Abstract
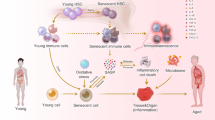
Sterile inflammation, also known as ‘inflammaging’, is a hallmark of tissue aging. Cellular senescence contributes to tissue aging, in part, through the secretion of proinflammatory factors collectively known as the senescence-associated secretory phenotype (SASP). The genetic variability of thioredoxin reductase 1 (TXNRD1) is associated with aging and age-associated phenotypes such as late-life survival, activity of daily living and physical performance in old age. TXNRD1’s role in regulating tissue aging has been attributed to its enzymatic role in cellular redox regulation. Here, we show that TXNRD1 drives the SASP and inflammaging through the cyclic GMP–AMP synthase (cGAS)–stimulator of interferon genes (STING) innate immune response pathway independently of its enzymatic activity. TXNRD1 localizes to cytoplasmic chromatin fragments and interacts with cGAS in a senescence-status-dependent manner, which is necessary for the SASP. TXNRD1 enhances the enzymatic activity of cGAS. TXNRD1 is required for both the tumor-promoting and immune surveillance functions of senescent cells, which are mediated by the SASP in vivo in mouse models. Treatment of aged mice with a TXNRD1 inhibitor that disrupts its interaction with cGAS, but not with an inhibitor of its enzymatic activity alone, downregulated markers of inflammaging in several tissues. In summary, our results show that TXNRD1 promotes the SASP through the innate immune response, with implications for inflammaging. This suggests that the TXNRD1–cGAS interaction is a relevant target for selectively suppressing inflammaging.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RNA-seq datasets of IMR90 cells have been deposited in the Gene Expression Omnibus (GEO) under accession number GSE202664. RNA-seq datasets of mouse tissues have been deposited in the GEO under accession number GSE204841. MS proteomics data have been deposited in the MassIVE (http://massive.ucsd.edu) and ProteomeXchange (http://www.proteomexchange.org) data repositories with accession numbers MSV000089650 and PXD034505, respectively. All other data supporting the findings of this study are available within this article and its supplementary materials. Source data are provided with this paper.
Code availability
The software and algorithms for data analyses used in this study are all well-established from previous work and are referenced throughout the article.
References
Munoz-Espin, D. & Serrano, M. Cellular senescence: from physiology to pathology. Nat. Rev. Mol. Cell Biol. 15, 482–496 (2014).
Hao, X., Wang, C. & Zhang, R. Chromatin basis of the senescence-associated secretory phenotype. Trends Cell Biol. https://doi.org/10.1016/j.tcb.2021.12.003 (2022).
Dou, Z. et al. Cytoplasmic chromatin triggers inflammation in senescence and cancer. Nature 550, 402–406 (2017).
Glück, S. et al. Innate immune sensing of cytosolic chromatin fragments through cGAS promotes senescence. Nat. Cell Biol. 19, 1061–1070 (2017).
Yang, H., Wang, H., Ren, J., Chen, Q. & Chen, Z. J. cGAS is essential for cellular senescence. Proc. Natl Acad. Sci. USA 114, E4612–E4620 (2017).
Takahashi, A. et al. Downregulation of cytoplasmic DNases is implicated in cytoplasmic DNA accumulation and SASP in senescent cells. Nat. Commun. 9, 1249 (2018).
Franceschi, C. & Campisi, J. Chronic inflammation (inflammaging) and its potential contribution to age-associated diseases. J. Gerontol. A 69, S4–S9 (2014).
López-Otin, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Arnér, E. S. J. Targeting the selenoprotein thioredoxin reductase 1 for anticancer therapy. Adv. Cancer Res. 136, 139–151 (2017).
Dato, S., De Rango, F., Crocco, P., Passarino, G. & Rose, G. Antioxidants and quality of aging: further evidences for a major role of TXNRD1 gene variability on physical performance at old age. Oxid. Med. Cell. Longev. 2015, 926067 (2015).
Dato, S. et al. Contribution of genetic polymorphisms on functional status at very old age: a gene-based analysis of 38 genes (311 SNPs) in the oxidative stress pathway. Exp. Gerontol. 52, 23–29 (2014).
