidney transcriptome revealed uridine diphosphate galactosyltransferase 8A (UGT8a), alkylglycerone phosphate synthase and fatty acyl-coenzyme A reductase 1 as potential enzymes responsible for the male-specific glycolipid biosynthesis in vivo, which would be relevant to sex dependency in kidney diseases. Inhibiting UGT8 reduced the levels of these glycolipids and the expression of inflammatory cytokines in the kidney. Our study provides a valuable resource for clarifying potential links between lipid metabolism and aging.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All raw MS data are available on the RIKEN DROP Met website (http://prime.psc.riken.jp/menta.cgi/prime/drop_index) under index number DM0044. The lipidomics results are recorded in Supplementary Data 1 and can be browsed from our RIKEN lipidomics database (http://prime.psc.riken.jp/menta.cgi/lipidomics/index). The raw RNA-sequencing data are available on the DNA Data Bank of Japan (DDBJ) web page under the identifier PRJDB14285. The raw 16S rDNA amplicon sequence data are available on the DDBJ webpage under the identifier PRJDB16347. The 16S rDNA and transcriptomics results are recorded in Supplementary Data 5 and 6, respectively. Source data are provided with this paper.
References
Harayama, T. & Riezman, H. Understanding the diversity of membrane lipid composition. Nat. Rev. Mol. Cell Biol. 19, 281–296 (2018).
Soehnlein, O. & Libby, P. Targeting inflammation in atherosclerosis—from experimental insights to the clinic. Nat. Rev. Drug Discov. 20, 589–610 (2021).
Snaebjornsson, M. T., Janaki-Raman, S. & Schulze, A. Greasing the wheels of the cancer machine: the role of lipid metabolism in cancer. Cell Metab. 31, 62–76 (2020).
Huby, T. & Gautier, E. L. Immune cell-mediated features of non-alcoholic steatohepatitis. Nat. Rev. Immunol. 22, 429–443 (2022).
Baek, J., He, C., Afshinnia, F., Michailidis, G. & Pennathur, S. Lipidomic approaches to dissect dysregulated lipid metabolism in kidney disease. Nat. Rev. Nephrol. 18, 38–55 (2022).
Johnson, A. A. & Stolzing, A. The role of lipid metabolism in aging, lifespan regulation, and age-related disease. Aging Cell 18, e13048 (2019).
Khosla, S., Farr, J. N., Tchkonia, T. & Kirkland, J. L. The role of cellular senescence in ageing and endocrine disease. Nat. Rev. Endocrinol. 16, 263–275 (2020).
Ferrucci, L. & Fabbri, E. Inflammageing: chronic inflammation in ageing, cardiovascular disease, and frailty. Nat. Rev. Cardiol. 15, 505–522 (2018).
Mutlu, A. S., Duffy, J. & Wang, M. C. Lipid metabolism and lipid signals in aging and longevity. Dev. Cell 56, 1394–1407 (2021).
Sacket, S. J., Chung, H. Y., Okajima, F. & Im, D. S. Increase in sphingolipid catabolic enzyme activity during aging. Acta Pharmacol. Sin. 30, 1454–1461 (2009).
Mielke, M. M. et al. Serum ceramides increase the risk of Alzheimer disease: the Women’s Health and Aging Study II. Neurology 79, 633–641 (2012).
Streeper, R. S. et al. Deficiency of the lipid synthesis enzyme, DGAT1, extends longevity in mice. Aging (Albany NY) 4, 13–27 (2012).
Su, L.-J. et al. Reactive oxygen species-induced lipid peroxidation in apoptosis, autophagy, and ferroptosis. Oxid. Med. Cell. Longev. 2019, 5080843 (2019).
Ponnappan, U., Holley, D. H. & Lipschitz, D. A. Effect of age on the fatty acid composition of phospholipids in human lymphocytes. Exp. Gerontol. 31, 125–133 (1996).
Rabini, R. A. et al. Reduced susceptibility to peroxidation of erythrocyte plasma membranes from centenarians. Exp. Gerontol. 37, 657–663 (2002).
Mitchell, T. W., Buffenstein, R. & Hulbert, A. J. Membrane phospholipid composition may contribute to exceptional longevity of the naked mole-rat (Heterocephalus glaber): a comparative study using shotgun lipidomics. Exp. Gerontol. 42, 1053–1062 (2007).
