Abstract
Whereas cholesterol is vital for cell growth, proliferation, and remodeling, dysregulation of cholesterol metabolism is associated with multiple age-related pathologies. Here we show that senescent cells accumulate cholesterol in lysosomes to maintain the senescence-associated secretory phenotype (SASP). We find that induction of cellular senescence by diverse triggers enhances cellular cholesterol metabolism. Senescence is associated with the upregulation of the cholesterol exporter ABCA1, which is rerouted to the lysosome, where it moonlights as a cholesterol importer. Lysosomal cholesterol accumulation results in the formation of cholesterol-rich microdomains on the lysosomal limiting membrane enriched with the mammalian target of rapamycin complex 1 (mTORC1) scaffolding complex, thereby sustaining mTORC1 activity to support the SASP. We further show that pharmacological modulation of lysosomal cholesterol partitioning alters senescence-associated inflammation and in vivo senescence during osteoarthritis progression in male mice. Our study reveals a potential unifying theme for the role of cholesterol in the aging process through the regulation of senescence-associated inflammation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data supporting the findings of this study are available within the paper and its Supplementary Tables. The RNA-seq data generated by this study are deposited in the Gene Expression Omnibus database under project accession number GSE222676. Publicly available datasets used in this study are as follows: GSE64553 (Extended Data Fig. 2b), GSE57218 (Extended Data Fig. 9a), GTEx v8 (Extended Data Fig. 10a) and Cancer Cell Line Encyclopedia (DepMap portal (Public 21Q1); Extended Data Fig. 10b). Source data are provided with this paper.
References
Zhu, J. & Thompson, C. B. Metabolic regulation of cell growth and proliferation. Nat. Rev. Mol. Cell Biol. 20, 436–450 (2019).
Goldstein, J. L. & Brown, M. S. A century of cholesterol and coronaries: from plaques to genes to statins. Cell 161, 161–172 (2015).
Levental, I., Levental, K. R. & Heberle, F. A. Lipid rafts: controversies resolved, mysteries remain. Trends Cell Biol. 30, 341–353 (2020).
Tall, A. R. & Yvan-Charvet, L. Cholesterol, inflammation and innate immunity. Nat. Rev. Immunol. 15, 104–116 (2015).
Huang, B., Song, B.-l. & Xu, C. Cholesterol metabolism in cancer: mechanisms and therapeutic opportunities. Nat. Metab. 2, 132–141 (2020).
Song, Y., Liu, J., Zhao, K., Gao, L. & Zhao, J. Cholesterol-induced toxicity: an integrated view of the role of cholesterol in multiple diseases. Cell Metab. 33, 1911–1925 (2021).
Riscal, R., Skuli, N. & Simon, M. C. Even cancer cells watch their cholesterol! Mol. Cell 76, 220–231 (2019).
Castellano, B. M. et al. Lysosomal cholesterol activates mTORC1 via an SLC38A9–Niemann-Pick C1 signaling complex. Science 355, 1306–1311 (2017).
Shin, H. R. et al. Lysosomal GPCR-like protein LYCHOS signals cholesterol sufficiency to mTORC1. Science 377, 1290–1298 (2022).
Johnson, A. A. & Stolzing, A. The role of lipid metabolism in aging, lifespan regulation, and age-related disease. Aging Cell 18, e13048 (2019).
Broadfield, L. A., Pane, A. A., Talebi, A., Swinnen, J. V. & Fendt, S.-M. Lipid metabolism in cancer: new perspectives and emerging mechanisms. Dev. Cell 56, 1363–1393 (2021).
Mutlu, A. S., Duffy, J. & Wang, M. C. Lipid metabolism and lipid signals in aging and longevity. Dev. Cell 56, 1394–1407 (2021).
Thelen, A. M. & Zoncu, R. Emerging roles for the lysosome in lipid metabolism. Trends Cell Biol. 27, 833–850 (2017).
Childs, B. G. et al. Senescent cells: an emerging target for diseases of ageing. Nat. Rev. Drug Discov. 16, 718–735 (2017).
McHugh, D. & Gil, J. Senescence and aging: causes, consequences, and therapeutic avenues. J. Cell Biol. 217, 65–77 (2018).
Campisi, J. et al. From discoveries in ageing research to therapeutics for healthy ageing. Nature 571, 183–192 (2019).
Gorgoulis, V. et al. Cellular senescence: defining a path forward. Cell 179, 813–827 (2019).