Soerensen, M. et al. Human longevity and variation in GH/IGF-1/insulin signaling, DNA damage signaling and repair and pro/antioxidant pathway genes: cross sectional and longitudinal studies. Exp. Gerontol. 47, 379–387 (2012).
Usategui-Martin, R. et al. Association between genetic variants in oxidative stress-related genes and osteoporotic bone fracture. The Hortega follow-up study. Gene 809, 146036 (2022).
El Assar, M. et al. Frailty is associated with lower expression of genes involved in cellular response to stress: results from the Toledo Study for Healthy Aging. J. Am. Med. Dir. Assoc. 18, 734 (2017).
Rodriguez-Mañas, L. & Fried, L. P. Frailty in the clinical scenario. Lancet 385, e7–e9 (2015).
Finkel, T. & Holbrook, N. J. Oxidants, oxidative stress and the biology of ageing. Nature 408, 239–247 (2000).
Stafford, W. C. et al. Irreversible inhibition of cytosolic thioredoxin reductase 1 as a mechanistic basis for anticancer therapy. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aaf7444 (2018).
Li, X.-D. et al. Pivotal roles of cGAS–cGAMP signaling in antiviral defense and immune adjuvant effects. Science 341, 1390–1394 (2013).
Liu, Z.-S. et al. G3BP1 promotes DNA binding and activation of cGAS. Nat. Immunol. 20, 18–28 (2019).
Gromer, S., Arscott, L. D., Williams, C. H. Jr, Schirmer, R. H. & Becker, K. Human placenta thioredoxin reductase. Isolation of the selenoenzyme, steady state kinetics, and inhibition by therapeutic gold compounds. J. Biol. Chem. 273, 20096–20101 (1998).
Lama, L. et al. Development of human cGAS-specific small-molecule inhibitors for repression of dsDNA-triggered interferon expression. Nat. Commun. 10, 2261 (2019).
Cheng, Q. et al. The selenium-independent inherent pro-oxidant NADPH oxidase activity of mammalian thioredoxin reductase and its selenium-dependent direct peroxidase activities. J. Biol. Chem. 285, 21708–21723 (2010).
Michalski, S. et al. Structural basis for sequestration and autoinhibition of cGAS by chromatin. Nature 587, 678–682 (2020).
Zhao, B. et al. The molecular basis of tight nuclear tethering and inactivation of cGAS. Nature 587, 673–677 (2020).
Pathare, G. R. et al. Structural mechanism of cGAS inhibition by the nucleosome. Nature 587, 668–672 (2020).
Herranz, N. & Gil, J. Mechanisms and functions of cellular senescence. J. Clin. Invest. 128, 1238–1246 (2018).
Rodier, F. & Campisi, J. Four faces of cellular senescence. J. Cell Biol. 192, 547–556 (2011).
Coppé, J.-P. et al. Senescence-associated secretory phenotypes reveal cell-nonautonomous functions of oncogenic RAS and the p53 tumor suppressor. PLoS Biol. 6, e301 (2008).
Kang, T.-W. et al. Senescence surveillance of pre-malignant hepatocytes limits liver cancer development. Nature 479, 547–551 (2011).
Gulen, M. F. et al. cGAS–STING drives ageing-related inflammation and neurodegeneration. Nature 620, 374–380 (2023).
Kerur, N. et al. cGAS drives noncanonical-inflammasome activation in age-related macular degeneration. Nat. Med. 24, 50–61 (2018).
Hou, Y. et al. NAD+ supplementation reduces neuroinflammation and cell senescence in a transgenic mouse model of Alzheimer’s disease via cGAS–STING. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2011226118 (2021).
Zhong, Z. et al. NF-κB restricts inflammasome activation via elimination of damaged mitochondria. Cell 164, 896–910 (2016).
Swanson, K. V., Deng, M. & Ting, J. P.-Y. The NLRP3 inflammasome: molecular activation and regulation to therapeutics. Nat. Rev. Immunol. 19, 477–489 (2019).
Schroder, K. & Tschopp, J. The inflammasomes. Cell 140, 821–832 (2010).
Rosenbaum, S. R., Wilski, N. A. & Aplin, A. E. Fueling the fire: inflammatory forms of cell death and implications for cancer immunotherapy. Cancer Discov. 11, 266–281 (2021).
