Abstract
The immune system faces the arduous task of defending the mucosal surfaces from invading pathogens, but must simultaneously repress responses against commensal organisms and other inert antigens that are abundant in the external environment, as inappropriate immune activation might expose the host to increased risk of autoimmunity. The behavior of individual immune cells is governed by the expression of transcription factors that are responsible for switching immune response genes on and off. T-bet (T-box expressed in T cells) has emerged as one of the key transcription factors responsible for controlling the fate of both innate and adaptive immune cells, and its expression in different immune cells found at mucosal surfaces is capable of dictating the critical balance between permitting robust host immunity and limiting susceptibility to autoimmunity and allergy.
Similar content being viewed by others
Introduction
Mucosal surfaces occupy vast anatomical areas, enabling them to perform a diverse range of specialized and complex functions, including gas exchange, reproduction, digestion, and absorption of nutrients. The immune system is charged with the challenging task of defending these extensive areas from infectious or neoplastic insults, yet at the same time must refrain from reacting to inert, bystander antigens that are abundant in the external environment, including inhaled aeroallergens, trillions of commensal microbes, as well as a dynamic load of food antigens. Inappropriate activation of the mucosal immune system by harmless environmental antigens wastes valuable resources and may result in collateral damage to host tissue with the potential to impair organ function. Consequently, mucosal immune responses are tightly regulated. The host must be capable of rapidly deploying aggressive, inflammatory responses in the event of pathogen exposure, while at the same time exercising restraint against nonpathogenic antigens by using regulatory mechanisms. The inflammatory and regulatory properties of mucosal immune cells are governed by cell-specific genetic programs, which are in turn controlled by transcription factors that dictate which immune response genes are switched on or off at any given moment.
T-bet (T-box expressed in T cells), encoded by the Tbx21 gene, has emerged as one of the key players governing the transcriptional control of mucosal immune responses. T-bet is a member of the T-box family of transcription factors and was first cloned in 2000.1 It has a critical role in regulating systemic, mucosal, and organ-specific immune responses, such that changes in T-bet expression can profoundly influence the balance between the development of autoimmunity or susceptibility to infection. T-bet-deficient mice are resistant to experimental models of autoimmunity, such as diabetes,2 encephalomyelitis,3 lupus4 and colitis,5 yet on the other hand, have heightened susceptibility to infections including mycobacteria,6 salmonella,7 leishmania,8 trypanosoma,9 and viruses.10, 11 Accordingly, T-bet must be rigorously regulated in a dynamic fashion to maintain the capacity to resist infection, without exposing the host to increased risk of autoimmunity, particularly at mucosal surfaces often in contact with commensal and pathogenic microorganisms.
T-bet expression is almost exclusively restricted to the immune system, and, to date, its expression has been detected in CD4+ T cells,1 CD8+ T cells1, B cells,1 monocytes,12 dendritic cells (DCs),12, 13 and natural killer (NK) cells,1 where it regulates cellular differentiation and the expression of cytokines, chemokines, cell surface receptors, and other important immune response proteins. Although most of the studies carried out so far have concentrated on the role of T-bet in T cells, its role in the innate immune system, as well as in coordinating the complex interplay between innate and adaptive immune components, may yet prove to be even more significant. It is especially intriguing that a single transcription factor seems to be a transcriptional activator in one cell type and a transcriptional repressor in another at the same genomic locus. The purpose of this review is to discuss T-bet-mediated transcriptional regulation of immune response genes involved in mucosal homeostasis, and how changes in T-bet expression and function can lead to the emergence of mucosal pathology (Table 1).
