Abstract
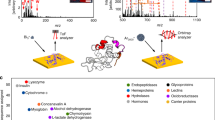
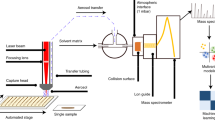
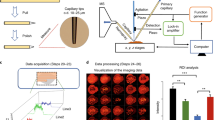
Rapid, sensitive, precise and accurate analysis of samples in their native in vivo environment is critical to better decipher physiological and physiopathological mechanisms. SpiderMass is an ambient mass spectrometry (MS) system designed for mobile in vivo and real-time surface analyses of biological tissues. The system uses a fibered laser, which is tuned to excite the most intense vibrational band of water, resulting in a process termed water-assisted laser desorption/ionization (WALDI). The water molecules act as an endogenous matrix in a matrix-assisted laser desorption ionization (MALDI)-like scenario, leading to the desorption/ionization of biomolecules (lipids, metabolites and proteins). The ejected material is transferred to the mass spectrometer through an atmospheric interface and a transfer line that is several meters long. Here, we formulate a three-stage procedure that includes (i) a laser system setup coupled to a Waters Q-TOF or Thermo Fisher Q Exactive mass analyzer, (ii) analysis of specimens and (iii) data processing. We also describe the optimal setup for the analysis of cell cultures, fresh-frozen tissue sections and in vivo experiments on skin. With proper optimization, the system can be used for a variety of different targets and applications. The entire procedure takes 1–2 d for complex samples.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All published data presented here are publicly available and can be found in the PRIDE Archive (https://www.ebi.ac.uk/pride/archive/). The dog sarcoma data are available from the ProteomeXchange Consortium (PXD010990), Real-time molecular diagnosis of tumors using water-assisted laser desorption/ionization mass spectrometry technology.
References
Takats, Z., Strittmatter, N. & McKenzie, J. S. Ambient mass spectrometry in cancer research. Adv. Cancer Res. 134, 231–256 (2017).
Zhang, J. et al. Nondestructive tissue analysis for ex vivo and in vivo cancer diagnosis using a handheld mass spectrometry system. Sci. Transl. Med. 9, eaan3968 (2017).
Fatou, B. et al. In vivo real-time mass spectrometry for guided surgery application. Sci. Rep. 6, 25919 (2016).
Balog, J. et al. Intraoperative tissue identification using rapid evaporative ionization mass spectrometry. Sci. Transl. Med. 5, 194ra93 (2013).
Huang, M.-Z., Cheng, S.-C., Cho, Y.-T. & Shiea, J. Ambient ionization mass spectrometry: a tutorial. Anal. Chim. Acta 702, 1–15 (2011).
Alberici, R. M. et al. Ambient mass spectrometry: bringing MS into the “real world”. Anal. Bioanal. Chem. 398, 265–294 (2010).
Takáts, Z., Wiseman, J. M., Gologan, B. & Cooks, R. G. Mass spectrometry sampling under ambient conditions with desorption electrospray ionization. Science 306, 471–473 (2004).
Laiko, V. V., Baldwin, M. A. & Burlingame, A. L. Atmospheric pressure matrix-assisted laser desorption/ionization mass spectrometry. Anal. Chem. 72, 652–657 (2000).
Laiko, V. V. et al. Desorption/ionization of biomolecules from aqueous solutions at atmospheric pressure using an infrared laser at 3 μm. J. Am. Soc. Mass Spectrom. 13, 354–361 (2002).
Nemes, P. & Vertes, A. Laser ablation electrospray ionization for atmospheric pressure molecular imaging mass spectrometry. Methods Mol. Biol. 656, 159–171 (2010).
Nemes, P., Woods, A. S. & Vertes, A. Simultaneous imaging of small metabolites and lipids in rat brain tissues at atmospheric pressure by laser ablation electrospray ionization mass spectrometry. Anal. Chem. 82, 982–988 (2010).
Sampson, J. S., Hawkridge, A. M. & Muddiman, D. C. Generation and detection of multiply-charged peptides and proteins by matrix-assisted laser desorption electrospray ionization (MALDESI) Fourier transform ion cyclotron resonance mass spectrometry. J. Am. Soc. Mass Spectrom. 17, 1712–1716 (2006).
Wu, C., Dill, A. L., Eberlin, L. S., Cooks, R. G. & Ifa, D. R. Mass spectrometry imaging under ambient conditions. Mass Spectrom. Rev. 32, 218–243 (2013).
Chen, H., Talaty, N. N., Takáts, Z. & Cooks, R. G. Desorption electrospray ionization mass spectrometry for high-throughput analysis of pharmaceutical samples in the ambient environment. Anal. Chem. 77, 6915–6927 (2005).