Navarro-Pando, J. M. et al. Inhibition of the NLRP3 inflammasome prevents ovarian aging. Sci. Adv. https://doi.org/10.1126/sciadv.abc7409 (2021).
McIlwain, D. R., Berger, T. & Mak, T. W. Caspase functions in cell death and disease. Cold Spring Harb. Perspect. Biol. 5, a008656 (2013).
Michaud, M. et al. Proinflammatory cytokines, aging, and age-related diseases. J. Am. Med. Dir. Assoc. 14, 877–882 (2013).
Cassidy, L. D. et al. Temporal inhibition of autophagy reveals segmental reversal of ageing with increased cancer risk. Nat. Commun. 11, 307 (2020).
Mitsui, A. et al. Overexpression of human thioredoxin in transgenic mice controls oxidative stress and life span. Antioxid. Redox Signal. 4, 693–696 (2002).
Nelson, G., Kucheryavenko, O., Wordsworth, J. & von Zglinicki, T. The senescent bystander effect is caused by ROS-activated NF-κB signalling. Mech. Ageing Dev. 170, 30–36 (2018).
Zhao, B. et al. Topoisomerase 1 cleavage complex enables pattern recognition and inflammation during senescence. Nat. Commun. 11, 908 (2020).
Nacarelli, T. et al. NAD+ metabolism governs the proinflammatory senescence-associated secretome. Nat. Cell Biol. 21, 397–407 (2019).
Liu, P. et al. m6A-independent genome-wide METTL3 and METTL14 redistribution drives the senescence-associated secretory phenotype. Nat. Cell Biol. 23, 355–365 (2021).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Garcia, M. F. et al. Structural features of transcription factors associating with nucleosome binding. Mol. Cell 75, 921–932 (2019).
Hao, X. et al. ADAR1 downregulation by autophagy drives senescence independently of RNA editing by enhancing p16INK4a levels. Nat. Cell Biol. 24, 1202–1210 (2022).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Acknowledgements
We thank F. Keeney at The Wistar Institute Imaging Facility for qualifying NLRP3 staining in mouse tissues. This work was supported by the US National Institutes of Health (grants R01CA160331 and R01CA276569 to R.Z.; P01AG031862 to P.D.A., S.L.B, R.M., D.S. and R.Z.; R50CA221838 to H.-Y.T.; and R50CA211199 to A.V.K.), US Department of Defense (OC220011 to X.H.) and Cancer Prevention and Research Institute of Texas (CPRIT Scholar in Cancer Research RR230005 to R.Z.). Support of core facilities was provided by the Cancer Center Support Grant (CCSG) CA010815 to The Wistar Institute and P30CA016672 to the University of Texas MD Anderson Cancer Center. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
X.H., B.Z. and R.Z. designed the experiments. X.H., B.Z., M.T., L.L., E.L.M., X.X., C.F., H.P. and H.-Y.T. performed the experiments and analyzed the data. A.V.K. performed the bioinformatics analysis. A.H., D.S., R.M., K.S.Z. and S.L.B. contributed key experimental materials and participated in the study design. X.H., H.-Y.T., A.V.K. and R.Z. wrote the paper. D.W.S., P.D.A., S.L.B. and R.Z. supervised the study. R.Z. conceived the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Masashi Narita for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purification of CCFs from oncogene-induced senescent cells.
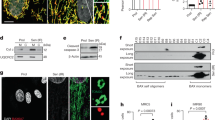
a-c, IMR90 cells were induced into senesce by oncogenic H-RASG12V and were subjected to SA-β-gal staining (a). SA-β-gal positive cells were quantified in the indicated groups (b). Expression of the indicated proteins in the indicated control senescent cells was analyzed by immunoblot (c). Scale bar = 100 μm. d, Schematics of the protocol used for purification of CCFs. e, Intensity heatmap of the thioredoxin-related proteins enriched in CCFs. Sorted by number of the detected peptides with two independent biological repeats. Data represent mean ± s.e.m. n = 3 biologically independent experiments unless otherwise stated. P-values were calculated using a two-tailed t test.