Nagpal, R. et al. Gut microbiome and aging: physiological and mechanistic insights. Nutr. Healthy Aging 4, 267–285 (2018).
Albouery, M. et al. Age-related changes in the gut microbiota modify brain lipid composition. Front. Cell. Infect. Microbiol. 9, 444 (2020).
Naoe, S., Tsugawa, H., Takahashi, M., Ikeda, K. & Arita, M. Characterization of lipid profiles after dietary intake of polyunsaturated fatty acids using integrated untargeted and targeted lipidomics. Metabolites 9, 241 (2019).
Weger, B. D. et al. The mouse microbiome is required for sex-specific diurnal rhythms of gene expression and metabolism. Cell Metab. 29, 362–382 (2019).
Yasuda, S. et al. Elucidation of gut microbiota-associated lipids using LC–MS/MS and 16S rRNA sequence analyses. iScience 23, 101841 (2020).
Ghorasaini, M. et al. Cross-laboratory standardization of preclinical lipidomics using differential mobility spectrometry and multiple reaction monitoring. Anal. Chem. 93, 16369–16378 (2021).
Tsugawa, H. et al. A lipidome atlas in MS-DIAL 4. Nat. Biotechnol. 38, 1159–1163 (2020).
Gonzalez-Covarrubias, V. Lipidomics in longevity and healthy aging. Biogerontology 14, 663–672 (2013).
Slade, E. et al. Age and sex are associated with the plasma lipidome: findings from the GOLDN study. Lipids Health Dis. 20, 30 (2021).
Beyene, H. B. et al. High-coverage plasma lipidomics reveals novel sex-specific lipidomic fingerprints of age and BMI: evidence from two large population cohort studies. PLoS Biol. 18, e3000870 (2020).
Eum, J. Y. et al. Aging-related lipidomic changes in mouse serum, kidney, and heart by nanoflow ultrahigh-performance liquid chromatography–tandem mass spectrometry. J. Chromatogr. A 1618, 460849 (2020).
Papsdorf, K. & Brunet, A. Linking lipid metabolism to chromatin regulation in aging. Trends Cell Biol. 29, 97–116 (2019).
Pollard, A. K., Ortori, C. A., Stöger, R., Barrett, D. A. & Chakrabarti, L. Mouse mitochondrial lipid composition is defined by age in brain and muscle. Aging (Albany NY) 9, 986–998 (2017).
Ding, J. et al. A metabolome atlas of the aging mouse brain. Nat. Commun. 12, 6021 (2021).
Tan, D. et al. A class of anti-inflammatory lipids decrease with aging in the central nervous system. Nat. Chem. Biol. 19, 187–197 (2023).
Ni, Z. X., Angelidou, G., Lange, M., Hoffmann, R. & Fedorova, M. LipidHunter identifies phospholipids by high-throughput processing of LC–MS and shotgun lipidomics datasets. Anal. Chem. 89, 8800–8807 (2017).
Koelmel, J. P. et al. LipidMatch: an automated workflow for rule-based lipid identification using untargeted high-resolution tandem mass spectrometry data. BMC Bioinformatics 18, 331 (2017).
Velagapudi, V. R. et al. The gut microbiota modulates host energy and lipid metabolism in mice. J. Lipid Res. 51, 1101–1112 (2010).
Grabner, G. F. et al. Metabolic regulation of the lysosomal cofactor bis(monoacylglycero)phosphate in mice. J. Lipid Res. 61, 995–1003 (2020).
Showalter, M. R. et al. The emerging and diverse roles of bis(monoacylglycero) phosphate lipids in cellular physiology and disease. Int. J. Mol. Sci. 21, 8067 (2020).
Jojima, K., Edagawa, M., Sawai, M., Ohno, Y. & Kihara, A. Biosynthesis of the anti-lipid-microdomain sphingoid base 4,14-sphingadiene by the ceramide desaturase FADS3. FASEB J. 34, 3318–3335 (2020).
Pergande, M. R. et al. Lipidomic analysis identifies age-disease-related changes and potential new biomarkers in brain-derived extracellular vesicles from metachromatic leukodystrophy mice. Lipids Health Dis. 21, 32 (2022).
Slomiany, B. L., Murty, V. L., Liau, Y. H. & Slomiany, A. Animal glycoglycerolipids. Prog. Lipid Res. 26, 29–51 (1987).