The Discovery of T-bet
The existence of distinct functional subsets of T helper cells, distinguished by patterns of cytokine secretion and cellular function, was first described in 1986.14 Th1 cells were characterized by interferon-γ (IFN-γ) production and Th2 cells produced interleukin 4 (IL-4) and IL-5. Subsequently, key roles for IL-12,15 STAT4,16, 17, 18, 19 and IFN-regulatory factor-1 (IRF-1)20, 21 in Th1 differentiation were described. However, the observation that STAT4−/− STAT6−/− double knockout T cells retained the ability to mount a polarized Th1 response, strongly hinted to the existence of an additional factor exerting transcriptional control over Th1 differentiation.22
Using a yeast one-hybrid screen, Szabo et al.1 identified this novel transcription factor as a protein bound to the IL-2 promoter in Th1 cells, but not in Th2. They identified and cloned a 62 kDa 530 amino-acid protein, containing a 186 amino acid DNA-binding domain homologous to members of the T-box family of transcription factors, and termed this new protein T-bet (Tbx21).
Gene Targets and Function
Forced expression of T-bet in naive T cells, or even in well-established Th2 cell lines, redirects these cells toward the Th1 lineage.1 The mechanisms whereby T-bet favors Th1 polarization are likely multifactorial, and includes direct transactivation of Th1-promoting genes, interaction with other transcription factors, epigenetic modification, and repression of Th2 differentiation.
In Th1 cells, T-bet binds at the promoter of more than 800 protein-encoding genes, which include cytokines, cytokine receptors, other transcription factors, and genes encoding proteins involved in cell metabolism, trafficking, and cellular differentiation.23 These gene targets are relatively well conserved between mouse and human.23
T-bet transactivates the Ifng gene through binding to multiple proximal and distal cis-acting enhancer elements.24, 25 Transactivation of other Th1-related genes, such TNF and IL18RAP, ensures that a coordinated and concerted Th1 response ensues.23 T-bet mediated negative regulation of GATA-3 has also been proposed as a potential mechanism responsible for T-bet-instigated Th1 differentiation.26 Baseline levels of GATA-3 are increased in T-bet−/− T cells, and retroviral overexpression of T-bet in Th2 cells reduces GATA-3 expression.26 Early inhibition of GATA-3 in naive T cells through blockade of the IL-4/STAT6 pathway, also culminates in Ifng promoter accessibility and induction of IFN-γ production, even in the absence of T-bet.26 T-bet binds to the promoter of RUNX1, the product of which binds to the IL-4 silencer and also directly represses GATA-3.23, 27, 28 T-bet also directly interacts with GATA-3 protein, interfering with its DNA binding.29 However, in contrast to the absence of T-bet seen in GATA-3+ Th2 cells, GATA-3 is both coexpressed with T-bet in T-bet+ Th1 cells and frequently binds to the same target genes in close proximity to T-bet.23 Interactions between T-bet and other transcription factors may also contribute to Th1 lineage specification. Recent data suggests that T-bet interacts with STAT4 at multiple Ifng distal enhancer sites and recruits RelA to the Ifng locus to facilitate efficient IFN-γ transcription.30
Mounting evidence is accruing regarding the role of T-bet in remodeling chromatin structure to transactivate or silence target genes. T-bet recruits histone 3 lysine 9 (H3K9) methyltransferase activity and induces lineage-specific, permissive H3K9 dimethyl modifications at the Ifng and Cxcr3 loci.31, 32, 33 T-bet-dependent permissive H3K9 methylation at Ifng is rapidly induced and sustained in Th1 differentiation.34 T-bet also physically interacts with the H3K27 demethylase, JMJD3.33 T-bet is functionally required and sufficient to alleviate the repressive H3K27 trimethylated state at Ifng and Cxcr3.33
T-bet also controls a transcriptional program determining lymphocyte homing to inflammatory sites by binding and activating numerous genes with critical functions in cell trafficking, including CXCR3, CCR5, ITGAL, and ICAM1.23, 35, 36 Indeed, the T-bet target genes CCL4 and CCRL2 are preferentially expressed in Th1 cells, whereas T-bet regulates the expression of CCL3 and CCL4 (RANTES) independent of IFN-γ.23 T-bet also controls expression of NKG7, a gene coding for a cell surface cytotoxic molecule preferentially expressed in polarized Th1 cells, independent of IFN-γ.23, 37