Talaty, N., Takáts, Z. & Cooks, R. G. Rapid in situ detection of alkaloids in plant tissue under ambient conditions using desorption electrospray ionization. Analyst 130, 1624–1633 (2005).
Takáts, Z., Wiseman, J. M. & Cooks, R. G. Ambient mass spectrometry using desorption electrospray ionization (DESI): instrumentation, mechanisms and applications in forensics, chemistry, and biology. J. Mass Spectrom. 40, 1261–1275 (2005).
Katona, M., Dénes, J., Skoumal, R., Tóth, M. & Takáts, Z. Intact skin analysis by desorption electrospray ionization mass spectrometry. Analyst 136, 835–840 (2011).
Hayashi, Y. et al. Intact metabolite profiling of mouse brain by probe electrospray ionization/triple quadrupole tandem mass spectrometry (PESI/MS/MS) and its potential use for local distribution analysis of the brain. Anal. Chim. Acta 983, 160–165 (2017).
Zaitsu, K. et al. Intact endogenous metabolite analysis of mice liver by probe electrospray ionization/triple quadrupole tandem mass spectrometry and its preliminary application to in vivo real-time analysis. Anal. Chem. 88, 3556–3561 (2016).
Zaitsu, K. et al. In vivo real-time monitoring system using probe electrospray ionization/tandem mass spectrometry for metabolites in mouse brain. Anal. Chem. 90, 4695–4701 (2018).
Liu, J., Cooks, R. G. & Ouyang, Z. Biological tissue diagnostics using needle biopsy and spray ionization mass spectrometry. Anal. Chem. 83, 9221–9225 (2011).
Song, Y., Liao, J., Zha, C., Wang, B. & Liu, C. C. A novel approach to determine the tyrosine concentration in human plasma by DART-MS/MS. Anal. Methods 7, 1600–1605 (2015).
Zhou, M., McDonald, J. F. & Fernández, F. M. Optimization of a direct analysis in real time/time-of-flight mass spectrometry method for rapid serum metabolomic fingerprinting. J. Am. Soc. Mass Spectrom 21, 68–75 (2010).
Schäfer, K.-C. et al. In vivo, in situ tissue analysis using rapid evaporative ionization mass spectrometry. Angew. Chem. Int. Edn 48, 8240–8242 (2009).
Phelps, D. L. et al. The surgical intelligent knife distinguishes normal, borderline and malignant gynaecological tissues using rapid evaporative ionisation mass spectrometry (REIMS). Br. J. Cancer 118, 1349–1358 (2018).
St John, E. R. et al. Rapid evaporative ionisation mass spectrometry of electrosurgical vapours for the identification of breast pathology: towards an intelligent knife for breast cancer surgery. Breast Cancer Res. 19, 59 (2017).
Balog, J. et al. In vivo endoscopic tissue identification by rapid evaporative ionization mass spectrometry (REIMS). Angew. Chem. Int. Edn 54, 11059–11062 (2015).
Sans, M. et al. Performance of the MasSpec Pen for rapid diagnosis of ovarian cancer. Clin. Chem. 65, 674–683 (2019).
Fournier, I. et al. Device for real-time in vivo molecular analysis. WO2016046748 (A1) (2014).
Schäfer, K.-C. et al. In situ, real-time identification of biological tissues by ultraviolet and infrared laser desorption ionization mass spectrometry. Anal. Chem. 83, 1632–1640 (2011).
Berkenkamp, S., Karas, M. & Hillenkamp, F. Ice as a matrix for IR-matrix-assisted laser desorption/ionization: mass spectra from a protein single crystal. Proc. Natl Acad. Sci. USA 93, 7003–7007 (1996).
Pirkl, A., Soltwisch, J., Draude, F. & Dreisewerd, K. Infrared matrix-assisted laser desorption/ionization orthogonal-time-of-flight mass spectrometry employing a cooling stage and water ice as a matrix. Anal. Chem. 84, 5669–5676 (2012).
Fatou, B. et al. Remote atmospheric pressure infrared matrix-assisted laser desorption-ionization mass spectrometry (remote IR-MALDI MS) of proteins. Mol. Cell. Proteomics 17, 1637–1649 (2018).
Saudemont, P. et al. Real-time molecular diagnosis of tumors using water-assisted laser desorption/ionization mass spectrometry technology. Cancer Cell 34, 840–851 (2018).
Fatou, B. et al. Real time and in vivo pharmaceutical and environmental studies with SpiderMass instrument. J. Biotechnol. 281, 61–66 (2018).
Fatou, B., Salzet, M. & Fournier, I. Real time human micro-organisms biotyping based on water-assisted laser desorption/ionization. EuroBiotech J. 3, 97–104 (2019).
Woolman, M. et al. Rapid determination of medulloblastoma subgroup affiliation with mass spectrometry using a handheld picosecond infrared laser desorption. Chem. Sci. 8, 6508–6519 (2017).