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Borghesan, M., Hoogaars, W. M. H., Varela-Eirin, M., Talma, N. & Demaria, M. A senescence-centric view of aging: implications for longevity and disease. Trends Cell Biol. 30, 777–791 (2020).
Kang, C. Senolytics and senostatics: a two-pronged approach to target cellular senescence for delaying aging and age-related diseases. Mol. Cells 42, 821–827 (2019).
van Deursen, J. M. Senolytic therapies for healthy longevity. Science 364, 636–637 (2019).
Kirkland, J. L. & Tchkonia, T. Cellular senescence: a translational perspective. eBioMedicine 21, 21–28 (2017).
Birch, J. & Gil, J. Senescence and the SASP: many therapeutic avenues. Genes Dev. 34, 1565–1576 (2020).
Xu, M. et al. Senolytics improve physical function and increase lifespan in old age. Nat. Med. 24, 1246–1256 (2018).
Jeon, O. H. et al. Systemic induction of senescence in young mice after single heterochronic blood exchange. Nat. Metab. 4, 995–1006 (2022).
Guerrero, A. et al. Cardiac glycosides are broad-spectrum senolytics. Nat. Metab. 1, 1074–1088 (2019).
Xu, Q. et al. The flavonoid procyanidin C1 has senotherapeutic activity and increases lifespan in mice. Nat. Metab. 3, 1706–1726 (2021).
Pavlova, N. N., Zhu, J. & Thompson, C. B. The hallmarks of cancer metabolism: still emerging. Cell Metab. 34, 355–377 (2022).
Wiley, C. D. & Campisi, J. From ancient pathways to aging cells—connecting metabolism and cellular senescence. Cell Metab. 23, 1013–1021 (2016).
Kim, S. G., Buel, G. R. & Blenis, J. Nutrient regulation of the mTOR complex 1 signaling pathway. Mol. Cells 35, 463–473 (2013).
Shimobayashi, M. & Hall, M. N. Making new contacts: the mTOR network in metabolism and signalling crosstalk. Nat. Rev. Mol. Cell Biol. 15, 155–162 (2014).
Herranz, N. et al. mTOR regulates MAPKAPK2 translation to control the senescence-associated secretory phenotype. Nat. Cell Biol. 17, 1205–1217 (2015).
Laberge, R.-M. et al. MTOR regulates the pro-tumorigenic senescence-associated secretory phenotype by promoting IL1A translation. Nat. Cell Biol. 17, 1049–1061 (2015).
Carroll, B. et al. Persistent mTORC1 signaling in cell senescence results from defects in amino acid and growth factor sensing. J. Cell Biol. 216, 1949–1957 (2017).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 168, 960–976 (2017).
Liu, G. Y. & Sabatini, D. M. mTOR at the nexus of nutrition, growth, ageing and disease. Nat. Rev. Mol. Cell Biol. 21, 183–203 (2020).
Wiley, C. D. & Campisi, J. The metabolic roots of senescence: mechanisms and opportunities for intervention. Nat. Metab. 3, 1290–1301 (2021).
Wiley, C. D. et al. Oxylipin biosynthesis reinforces cellular senescence and allows detection of senolysis. Cell Metab. 33, 1124–1136.e5 (2021).
Hernandez-Segura, A. et al. Unmasking transcriptional heterogeneity in senescent cells. Curr. Biol. 27, 2652–2660.e4 (2017).
Hernandez-Segura, A., Nehme, J. & Demaria, M. Hallmarks of cellular senescence. Trends Cell Biol. 28, 436–453 (2018).
Liu, S. et al. Simvastatin suppresses breast cancer cell proliferation induced by senescent cells. Sci. Rep. 5, 17895 (2016).
Weiss, W. A., Taylor, S. S. & Shokat, K. M. Recognizing and exploiting differences between RNAi and small-molecule inhibitors. Nat. Chem. Biol. 3, 739–744 (2007).
Housden, B. E. et al. Loss-of-function genetic tools for animal models: cross-species and cross-platform differences. Nat. Rev. Genet. 18, 24–40 (2017).
Kang, C. et al. The DNA damage response induces inflammation and senescence by inhibiting autophagy of GATA4. Science 349, aaa5612 (2015).
Moon, S.-H. et al. p53 represses the mevalonate pathway to mediate tumor suppression. Cell 176, 564–580.e19 (2019).
