Abstract
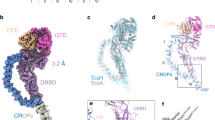
The evolution of virulence traits is central for the emergence or re-emergence of microbial pathogens and for their adaptation to a specific host1,2,3,4,5. Typhoid toxin is an essential virulence factor of the human-adapted bacterial pathogen Salmonella Typhi6,7, the cause of typhoid fever in humans8,9,10,11,12. Typhoid toxin has a unique A2B5 architecture with two covalently linked enzymatic ‘A’ subunits, PltA and CdtB, associated with a homopentameric ‘B’ subunit made up of PltB, which has binding specificity for the N-acetylneuraminic acid (Neu5Ac) sialoglycans6,13 prominently present in humans14. Here, we examine the functional and structural relationship between typhoid toxin and ArtAB, an evolutionarily related AB5 toxin encoded by the broad-host Salmonella Typhimurium15. We find that ArtA and ArtB, homologues of PltA and PltB, can form a functional complex with the typhoid toxin CdtB subunit after substitution of a single amino acid in ArtA, while ArtB can form a functional complex with wild-type PltA and CdtB. We also found that, after addition of a single-terminal Cys residue, a CdtB homologue from cytolethal distending toxin can form a functional complex with ArtA and ArtB. In line with the broad host specificity of S. Typhimurium, we found that ArtB binds human glycans, terminated in N-acetylneuraminic acid, as well as glycans terminated in N-glycolylneuraminic acid (Neu5Gc), which are expressed in most other mammals14. The atomic structure of ArtB bound to its receptor shows the presence of an additional glycan-binding site, which broadens its binding specificity. Despite equivalent toxicity in vitro, we found that the ArtB/PltA/CdtB chimaeric toxin exhibits reduced lethality in an animal model, indicating that the host specialization of typhoid toxin has optimized its targeting mechanisms to the human host. This is a remarkable example of a toxin evolving to broaden its enzymatic activities and adapt to a specific host.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
01 November 2017
The original version of this Letter has been modified in the abstract and main text to better reflect the distribution of Neu5Ac sialoglycans in humans. Additionally, co-author Lingquan Deng’s present address has been further clarified.
References
Jackson, R., Johnson, L., Clarke, S. & Arnold, D. Bacterial pathogen evolution: breaking news. Trends Genet. 27, 32–40 (2011).
Donnenberg, M. S. & Whittam, T. S. Pathogenesis and evolution of virulence in enteropathogenic and enterohemorrhagic Escherichia coli. J. Clin. Invest. 107, 539–548 (2001).
Bentley, S. & Parkhill, J. Genomic perspectives on the evolution and spread of bacterial pathogens. Proc. Biol. Sci. 282, 20150488 (2015).
Alizon, S. & Michalakis, Y. Adaptive virulence evolution: the good old fitness-based approach. Trends Ecol. Evol. 30, 248–254 (2015).
Daugherty, M. & Malik, H. Rules of engagement: molecular insights from host–virus arms races. Annu. Rev. Genet. 46, 677–700 (2012).
Song, J., Gao, X. & Galan, J. E. Structure and function of the Salmonella Typhi chimaeric A2B5 typhoid toxin. Nature 499, 350–354 (2013).
Galán, J. Typhoid toxin provides a window into typhoid fever and the biology of Salmonella Typhi. Proc. Natl Acad. Sci. USA 113, 6338–6344 (2016).
Parry, C., Hien, T. T., Dougan, G., White, N. & Farrar, J. Typhoid fever. N. Engl. J. Med. 347, 1770–1782 (2002).
Crump, J. & Mintz, E. Global trends in typhoid and paratyphoid fever. Clin. Infect. Dis. 50, 241–246 (2010).
Raffatellu, M., Wilson, R., Winter, S. & Bäumler, A. Clinical pathogenesis of typhoid fever. J. Infect. Dev. Ctries 2, 260–266 (2008).
Wain, J., Hendriksen, R., Mikoleit, M., Keddy, K. & Ochiai, R. Typhoid fever. Lancet 385, 1136–1145 (2015).
Dougan, G. & Baker, S. Salmonella enterica serovar Typhi and the pathogenesis of typhoid fever. Annu. Rev. Microbiol. 68, 317–336 (2014).