Walker, A. et al. Sulfonolipids as novel metabolite markers of Alistipes and Odoribacter affected by high-fat diets. Sci. Rep. 7, 11047 (2017).
Vital, M., Rud, T., Rath, S., Pieper, D. H. & Schluter, D. Diversity of bacteria exhibiting bile acid-inducible 7α-dehydroxylation genes in the human gut. Comput. Struct. Biotechnol. J. 17, 1016–1019 (2019).
Zhang, Q. et al. Genetic mapping of microbial and host traits reveals production of immunomodulatory lipids by Akkermansia muciniphila in the murine gut. Nat. Microbiol. 8, 424–440 (2023).
Brejchova, K. et al. Understanding FAHFAs: from structure to metabolic regulation. Prog. Lipid Res. 79, 101053 (2020).
Patel, R. et al. ATGL is a biosynthetic enzyme for fatty acid esters of hydroxy fatty acids. Nature 606, 968–975 (2022).
Wang, Y. et al. Sex differences in transcriptomic profiles in aged kidney cells of renin lineage. Aging (Albany NY) 10, 606–621 (2018).
Sembach, F. E. et al. Impact of sex on diabetic nephropathy and the renal transcriptome in UNx db/db C57BLKS mice. Physiol. Rep. 7, e14333 (2019).
Braun, F. et al. Altered lipid metabolism in the aging kidney identified by three layered omic analysis. Aging (Albany NY) 8, 441–457 (2016).
Reimand, J. et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat. Protoc. 14, 482–517 (2019).
Franceschi, C., Garagnani, P., Parini, P., Giuliani, C. & Santoro, A. Inflammaging: a new immune-metabolic viewpoint for age-related diseases. Nat. Rev. Endocrinol. 14, 576–590 (2018).
Carrero, J. J., Hecking, M., Chesnaye, N. C. & Jager, K. J. Sex and gender disparities in the epidemiology and outcomes of chronic kidney disease. Nat. Rev. Nephrol. 14, 151–164 (2018).
Zou, Z. N., Ohta, T., Miura, F. & Oki, S. ChIP-Atlas 2021 update: a data-mining suite for exploring epigenomic landscapes by fully integrating ChIP-seq, ATAC-seq and Bisulfite-seq data. Nucleic Acids Res. 50, W175–W182 (2022).
Martovetsky, G., Tee, J. B. & Nigam, S. K. Hepatocyte nuclear factors 4α and 1α regulate kidney developmental expression of drug-metabolizing enzymes and drug transporters. Mol. Pharmacol. 84, 808–823 (2013).
Chamouton, J. & Latruffe, N. PPARα/HNF4α interplay on diversified responsive elements. Relevance in the regulation of liver peroxisomal fatty acid catabolism. Curr. Drug Metab. 13, 1436–1453 (2012).
Harris, A. N., Castro, R. A., Lee, H.-W., Verlander, J. W. & Weiner, I. D. Role of the renal androgen receptor in sex differences in ammonia metabolism. Am. J. Physiol. Renal Physiol. 321, F629–F644 (2021).
O’Brown, Z. K., Van Nostrand, E. L., Higgins, J. P. & Kim, S. K. The inflammatory transcription factors NFκB, STAT1 and STAT3 drive age-associated transcriptional changes in the human kidney. PLoS Genet. 11, e1005734 (2015).
Liu, M. et al. Androgen–STAT3 activation may contribute to gender disparity in human simply renal cysts. Int. J. Clin. Exp. Pathol. 6, 686–694 (2013).
Iida, K. et al. A possible role of vitamin D receptors in regulating vitamin D activation in the kidney. Proc. Natl Acad. Sci. USA 92, 6112–6116 (1995).
Cozzolino, M. & Malindretos, P. The role of vitamin D receptor activation in chronic kidney disease. Hippokratia 14, 7–9 (2010).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Molenaar, M. R. et al. LION/web: a web-based ontology enrichment tool for lipidomic data analysis. Gigascience 8, giz061 (2019).
Muralidharan, S. et al. A reference map of sphingolipids in murine tissues. Cell Rep. 35, 109250 (2021).
van der Bijl, P., Strous, G. J., Lopes-Cardozo, M., Thomas-Oates, J. & van Meer, G. Synthesis of non-hydroxy-galactosylceramides and galactosyldiglycerides by hydroxy-ceramide galactosyltransferase. Biochem. J. 317, 589–597 (1996).