Plummer, A. M., Culbertson, A. T. & Liao, M. The ABCs of sterol transport. Annu Rev. Physiol. 83, 153–181 (2021).
Settembre, C. & Ballabio, A. Lysosome: regulator of lipid degradation pathways. Trends Cell Biol. 24, 743–750 (2014).
Lawrence, R. E. & Zoncu, R. The lysosome as a cellular centre for signalling, metabolism and quality control. Nat. Cell Biol. 21, 133–142 (2019).
Ballabio, A. & Bonifacino, J. S. Lysosomes as dynamic regulators of cell and organismal homeostasis. Nat. Rev. Mol. Cell Biol. 21, 101–118 (2020).
Shin, H. R. & Zoncu, R. The lysosome at the intersection of cellular growth and destruction. Dev. Cell 54, 226–238 (2020).
Narita, M. et al. Spatial coupling of mTOR and autophagy augments secretory phenotypes. Science 332, 966–970 (2011).
Rabanal-Ruiz, Y. et al. mTORC1 activity is supported by spatial association with focal adhesions. J. Cell Biol. 220, e202004010 (2021).
Ma, X. M. & Blenis, J. Molecular mechanisms of mTOR-mediated translational control. Nat. Rev. Mol. Cell Biol. 10, 307–318 (2009).
Favari, E. et al. Probucol inhibits ABCA1-mediated cellular lipid efflux. Arterioscler Thromb. Vasc. Biol. 24, 2345–2350 (2004).
Abu-Remaileh, M. et al. Lysosomal metabolomics reveals V-ATPase- and mTOR-dependent regulation of amino acid efflux from lysosomes. Science 358, 807–813 (2017).
Nieland, T. J. F. et al. Cross-inhibition of SR-BI- and ABCA1-mediated cholesterol transport by the small molecules BLT-4 and glyburide. J. Lipid Res. 45, 1256–1265 (2004).
Ramos, Y. F. M. et al. Genes involved in the osteoarthritis process identified through genome wide expression analysis in articular cartilage; the RAAK study. PLoS ONE 9, e103056 (2014).
Lee, Y. et al. Coordinate regulation of the senescent state by selective autophagy. Dev. Cell 56, 1512–1525.e7 (2021).
Jeon, O. H., David, N., Campisi, J. & Elisseeff, J. H. Senescent cells and osteoarthritis: a painful connection. J. Clin. Invest. 128, 1229–1237 (2018).
Kang, D. et al. Stress-activated miR-204 governs senescent phenotypes of chondrocytes to promote osteoarthritis development. Sci. Transl. Med. 11, eaar6659 (2019).
Jeon, O. H. et al. Local clearance of senescent cells attenuates the development of post-traumatic osteoarthritis and creates a pro-regenerative environment. Nat. Med. 23, 775–781 (2017).
Di Micco, R., Krizhanovsky, V., Baker, D. & d’Adda di Fagagna, F. Cellular senescence in ageing: from mechanisms to therapeutic opportunities. Nat. Rev. Mol. Cell Biol. 22, 75–95 (2021).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Crouse, J. R., Grundy, S. M. & Ahrens, E. H.Jr. Cholesterol distribution in the bulk tissues of man: variation with age. J. Clin. Invest. 51, 1292–1296 (1972).
Gostynski, M. et al. Analysis of the relationship between total cholesterol, age, body mass index among males and females in the WHO MONICA Project. Int. J. Obes. Relat. Metab. Disord. 28, 1082–1090 (2004).
Yi, S.-W., Yi, J.-J. & Ohrr, H. Total cholesterol and all-cause mortality by sex and age: a prospective cohort study among 12.8 million adults. Sci. Rep. 9, 1596 (2019).
Choi, W.-S. et al. The CH25H–CYP7B1–RORα axis of cholesterol metabolism regulates osteoarthritis. Nature 566, 254–258 (2019).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Liu, F. et al. Long non-coding RNA SNHG6 couples cholesterol sensing with mTORC1 activation in hepatocellular carcinoma. Nat. Metab. 4, 1022–1040 (2022).
Wang, Y., Xu, Q., Sack, L., Kang, C. & Elledge, S. J. A gain-of-function senescence bypass screen identifies the homeobox transcription factor DLX2 as a regulator of ATM–p53 signaling. Genes Dev. 30, 293–306 (2016).