Deng, L. et al. Host adaptation of a bacterial toxin from the human pathogen Salmonella Typhi. Cell 159, 1290–1299 (2014).
Varki, N., Strobert, E., Dick, E. J., Benirschke, K. & Varki, A. Biomedical differences between human and nonhuman hominids: potential roles for uniquely human aspects of sialic acid biology. Annu. Rev. Pathol. 6, 365–393 (2011).
Saitoh, M. et al. The artAB genes encode a putative ADP-ribosyltransferase toxin homologue associated with Salmonella enterica serovar Typhimurium DT104. Microbiology 151, 3089–3096 (2005).
Lara-Tejero, M. & Galan, J. E. Cytolethal distending toxin: limited damage as a strategy to modulate cellular functions. Trends Microbiol. 10, 147–152 (2002).
Desai, P. et al. Evolutionary genomics of Salmonella enterica subspecies. mBio 4, e00579-12 (2013).
Wang, C. et al. Complete genome sequence of Salmonella enterica subspecies arizonae str. RKS2983. Stand. Genomic Sci. 10, 30 (2015).
Byres, E. et al. Incorporation of a non-human glycan mediates human susceptibility to a bacterial toxin. Nature 456, 648–652 (2008).
Hedlund, M. et al. N-glycolylneuraminic acid deficiency in mice: implications for human biology and evolution. Mol. Cell. Biol. 27, 4340–4346 (2007).
Beddoe, T., Paton, A., Le Nours, J., Rossjohn, J. & Paton, J. Structure, biological functions and applications of the AB5 toxins. Trends Biochem. Sci. 35, 411–418 (2010).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
DeLano, W. L. The PyMOL molecular graphics system (Schrödinger, 2002); http://www.pymol.org
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Padler-Karavani, V. et al. Rapid evolution of binding specificities and expression patterns of inhibitory CD33-related Siglecs in primates. FASEB J. 28, 1280–1293 (2014).
Spano, S., Ugalde, J. E. & Galan, J. E. Delivery of a Salmonella Typhi exotoxin from a host intracellular compartment. Cell Host Microbe 3, 30–38 (2008).
Acknowledgements
The authors thank J. Wang for suggestions and for providing help with structure refinement, and the Galan Laboratory for careful review of the manuscript. G.S. is supported in part by a Postdoctoral Fellowship from the EMBO (ALTF 172-2015). Crystal screening was conducted at the Yale Macromolecular X-ray Core Facility (1S10OD018007-01). This work was supported by National Institutes of Health grants AI079022 (to J.E.G.) and GM32373 (to A.V.).
Author information
Authors and Affiliations
Contributions
X.G., G.S., L.D., A.V. and J.E.G. designed the research and analysed data. X.G., G.S. and L.D. performed the research. Y.N.-M., H.Y. and X.C. provided critical reagents. X.G. and J.E.G. wrote the manuscript with input from all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A correction to this article is available online at https://doi.org/10.1038/s41564-017-0070-x.
Electronic supplementary material
Supplementary Information
Supplementary Figures 1–29, Supplementary Table 6.
Supplementary Table 1
ArtB and PltB-binding to a customized sialoglycan microarray.
Supplementary Table 2
Analysis of fine ligand specificity of ArtB and PltB.
Supplementary Table 3
Binding of ArtB andmutant derivative to a customized sialoglycan microarray.
Supplementary Table 4
Analysis of the Neu5Acα6Gal/GalNAc-binding specificity of ArtB mutants.
Supplementary Table 5
Analysis of fine ligand specificity of ArtB and its mutants.
Rights and permissions
About this article
Cite this article
Gao, X., Deng, L., Stack, G. et al. Evolution of host adaptation in the Salmonella typhoid toxin. Nat Microbiol 2, 1592–1599 (2017). https://doi.org/10.1038/s41564-017-0033-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-017-0033-2
This article is cited by
-
The phylogenomics of CRISPR-Cas system and revelation of its features in Salmonella
Scientific Reports (2020)
-
Alternate subunit assembly diversifies the function of a bacterial toxin
Nature Communications (2019)
-
Cryo-EM structures of the pore-forming A subunit from the Yersinia entomophaga ABC toxin
Nature Communications (2019)
-
Decoding glycan recognition by bacterial toxins
Nature Microbiology (2018)