Extended Data Fig. 2 TXNRD1 localizes into CCFs during senescence.
a, Representative images of immunostaining for γH2AX and TXNRD1 in control and oncogenic RAS-induced senescent IMR90 cells. The arrow indicates an example of γH2AX and TXNRD1 positive CCFs in RAS-induced senescent cells. Scale bar = 10 μm. b, Representative images of immunostaining for γH2AX, cGAS and TXNRD1 in young (PD26) and replicative senescent (PD72) IMR90 cells. The arrow indicates an example of γH2AX, cGAS and TXNRD1 positive CCFs in replicative senescent cells. Scale bar = 10 μm. c, Quantification of (a) and (b). d,e, Representative images of immunostaining for γH2AX, cGAS and TXNRD1 in control and oncogenic RAS-induced senescent BJ human fibroblasts. The arrows indicate examples of γH2AX, cGAS and TXNRD1 positive CCFs in RAS-induced senescent cells (d). γH2AX-positive CCFs that are positive for TXNRD1 in senescent BJ cells were quantified (e). Scale bar = 10 μm. Data represent mean ± s.e.m. n = 3 biologically independent experiments.
Extended Data Fig. 3 Inhibition of TXNRD1 doesn’t affect senescence-associated cell growth arrest.
a, Immunoblot of the indicated proteins in oncogenic RAS-induced senescent IMR90 cells with or without TXNRD1 knockdown. b, γH2AX positive CCFs were quantified in oncogenic RAS-induced senescent IMR90 cells with or without TXNRD1 knockdown. n = 4 biologically independent experiments. c, Immunoblot of the indicated protein in control proliferating and oncogenic RAS-induced senescent IMR90 cells with or without TXNRD1 knockdown or treatment with a pharmacological TXNRD1 inhibitor Tri-1 (5 μM). d, e, Control and oncogenic RAS-induced senescent IMR90 cells with or without TXNRD1 knockdown or treatment with a pharmacological TXNRD1 inhibitor Tri-1 (5 μM) were subjected to SA-β-gal staining or colony formation assays (d). SA-β-gal positive cells were quantified in the indicated groups (e). Scale bar = 100 μm. n = 3 biologically independent experiments. Data represent mean ± s.e.m.
Extended Data Fig. 4 TXNRD1 is required for cGAS-STING activation during therapy-induced senescence.
a, Immunoblot of the indicated proteins in cisplatin-induced senescent PEO1 cells with or without TXNRD1 knockdown. b-d, Representative images of immunostaining for cGAS and γH2AX in cisplatin-induced senescent PEO1 cells with or without TXNRD1 knockdown or treatment with Tri-1 or vehicle control (b). White arrows indicate examples of cGAS and γH2AX positive CCFs in control cells, while the yellow arrows indicate examples of cGAS negative, γH2AX positive CCFs in TXNRD1 knockdown or Tri-1 treated cells. γH2AX-positive CCFs that are positive for cGAS from senescent PEO1 cells with or without TXNRD1 knockdown were quantified (c). γH2AX-positive CCFs that are positive for cGAS from senescent PEO1 cells with or without TXNRD1 inhibitor Tri-1 treatment were quantified (d). Scale bar = 10 μm. e, f, Immunoblot of the indicated protein in control and cisplatin-induced senescent PEO1 cells with or without TXNRD1 knockdown or treatment with a pharmacological TXNRD1 inhibitor Tri-1 (5 μM) (e). In addition, cellular 2’ 3’-cGAMP levels were measured in the indicated cells (f). Data represent mean ± s.e.m. n = 3 biologically independent experiments. P-values were calculated using a two-tailed t test.
Extended Data Fig. 5 TXNRD1 inhibition during senescence induction suppresses the SASP.
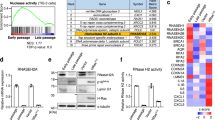
a, Schematic of experimental design for determining the effects of TXNRD1 inhibition during induction of senescence in IMR90 cells. b, Heatmap of the SASP genes that were significantly suppressed by both TXNRD1 knockdown and Tri-1 treatment based on RNA-seq analysis. The relative expression levels per replicate and average fold change differences are shown (n = 3 biologically independent experiments). c, d, Expression of the indicated SASP genes in control and oncogenic RAS-induced senescent IMR90 cells with or without TXNRD1 knockdown (c) or Tri-1 treatment (d) was determined by RT-qPCR. n = 4 biologically independent experiments. e, f, Expression of the indicated proteins in oncogenic RAS-induced senescent IMR90 cells with or without TXNRD1 knockout was determined by immunoblot (e), and expression of the indicated SASP genes was determined by RT-qPCR (f). Data represent mean ± s.e.m. n = 3 biologically independent experiments unless otherwise stated. P-values were calculated using a two-tailed t test.