Meerbrey, K. L. et al. The pINDUCER lentiviral toolkit for inducible RNA interference in vitro and in vivo. Proc. Natl Acad. Sci. USA 108, 3665–3670 (2011).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Coppé, J.-P., Desprez, P.-Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. 5, 99–118 (2010).
Meng, Y., Heybrock, S., Neculai, D. & Saftig, P. Cholesterol handling in lysosomes and beyond. Trends Cell Biol. 30, 452–466 (2020).
Kaushik, S., Massey, A. C. & Cuervo, A. M. Lysosome membrane lipid microdomains: novel regulators of chaperone-mediated autophagy. EMBO J. 25, 3921–3933 (2006).
Choi, R.-Y., Ji, M., Lee, M.-K. & Paik, M.-J. Metabolomics study of serum from a chronic alcohol-fed rat model following administration of defatted Tenebrio molitor larva fermentation extract. Metabolites 10, 436 (2020).
Zeng, H. et al. mTORC1 couples immune signals and metabolic programming to establish Treg-cell function. Nature 499, 485–490 (2013).
Wang, J.-Q. et al. Inhibition of ASGR1 decreases lipid levels by promoting cholesterol excretion. Nature 608, 413–420 (2022).
Lee, Y. et al. A flow-cytometry-based assessment of global protein synthesis in human senescent cells. STAR Protoc. 2, 100809 (2021).
Glasson, S. S., Chambers, M. G., Van Den Berg, W. B. & Little, C. B. The OARSI histopathology initiative – recommendations for histological assessments of osteoarthritis in the mouse. Osteoarthritis Cartilage 18(Suppl. 3), S17–S23 (2010).
Acknowledgements
We thank J.B. Kim and N. Kim for sharing reagents. We are also grateful to all of the members of the Kang lab and the Kim lab for their support. This work was supported by the Suh Kyungbae Foundation (SUHF-17020068), the National Research Foundation of Korea (NRF-2020R1A5A1018081), and the Samsung Science & Technology Foundation (SSTF-BA2201-09) to C.K. J.-H.K. was supported by the Suh Kyungbae Foundation (SUHF-18010068) and the National Research Foundation of Korea (NRF-2020R1A2C2012300).
Author information
Authors and Affiliations
Contributions
K.R. and C.K. conceived and designed the project. J.N. and J.-H.K. conceived and designed male mouse OA experiments. K.R., Y.K., Y.J. and C.K. performed most of the experiments and analyzed the data. J.N., D.K. and J.-H.K. performed and analyzed male mouse OA experiments. M.J. and M.-J.P. performed GC–MS/MS and amino acid profiling analysis. J.-H.K. and C.K. supervised the project. H.C. and M.-S.K. provided multiple types of senescent samples for analysis. M.-S.K. performed ChIP experiments. Y.L. performed global protein synthesis analyses. J.K. performed GSVA. J.C. provided intellectual contribution on the project. K.R., J.N., Y.K., Y.J., J.-H.K. and C.K. wrote the paper. Y.K. and Y.J. contributed equally to this work. All authors discussed the results and commented on the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Christopher Wiley and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Christoph Schmitt, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cholesterol metabolism is altered in DNA damage-induced, replicative, and oncogene-induced senescence.
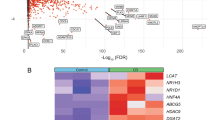
a, Schematic representation of the gene expression profiling for DNA damage-induced senescence followed by GSEA with a specific focus on metabolic pathway. VMH denotes the virtual metabolic human database. PRO and SEN denote sham-irradiated proliferating and DNA damage-induced senescent cells, respectively. DEGs denotes differentially expressed genes. b, GSEA of differentially expressed genes in replicative senescent MRC5 cells for the cholesterol homeostasis pathway [left, false discovery rate (q-value) and normalized enrichment score (NES) calculated by GSEA]. Abundance of the indicated mRNAs analyzed by qRT-PCR upon replicative senescence (middle, n = 3 biologically independent samples, mean ± SEM, two-sided unpaired t test). PRO and SEN (REP) denote proliferating and replicative senescent IMR90 cells, respectively. Enzymatic measurement of cholesterol upon replicative senescence in IMR90 cells (right, n = 3 biologically independent samples, mean ± SEM, two-sided unpaired t test). c, Abundance of the indicated mRNAs analyzed by qRT-PCR upon oncogene-induced senescence (left, n = 3 biologically independent samples, mean ± SEM, two-sided unpaired t test). PRO and SEN (OIS) denote sham-treated and doxycycline-treated IMR90 cells carrying a Dox-inducible (Tet-On) vector expressing H-RasV12. Enzymatic measurement of cholesterol upon oncogene-induced senescence (right, n = 4 biologically independent samples, mean ± SEM, two-sided unpaired t test). d, Enzymatic measurement of cholesterol upon knockdown of the indicated genes during senescence (mean ± SEM, n = 3 biologically independent samples, one-way ANOVA test with Tukey’s multiple comparisons test).