Hayashi, T., Hayashi, E., Fujimoto, M., Sprong, H. & Su, T.-P. The lifetime of UDP-galactose:ceramide galactosyltransferase is controlled by a distinct endoplasmic reticulum-associated degradation (ERAD) regulated by sigma-1 receptor chaperones. J. Biol. Chem. 287, 43156–43169 (2012).
Eckhardt, M. Fatty acid 2-hydroxylase and 2-hydroxylated sphingolipids: metabolism and function in health and diseases. Int. J. Mol. Sci. 24, 4908 (2023).
Lu, C.-L. et al. Indoxyl-sulfate-induced redox imbalance in chronic kidney disease. Antioxidants (Basel) 10, 936 (2021).
Strott, C. A. & Higashi, Y. Cholesterol sulfate in human physiology: what’s it all about? J. Lipid Res. 44, 1268–1278 (2003).
Stofan, M. & Guo, G. L. Bile acids and FXR: novel targets for liver diseases. Front. Med. (Lausanne) 7, 544 (2020).
Rouillard, A. D. et al. The harmonizome: a collection of processed datasets gathered to serve and mine knowledge about genes and proteins. Database (Oxford) 2016, baw100 (2016).
Dayama, G., Priya, S., Niccum, D. E., Khoruts, A. & Blekhman, R. Interactions between the gut microbiome and host gene regulation in cystic fibrosis. Genome Med. 12, 12 (2020).
Ohsaka, F. et al. Gut commensals suppress interleukin-2 production through microRNA-200/BCL11B and microRNA-200/ETS-1 axes in lamina propria leukocytes of murine large intestine. Biochem. Biophys. Res. Commun. 534, 808–814 (2021).
Kolter, T. & Sandhoff, K. Lysosomal degradation of membrane lipids. FEBS Lett. 584, 1700–1712 (2010).
Johmura, Y. et al. Senolysis by glutaminolysis inhibition ameliorates various age-associated disorders. Science 371, 265–270 (2021).
Babenko, N. A., Garkavenko, V. V., Storozhenko, G. V. & Timofiychuk, O. A. Role of acid sphingomyelinase in the age-dependent dysregulation of sphingolipids turnover in the tissues of rats. Gen. Physiol. Biophys. 35, 195–205 (2016).
Medoh, U. N. et al. The Batten disease gene product CLN5 is the lysosomal bis(monoacylglycero)phosphate synthase. Science 381, 1182–1189 (2023).
Parker, B. J., Wearsch, P. A., Veloo, A. C. M. & Rodriguez-Palacios, A. The genus Alistipes: gut bacteria with emerging implications to inflammation, cancer, and mental health. Front. Immunol. 11, 906 (2020).
Almsherqi, Z. A. Potential role of plasmalogens in the modulation of biomembrane morphology. Front. Cell Dev. Biol. 9, 673917 (2021).
Tadano-Aritomi, K. et al. Kidney lipids in galactosylceramide synthase-deficient mice: absence of galactosylsulfatide and compensatory increase in more polar sulfoglycolipids. J. Lipid Res. 41, 1237–1243 (2000).
Honke, K. et al. Paranodal junction formation and spermatogenesis require sulfoglycolipids. Proc. Natl Acad. Sci. USA 99, 4227–4232 (2002).
Stormo, G. D. Modeling the specificity of protein–DNA interactions. Quant. Biol. 1, 115–130 (2013).
Thelen, A. M. & Zoncu, R. Emerging roles for the lysosome in lipid metabolism. Trends Cell Biol. 27, 833–850 (2017).
Yu, Z. et al. Human serum metabolic profiles are age dependent. Aging Cell 11, 960–967 (2012).
Montoliu, I. et al. Serum profiling of healthy aging identifies phospho- and sphingolipid species as markers of human longevity. Aging (Albany NY) 6, 9–25 (2014).
Jarrell, Z. R. et al. Plasma acylcarnitine levels increase with healthy aging. Aging (Albany NY) 12, 13555–13570 (2020).
Folz, J. et al. Human metabolome variation along the upper intestinal tract. Nat. Metab. 5, 777–788 (2023).
Akiyama, H. et al. Galabiosylceramide is present in human cerebrospinal fluid. Biochem. Biophys. Res. Commun. 536, 73–79 (2021).