Extended Data Fig. 6 TXN knockdown does not affect cGAS localization or activity.
a–d, Expression of the indicated proteins in IMR90 cells induced to undergo senescence by oncogenic RAS expressing shControl or shTXN (a). The indicated cells were stained for γH2AX and cGAS. DAPI counter staining was used to visualized nuclei. Arrows point to examples of cGAS positive CCFs (b), which was quantified (c). Further, 2’3’-cGAMP levels in the indicated cells were quantified (d). Scale bar = 10 μm. Data represent mean ± s.e.m. n = 3 biologically independent experiments.
Extended Data Fig. 7 TXNRD1 is required for SASP function in vivo.
a, TOV21G ovarian cancer cell growth in conditioned medium collected from control and senescent IMR90 cells with or without TXNRD1 knockdown or Tri-1 treatment. After 7 days of incubation, the cell numbers were determined and normalized to the numbers of the cells cultured in conditioned media collected from control proliferating IMR90 cells. Data represent mean ± s.e.m. n = 3 biologically independent experiments unless otherwise stated. P-values were calculated using a two-tailed t test. b, c, TOV21G and oncogene-induced senescent IMR90 cells with or without TXNRD1 inhibition were subcutaneously co-injected into the right dorsal flank of 6–8-week-old NSG female mice (n = 5 biologically independent mice per group). Shown are images of tumors dissected in the indicated groups at the end of experiments (b). Tumor growth in the indicated treatment groups was measured at the indicated time points (c). Data represent mean ± s.e.m. d, Validation of Txnrd1 knockdown by immunostaining in mouse NIH 3T3 cells. Arrows point to dsRed-expressing shRen control, shTxnrd1 #1 and shTxnrd1 #2. Scale bars = 10 μm. The experiment was repeated twice with similar results.
Extended Data Fig. 8 Tri-1 suppresses NLRP3 positivity in replicative senescent cells.
a, b, Knockdown of Caspase 1 (a) and GSDMD (b) in senescent IMR90 cells was validated by RT-qPCR. n = 4 biologically independent experiments. c, Expression of IL1β in ER-RAS induced (by 4-OHT) senescent IMR90 cells with or without the knockdown of GSDMD or Caspase 1 was determined by RT-qPCR analysis. n = 3 biologically independent experiments. d, e, Representative images of immunostaining for NLRP3 in ER-RAS induced (by 4-OHT) senescent IMR90 cells with or without the indicated treatments (d). NLRP3 positive cells were quantified (e). n = 3 biologically independent experiments. Scale bars = 10 μm. f, g, Representative images of immunostaining for ASC to visualize inflammasome formation in the indicated control and ER-RAS induced (by 4-OHT) senescent IMR90 cells (f). ASC speck positive cells were quantified (g). n = 3 biologically independent experiments. Nigericin, a known inducer of inflammasome formation, was used as a positive control (10 μM for 4 hours). Scale bars = 10 μm. h, Expression of Txnrd1 in young (4 months) and aged mice (22 months) with or without Tri-1 treatment was determined by RT-qPCR analysis. n = 4 biologically independent mice per group. i, j, Immunoblot of the indicated proteins in the ovary tissues harvested from young (4 months) and aged mice (22 months) (i). The intensity of the indicated proteins was quantified by NIH ImageJ software and normalized against a loading control β-actin expression (j). n = 5 biologically independent mice per group. Data represent mean ± s.e.m. P-values were calculated using a two-tailed t test.