Zou, J. et al. Ambient mass spectrometry imaging with picosecond infrared laser ablation electrospray ionization (PIR-LAESI). Anal. Chem. 87, 12071–12079 (2015).
Woolman, M. et al. Optimized mass spectrometry analysis workflow with polarimetric guidance for ex vivo and in situ sampling of biological tissues. Sci. Rep. 7, 468 (2017).
Strohalm, M., Hassman, M., Košata, B. & Kodíček, M. mMass data miner: an open source alternative for mass spectrometric data analysis. Rapid Commun. Mass Spectrom. 22, 905–908 (2008).
Strohalm, M., Kavan, D., Novák, P., Volný, M. & Havlíček, V. mMass 3: a cross-platform software environment for precise analysis of mass spectrometric data. Anal. Chem. 82, 4648–4651 (2010).
Gagnon, H. et al. Proprotein convertase 1/3 (PC1/3) in the rat alveolar macrophage cell line NR8383: localization, trafficking and effects on cytokine secretion. PLoS ONE 8, e61557 (2013).
Duhamel, M. et al. Molecular consequences of proprotein convertase 1/3 (PC1/3) inhibition in macrophages for application to cancer immunotherapy: a proteomic study. Mol. Cell. Proteomics 14, 2857–2877 (2015).
Duhamel, M. et al. Paclitaxel treatment and proprotein convertase 1/3 (PC1/3) knockdown in macrophages is a promising antiglioma strategy as revealed by proteomics and cytotoxicity studies. Mol. Cell. Proteomics 17, 1126–1143 (2018).
Acknowledgements
We thank all members of SATT-Nord involved in SpiderMass and, in particular, E. Rollet and F.-X. Denimal for their support of the project. This work was funded by the Ministère de l’Enseignement Supérieur, de la Recherche et de l’Innovation, Université de Lille and Inserm. The project was also funded by ANR-14-CE17-0021 REALITY’MS (I.F.), Inserm Programme PhysiCancer SPIDERMASS (M.S.), Région Hauts de France-EU FEDER O’DREAMS (D.T., I.F. and M.S.) and SIRIC ONCOLille grant INCa-DGOS-Inserm 6041aa (D.T., I.F. and M.S.). SpiderMass was awarded Best Breakthrough Innovation Prize by the MATWIN 2015 international board.
Author information
Authors and Affiliations
Contributions
N.O., P.S. and J.B. wrote the original draft of the manuscript. P.S., N.O. and J.-P.G. carried out the experiments. P.S. and I.F. developed the technique. Q.P. collected the samples and performed the histology. Y.-M.R. did histology and validated diagnostics. P.S., J.B. and I.F. analyzed the data. I.F., M.S. and D.T. designed the study. I.F., M.S. and Z.T. corrected the manuscript. I.F. and M.S. supervised the project and provided the funding.
Corresponding authors
Ethics declarations
Competing interests
J.B. is an employee of Waters Research Center. M.S. and I.F. are inventors on a patent (priority no. WO2015IB57301 20150922) related to part of the described protocol. Q.P. is an employee of OCR. D.T. is founder and CEO of OCR. Z.T. is a consultant for Waters Corporation. The system is under protection by patent CA2961491 A1 (29). The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Fatou, B. et al. Sci. Rep. 6, 25919 (2016): https://www.nature.com/articles/srep25919
Fatou, B. et al. J. Biotechnol. 281, 61–66 (2018): https://www.sciencedirect.com/science/article/pii/S0168165618305169
Saudemont, P. et al. Cancer Cell 34, 840–851.E4 (2018): https://www.cell.com/cancer-cell/fulltext/S1535-6108(18)30423-9
Supplementary information
Rights and permissions
About this article
Cite this article
Ogrinc, N., Saudemont, P., Balog, J. et al. Water-assisted laser desorption/ionization mass spectrometry for minimally invasive in vivo and real-time surface analysis using SpiderMass. Nat Protoc 14, 3162–3182 (2019). https://doi.org/10.1038/s41596-019-0217-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0217-8
This article is cited by
-
Human robotic surgery with intraoperative tissue identification using rapid evaporation ionisation mass spectrometry
Scientific Reports (2024)
-
Recent advances in entirely hand-held ionization sources for mass spectrometry
Analytical and Bioanalytical Chemistry (2024)
-
Spatial analysis of the glioblastoma proteome reveals specific molecular signatures and markers of survival
Nature Communications (2022)
-
Rapid ex vivo molecular fingerprinting of biofluids using laser-assisted rapid evaporative ionization mass spectrometry
Nature Protocols (2021)
-
Metabolic profiles of human brain parenchyma and glioma for rapid tissue diagnosis by targeted desorption electrospray ionization mass spectrometry
Analytical and Bioanalytical Chemistry (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.