Nowak, A., Beuschlein, F., Sivasubramaniam, V., Kasper, D. & Warnock, D. G. Lyso-Gb3 associates with adverse long-term outcome in patients with Fabry disease. J. Med. Genet. 59, 287–293 (2022).
Fan, Y. & Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 19, 55–71 (2021).
Tsugawa, H., Rai, A., Saito, K. & Nakabayashi, R. Metabolomics and complementary techniques to investigate the plant phytochemical cosmos. Nat. Prod. Rep. 38, 1729–1759 (2021).
McDonald, J. G. et al. Introducing the Lipidomics Minimal Reporting Checklist. Nat. Metab. 4, 1086–1088 (2022).
Okahashi, N., Ueda, M., Yasuda, S., Tsugawa, H. & Arita, M. Global profiling of gut microbiota-associated lipid metabolites in antibiotic-treated mice by LC–MS/MS-based analyses. STAR Protoc. 2, 100492 (2021).
da Costa, E., Amaro, H. M., Melo, T., Guedes, A. C. & Domingues, M. R. Screening for polar lipids, antioxidant, and anti-inflammatory activities of Gloeothece sp. lipid extracts pursuing new phytochemicals from cyanobacteria. J. Appl. Phycol. 32, 3015–3030 (2020).
Moore, E. K. et al. Novel mono-, di-, and trimethylornithine membrane lipids in northern wetland planctomycetes. Appl. Environ. Microbiol. 79, 6874–6884 (2013).
Guo, L., Amarnath, V. & Davies, S. S. A liquid chromatography–tandem mass spectrometry method for measurement of N-modified phosphatidylethanolamines. Anal. Biochem. 405, 236–245 (2010).
Munger, L. H., Boulos, S. & Nystrom, L. UPLC–MS/MS based identification of dietary steryl glucosides by investigation of corresponding free sterols. Front. Chem. 6, 342 (2018).
Kessner, D., Chambers, M., Burke, R., Agus, D. & Mallick, P. ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics 24, 2534–2536 (2008).
Kato, T. et al. Multiple omics uncovers host–gut microbial mutualism during prebiotic fructooligosaccharide supplementation. DNA Res. 21, 469–480 (2014).
Maki, K. A., Wolff, B., Varuzza, L., Green, S. J. & Barb, J. J. Multi-amplicon microbiome data analysis pipelines for mixed orientation sequences using QIIME2: assessing reference database, variable region and pre-processing bias in classification of mock bacterial community samples. PLoS ONE 18, e0280293 (2023).
Acknowledgements
This work was supported by the Japan Science and Technology Agency (JST) ERATO ‘Arita Lipidome Atlas Project’ (JPMJER2101 to H.T. and M.A.), RIKEN Aging Project (M.A. and A.M.), Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Scientific Research on Innovative Areas ‘Biology of LipoQuality’ (15H05897 and 15H05898 to M.A.), JSPS KAKENHI (21K18216 to H.T., 22K11718 to T.I., 20H00495 to M.A.), National Cancer Center Research and Development Fund (2020-A-9 to H.T.), AMED Moonshot Research and Development Program (JP22zf0127007 to M.A.), AMED NEDDTrim (22ae0121036h0002 to M.A.), AMED Japan Program for Infectious Diseases Research and Infrastructure (21wm0325036h0001 to H.T. and M.A.), AMED Brain/MINDS (JP15dm0207001 to H.T.), JST National Bioscience Database Center (JPMJND2305 to H.T.) and Takeda Science Foundation (M.A.). The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
H.T., T.I., A.M. and M.A. designed the study. H.T. and M.T. developed an MS-DIAL annotation system. T.I. prepared the biological samples, and T.I. and A.H. performed the LC–MS experiments. K.O. and S.I. performed the kidney transcriptome analysis. N.S.-T. and H.O. performed the microbiome analysis. Y.Y. developed the RIKEN lipidomics database. H.T. and M.A. wrote the paper. All authors have thoroughly discussed this project and helped improve the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Naama Geva-Zatorsky, Marcelo Mori, Martin Denzel and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Summary of experimental design and lipid profiling in this study.
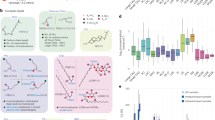
(a) Four types of mice were prepared: male and SPF (specific pathogen-free), male and GF (germ free), female and SPF, and female and GF. Total 13 biospecimens were harvested at 2 months, 12 months, 19 months, and 24 months. (b) The lipid extraction was performed for the optimal volume of biological samples. The untargeted lipidomics data was obtained using our experimental condition. The mass spectrometry data was analyzed by the MS-DIAL 4.20 algorithm with the updated lipid libraries. Panel a icons from iStock.