Extended Data Fig. 9 Tri-1 and auranofin do not affect p16 and p53 signatures in aged mouse ovaries.
a, Heatmap of the SASP genes that were significantly upregulated in ovaries from aged mice (22 months) compared with young mice (4 months) (n = 10 biologically independent mice in young group, n = 9 biologically independent mice in aged group). b, Heatmap of the SASP genes that were significantly suppressed by Tri-1 treatment in aged mouse ovaries (n = 4 biologically independent mice per group). c-e, Ingenuity Pathway Analysis of the 1920 genes that were significantly different in aged vs young mice ovaries. Common gene expression changes induced by Tri-1 and auranofin treatments showed expected common inhibition of GSR and TXNRD1 regulators (c). Transcription factors with altered activity were listed with p53 and p16 among them (d). P values were calculated by a Fisher Exact Test estimated by Ingenuity Pathway Analysis Software. Both these two age-associated signatures were not affected with either Tri-1 or auranofin treatment in the aged mice (e). P values were calculated by hypergeometrical test.
Extended Data Fig. 10 Suppression of SASP by Tri-1 treatment in aged mouse ovary.
a, Expression of the indicated SASP genes in ovaries from young (4 months) or aged (22 months) mice treated with Tri-1 or vehicle control was determined by RT-qPCR. n = 4 biologically independent mice per group. b,c, Immunoblot of the indicated proteins in the ovary tissues harvested from aged mice (22 months) with or without Tri-1 or auranofin treatments (b). The intensity of the indicated proteins was quantified by NIH ImageJ software and normalized against a loading control β-actin expression (c). n = 5 biologically independent mice per group. d, Expression of p16 in ovaries from young (4 months) or aged (22 months) mice treated with Tri-1 or vehicle control was determined by RT-qPCR. n = 4 biologically independent mice per group. Data represent mean ± s.e.m. P-values were calculated using a two-tailed t test.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12 and unprocessed western blot and gel source data for supplementary figures.
Supplementary Tables 1 and 2
Supplementary Table 1: List of proteins identified with at least five peptides and at least fivefold higher intensity in purified CCFs than in the control condition. Supplementary Table 2: List of primers used in this study.
Supplementary Data 1
Numerical source data for Supplementary Fig. 1.
Supplementary Data 2
Numerical source data for Supplementary Fig. 2.
Supplementary Data 3
Numerical source data for Supplementary Fig. 3.
Supplementary Data 4
Numerical source data for Supplementary Fig. 4.
Supplementary Data 5
Numerical source data for Supplementary Fig. 5.
Supplementary Data 6
Numerical source data for Supplementary Fig. 7.
Supplementary Data 7
Numerical source data for Supplementary Fig. 8.
Supplementary Data 8
Numerical source data for Supplementary Fig. 11.
Supplementary Data 9
Numerical source data for Supplementary Fig. 12.
Source data
Source Data Fig. 1
Unprocessed western blots for Fig. 1.
Source Data Fig. 1
Numerical source data for Fig. 1.
Source Data Fig. 2
Unprocessed western blots for Fig. 2.
Source Data Fig. 2
Unprocessed EMSA gel scan for Fig. 2l.
Source Data Fig. 2
Numerical source data for Fig. 2.
Source Data Fig. 3
Numerical source data for Fig. 3.
Source Data Fig. 4
Unprocessed western blots for Fig. 4.
Source Data Fig. 4
Numerical source data for Fig. 4.
Source Data Extended Data Fig. 1
Unprocessed western blots for Extended Data Fig. 1.
Source Data Extended Data Fig. 1
Numerical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Numerical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Unprocessed western blots for Extended Data Fig. 3.
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Unprocessed western blots for Extended Data Fig. 4.
Source Data Extended Data Fig. 4
Numerical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Unprocessed western blots for Extended Data Fig. 5.
Source Data Extended Data Fig. 5
Numerical source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Unprocessed western blots for Extended Data Fig. 6.
Source Data Extended Data Fig. 6
Numerical source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Numerical source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Unprocessed western blots for Extended Data Fig. 8.
Source Data Extended Data Fig. 8
Statistical source data for Extended Data Fig. 8.
Source Data Extended Data Fig. 10
Unprocessed western blots for Extended Data Fig. 10.
Source Data Extended Data Fig. 10
Statistical source data for Extended Data Fig. 10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hao, X., Zhao, B., Towers, M. et al. TXNRD1 drives the innate immune response in senescent cells with implications for age-associated inflammation. Nat Aging 4, 185–197 (2024). https://doi.org/10.1038/s43587-023-00564-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-023-00564-1