Extended Data Fig. 2 Cholesterol metabolism is associated with the SASP during DNA damage-induced and oncogene-induced senescence.
a, Enzymatic measurement of cholesterol upon MβCD treatment for 2 days in DMEM supplemented with 15% FBS during DNA damage-induced senescence (n = 3 biologically independent samples, mean ± SEM, two-sided unpaired t test). b, Abundance of the indicated mRNAs analyzed by qRT-PCR upon cholesterol depletion (left, n = 3 biologically independent samples, mean ± SEM, one-way ANOVA test with Tukey’s multiple comparisons test). ChDS denotes cholesterol depletion by incubating the cells with DMEM supplemented with 15% charcoal-stripped FBS. Abundance of the indicated proteins analyzed by immunoblotting upon senescence induction in BJ cells (right, n = 2 biologically independent experiments). The cells were starved for amino acids and serum, and then restimulated for the indicated time. c, Abundance of the indicated mRNAs analyzed by qRT-PCR (left, n = 3 biologically independent samples, mean ± SEM, one-way ANOVA test with Tukey’s multiple comparisons test). Abundance of the indicated proteins analyzed by immunoblotting (right, n = 3 biologically independent experiments). Lov denotes lovastatin. d-e, Abundance of the indicated mRNAs analyzed by qRT-PCR (mean ± SEM, n = 3 biologically independent samples, one-way ANOVA test with Tukey’s multiple comparisons test). Chol denotes cholesterol stimulation.
Extended Data Fig. 3 ABCA1 plays a key role in SASP regulation.
a, Abundance of the indicated mRNAs analyzed by qRT-PCR upon senescence induction in IMR90 cells expressing the indicated shRNAs (mean ± SEM, n = 6 and 4 biologically independent samples for IL1A/IL1B/IL8 and HMGCR/LRP1/SCARB1, respectively, RM one-way ANOVA test with Tukey’s multiple comparisons test). b, Abundance of the indicated proteins analyzed by immunoblotting upon senescence induction in BJ cells (top, n = 2 biologically independent experiments). Abundance of the indicated mRNAs analyzed by qRT-PCR upon senescence induction in BJ cells (bottom, n = 2 biologically independent samples, mean ± SEM). PRO and SEN denote sham-treated proliferating and DNA damage-induced senescent cells, respectively. C denotes control ABCA1 WT cells. c, Abundance of the indicated proteins analyzed by immunoblotting (left, n = 3 biologically independent experiments). Abundance of the indicated mRNAs analyzed by qRT-PCR upon oncogene-induced senescence (right, n = 3 biologically independent samples, mean ± SEM, one-way ANOVA test with Tukey’s multiple comparisons test). d, Abundance of the indicated mRNAs analyzed by qRT-PCR in IMR90 cells stimulated with or without cholesterol during senescence (mean ± SEM, n = 3 biologically independent samples, one-way ANOVA test with Tukey’s multiple comparisons test).
Extended Data Fig. 4 The GATA4-NF-kB axis induces ABCA1 during senescence.