Extended Data Fig. 2 Lipid profiling result of AIN-93M chow.
The x- and y-axis show the lipid subclass name and the log10 transformed value of normalized peak height. Each dot denotes each lipid molecule. The elements of the box plot are defined as follows: center line, median; box limits, upper and lower quartiles; and whiskers, 1.5x interquartile range. The number of molecules included in each lipid subclass is described in Supplementary Data 1.
Extended Data Fig. 3 Relationship between fecal microbiome and lipidome.
(a) The scatter plot of principal coordinates analysis (PCoA) using the unweighted UniFrac distance. The sky- and dark-blue colors denote 2- and 24-month-old mice, respectively. The circle and diamond shapes denote male and female, respectively. (b) The correlations between Alistipes and sulfonolipid (SL). The 95% confidence interval is also described by the gray color. The correlation coefficient test was performed to calculate the p-value (two-sided). (c) Bacteria relative abundances between aged- and young mice. The left- and right panels show the results from 16S rRNA and qPCR data, respectively. Student t-test was used to calculate the p-value (two-sided). N = 6 biologically independent samples where the results of male and female mice are recognized in the same group to calculate the p-value.
Extended Data Fig. 4 The significantly changed lipids and their related metabolic pathways in kidney tissue.
(a) The metabolic pathway of ether-linked (alkylacyl) glycerolipids and glycerophospholipids. The level of ether PE is calculated from the molecules annotated as plasmalogen type in the positive ion mode. (b) The significant genes related to the alkylacyl glycerolipids, and the other UGT genes, which were expected to be related to glycosyl lipid metabolism. The definition of symbol and color in the dot plot is the same as in Fig. 2. The p-value was calculated by Dunnett’s test (two-sided). *P < 0.05, **P < 0.01, and ***P < 0.001 against 2 months. N = 6 biologically independent samples where the results of SPF and GF mice are recognized in the same group.
Extended Data Fig. 5 The correlation of genes with the lipid metabolites associated with gut microbiota.
The definitions of statistical significance with months (aging), SPF/GF, and gene-lipid correlation are the same as those in Fig. 6. For phosphatidylcholine (PC), the molecules containing 17:0 and 17:1 were investigated even if several molecules were not included in the lipid clusters. The yellow color of SPF/GF means the significance. Several bile acids were identified by authentic standards. Dot plots indicate the significantly changed genes only in SPF mice with aging. The p-value was calculated by Dunnett’s test (two-sided). *P < 0.05, **P < 0.01 against 2 months. N = 6 biologically independent samples where the results of male and female mice are recognized in the same group.
Supplementary information
Supplementary Information
Legends of Supplementary Tables, legends of Supplementary Data, and Supplementary Note showing the Lipidomics Minimal Reporting Checklist.
Supplementary Tables
Supplementary Tables 1–8. The legends are also listed in the Supplementary Information.
Supplementary Data 1
Lipidome results of biological samples.
Supplementary Data 2
Annotation results from LipidHunter and LipidMatch.
Supplementary Data 3
MS/MS spectral annotation for odd-chain fatty acid-containing lipids and oxidized phospholipids.
Supplementary Data 4
VIP values generated in OPLS-R.
Supplementary Data 5
Fecal microbiome results.
Supplementary Data 6
Kidney transcriptome results.
Source data
Source Data Fig. 3
Data and source code to generate Fig. 3.
Source Data Fig. 4
Required information to create Fig. 4.
Source Data Fig. 5
Required information to create Fig. 5.
Source Data Fig. 6
Required information to create Fig. 6.
Source Data Fig. 8
Required information to create Fig. 8.
Source Data Extended Data Fig. 1
Required information to create Extended Data Fig. 1.
Source Data Extended Data Fig. 3
Required information to create Extended Data Fig. 3.
Source Data Extended Data Fig. 5
Required information to create Extended Data Fig. 5.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tsugawa, H., Ishihara, T., Ogasa, K. et al. A lipidome landscape of aging in mice. Nat Aging (2024). https://doi.org/10.1038/s43587-024-00610-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43587-024-00610-6