a, Abundance of the indicated proteins analyzed by immunoblotting upon senescence induction in IMR90 cells (left, n = 3 biologically independent experiments). Abundance of the indicated mRNAs analyzed by qRT-PCR upon senescence induction in IMR90 cells (right, from 1 biologically independent experiment, two times each experiment was repeated independently with similar results). b, Abundance of the indicated proteins analyzed by immunoblotting upon senescence induction in BJ cells expressing the PmiR-146a-GFP SASP reporter (n = 1 biologically independent experiment). c, Abundance of the indicated proteins analyzed by immunoblotting upon replicative senescence in IMR90 cells (left, n = 2 and 3 biologically independent samples for PRO and SEN (REP), respectively). Abundance of ABCA1 mRNA analyzed by qRT-PCR upon replicative senescence in IMR90 cells (right, n = 2 and 3 biologically independent samples for PRO and SEN (REP), respectively, mean ± SEM, two-sided unpaired t test). PRO and SEN (REP) denote proliferating and replicative senescent IMR90 cells. d, Schematic representation of the promoter region of ABCA1 with the indicated qPCR primers (left). Potential RELA and GATA4 motifs are shown. IMR90 cells carrying a Dox-inducible (Tet-On) vector expressing HA-GATA4 were cross linked with formaldehyde and protein extracts were immunoprecipitated with either HA or RELA antibodies. DNA was eluted, the cross-links were reversed, and the DNA was analyzed by qPCR using the indicated primers. Signals obtained from ChIP were normalized to signals obtained from an input sample (right, n = 3 biologically independent samples, % Input, mean ± SEM, two-sided unpaired t test). e, Abundance of the indicated proteins analyzed by immunoblotting upon senescence induction in IMR90 cells carrying a Dox-inducible (Tet-On) vector expressing dominant negative p53 (Tet-p53DN) (left, n = 3 biologically independent samples). Abundance of the indicated mRNAs analyzed by qRT-PCR (right, n = 3 biologically independent samples, mean ± SEM, two-way ANOVA test with Tukey’s multiple comparisons test).
Extended Data Fig. 5 ABCA1 is localized in the lysosome.
a, Immunocytochemistry of the indicated proteins (left, n = 2 biologically independent experiments). Scale bar, 50 μm. Abundance of the indicated proteins analyzed by immunoblotting upon LysoIP (right, n = 2 biologically independent experiments). PRO and SEN denote sham-treated proliferating and senescent cells. b, Representative images of cellular cholesterol and lysosome staining with filipin and lysotracker red (Lyso), respectively, upon GATA4 induction (left). Scale bar, 50 μm. Pearson’s correlation coefficient for colocalization of cellular cholesterol and lysosome (right, n = 2 biologically independent experiments, 5-7 randomly chosen fields from each experiment, box plots show the 1st and 3rd quartiles; whiskers show the minimum and maximum values, two-sided unpaired t test).
Extended Data Fig. 6 Pharmacological modulation of lysosomal cholesterol affects the SASP.
a, Abundance of the indicated mRNAs analyzed by qRT-PCR (n = 3 biologically independent samples, mean ± SEM, one-way ANOVA test with Tukey’s multiple comparisons test). SEN (OIS) and Prob denote oncogene-induced senescence and probucol, respectively. b, Abundance of the indicated mRNAs analyzed by qRT-PCR (n = 3 biologically independent samples, mean ± SEM, one-way ANOVA test with Tukey’s multiple comparisons test). Glyb denotes glyburide. c, Pearson’s correlation coefficient for colocalization of cellular cholesterol and lysosome (left, n = 2 biologically independent experiments, 10–20 randomly chosen fields from each experiment, box plots show the 1st and 3rd quartiles; whiskers show the minimum and maximum values, one-way ANOVA test with Tukey’s multiple comparisons test). Abundance of the indicated mRNAs analyzed by qRT-PCR (right, n = 3 biologically independent samples, mean ± SEM, one-way ANOVA test with Tukey’s multiple comparisons test). SEN denotes DNA damage-induced senescence. d, Abundance of the indicated mRNAs analyzed by qRT-PCR (mean ± SEM, n = 3 biologically independent samples, one-way ANOVA test with Tukey’s multiple comparisons test).
Extended Data Fig. 7 ABCA1 sustains a lysosomal mTORC1 recruitment during senescence.
Immunocytochemistry of the indicated proteins (left). Scale bar, 50 μm. Pearson’s correlation coefficient for colocalization of mTOR and LAMP2 in the indicated cell lines during senescence upon amino acid starvation (2 h) and refeeding (30 m) (right, n = 3 biologically independent experiments, 10–23 randomly chosen individual cells from each experiment, box plots show the 1st and 3rd quartiles; whiskers show the minimum and maximum values, one-way ANOVA test with Tukey’s multiple comparisons test). PRO and SEN denote sham-treated proliferating and senescent cells.
Extended Data Fig. 8 Lysosomal amino acid levels do not change during senescence.
Absolute quantification of amino acids upon LysoIP by Gas chromatography-tandem mass spectrometry (GC-MS/MS, n = 3 biologically independent samples, mean ± SEM, two-way ANOVA test with Tukey’s multiple comparisons test). WT and KO denote ABCA1 wild-type and knock-out cells. PRO and SEN denote sham-treated proliferating and senescent cells.
Extended Data Fig. 9 Cholesterol metabolism and ABCA1 modulate chondrocyte senescence and OA progression.
a, GSEA of differentially expressed genes in human OA for the cholesterol homeostasis pathway (left, top) and the mTOR signaling pathway (left, bottom). False discovery rate (q-value) and normalized enrichment score (NES) calculated by GSEA. Pearson’s correlation analysis of expression of ABCA1 and the indicated genes in human OA (right, n = 73 biologically independent individuals). b, Abundance of the indicated mRNAs analyzed by qRT-PCR (n = 6 biologically independent samples, mean ± SEM, two-sided unpaired t test). PRO and SEN denote proliferating and senescent chondrocytes, respectively. c, Abundance of the indicated mRNAs analyzed by qRT-PCR (n = 3 biologically independent samples, mean ± SEM, one-way ANOVA test with Tukey’s multiple comparisons test). Lov denotes lovastatin. d, Abundance of the indicated proteins analyzed by immunoblotting (n = 2 biologically independent experiments). ChDS denotes cholesterol depletion by incubating the cells with DMEM supplemented with charcoal-stripped FBS. The cells were starved for amino acids and serum, and then restimulated for the indicated time. e, Representative images of immunohistochemistry of the indicated proteins in cartilage sections of male mice. Scale bar, 50 μm. Tidemarks were represented by dotted lines. f, The percentage of weight placed on the DMM-operated limb versus the contralateral limb of sham- and DMM-operated mice treated with or without probucol or glyburide determined using a static weight bearing test (left). Paw withdrawal threshold in response to von Frey filaments in sham-operated and DMM-operated mice treated with or without probucol or glyburide (right). n = 5, 6, 9, 9, 5, 5, 7, and 8 biologically independent animals for Sham w/o Prob, Sham w/ Prob, DMM w/o Prob, DMM w/ Prob, Sham w/o Glyb, Sham w/ Glyb, DMM w/o Glyb, and DMM w/ Glyb, box plots show the 1st and 3rd quartiles; whisker show the minimum and maximum values, two-way ANOVA test with Holm-Sidak’s multiple comparisons test.
Extended Data Fig. 10 Expression of ABCA1 is associated with the SASP signature and inflammation in multiple tissues during human aging as well as in cancer.
a, Correlation analysis of the SASP GSVA scores and ABCA1 expression in human aging. Samples from the indicated tissues in the Genotype-Tissue Expression (GTEx) database were divided into two age groups [‘Young’ (20~39 years old) and ‘Old’ (60~79 years old)], and the Pearson’s correlation analysis between the SASP GSVA scores and ABCA1 expression was performed for each age group (two-tailed, n = 212, 73, 155, 17, 180, 33, 120, 9, 155, 16, 52, and 30 biologically independent individuals for lung old, lung young, pituitary old, pituitary young, heart old, heart young, brain frontal cortex old, brain frontal cortex young, brain cerebellum old, brain cerebellum young, salivary gland old, and salivary gland young, respectively). b, Box plot analysis comparing the abundance of the indicated genes between ABCA1low and ABCA1high cells (n = 347 and 345 biologically independent cell lines for ABCA1low and ABCA1high, respectively, box plots show the 1st and 3rd quartiles; whiskers show the minimum and maximum values, two-sided unpaired t-test). Cells were categorized into 2 groups [ABCA1low (bottom 25%) and ABCA1high (top 25%)] according to ABCA1 expression in the cancer cell line encyclopedia.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Tables
Supplementary Tables 1–9.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data and unprocessed western blots.
Source Data Extended Data Fig. 9
Unprocessed western blots.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Roh, K., Noh, J., Kim, Y. et al. Lysosomal control of senescence and inflammation through cholesterol partitioning. Nat Metab 5, 398–413 (2023). https://doi.org/10.1038/s42255-023-00747-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-023-00747-5
This article is cited by
-
Cholesterol biosynthetic pathway induces cellular senescence through ERRα
npj Aging (2024)
-
The senescence-associated secretory phenotype and its physiological and pathological implications
Nature Reviews Molecular Cell Biology (2024)
-
A homoeostatic switch causing glycerol-3-phosphate and phosphoethanolamine accumulation triggers senescence by rewiring lipid metabolism
Nature Metabolism (2024)
-
Cholesterol drives inflammatory senescence
Nature Metabolism (2023)