Th17 Cells
T-bet indirectly inhibits Th17 development through IFN-γ-mediated inhibition of IL-23R expression.38 Both T-bet−/− and STAT4−/− mice have decreased Th1 and increased Th17 cells.38, 39, 40 In T-bet−/− STAT6−/− double knockout mice, impairment in Th1 ontogeny can be partially compensated for by eomesodermin, the T-box family transcription factor that regulates IFN-γ production in Tc1 cells.41 These cells are highly sensitive to Th17 conversion by IL-6 and TGF-β.41 A complex interplay between T-bet and Th17 cells exists in disease models. Even though spontaneous asthma in T-bet−/− mice is a Th2-mediated disease, adoptively transferred T-bet−/− CD4+ cells preferentially convert to Th17, indicating the importance of T-bet in limiting Th17 ontogeny.42 In experimental autoimmune encephalitis, T-bet seems to be required for disease not only in Th1 cells but also in encephalitogenic Th17 cells.43 It is not certain whether T-bet confers pathogenic properties to Th17 cells, or whether these in vivo observations reflect the natural plasticity of the Th17 response, as T cells cultured in vitro under conditions favoring Th17 differentiation covert into Th1 cells in vivo.44, 45 The issue of T-cell Th1 and Th17 plasticity, and indeed pathogenicity, is especially intriguing in the context of intestinal inflammation, as adoptive transfer of T-bet−/− T cells fails to induce colitis,5 although it has yet to be established whether these cells differentiate into IL-17-secreting cells in the gut.
Regulatory T Cells
T-bet is also expressed in murine regulatory T cells (Tregs). Until recently, the function of T-bet in Tregs was unknown. Koch et al.46 proposed that T-bet expression in Tregs is required for an adequate Treg response to type 1 inflammation. It was observed that T-bet was upregulated in Tregs, in both experimental models of type 1 inflammation and Mycobacterial infection. Similar to Th1 cells, T-bet in Tregs was induced by Th1 conditions and required intact STAT1 and IFN-γR. However, “physiological” expression of T-bet in Tregs did not induce IFN-γ production, nor transform them into Th1 cells. Rather, T-bet+ Foxp3+ double-positive Tregs maintained their suppressive phenotype. Cxcr3 is a transcriptional target of T-bet in both mice35 and humans,23 allowing Th1 migration to inflammatory sites. Cxcr3 was induced by T-bet in Tregs, facilitating Treg migration to sites of type 1 inflammation. Genetic deficiency of T-bet also resulted in impaired Treg proliferation in response to type 1 inflammation. Indeed, T-bet was required to allow Foxp3+ Tregs to traffic to the lungs, draining lymph nodes and spleens of Mycobacterium tuberculosis-infected mice, and to control infection. T-bet was also required in adoptively transferred Tregs to control the Th1-mediated autoimmunity that occurred in Foxp3−/− scurfy mice. Although T-bet expression is functionally important in a subset of Foxp3+ Tregs, tolerogenic Foxp3− T-bet+ Tregs can also be induced from mucosal Th1 cells.47, 48 A population of tolerogenic T-bet+ Th1 cells has been identified in Toxoplasma gondii infection. Toxoplasma causes a highly polarized IL-12-dependent Th1 immune response, requiring IL-10 to prevent lethal immunopathology.49 Jankovic et al. found that the major CD4+ IL-10-producing population were IFN-γ+ T-bet+ Foxp3− Th1 cells.47 In addition to limiting a Th1-mediated immune response, these cells retain the ability to produce IFN-γ and promote efficient intracellular killing. IL-10 production by Th1 cells was inducible (on T-cell activation), transient, and limited to 10–15% of the Th1 population. Because TLR signaling is inhibitory to natural Treg activity, activated T-bet+ Th1 cells may provide a necessary source of IL-10 in Th1-mediated diseases such as T. gondii infection, that express potent TLR agonists.50
Tolerogenic mucosal T-bet+ CD4+ cells can also be induced by repeated intranasal administration of soluble peptide to myelin basic protein-specific TCR transgenic mice.48 Even though these former mucosal Th1 cells continue to express T-bet and some retain their capacity to produce IFN-γ, they exhibit both clonal anergy and IL-10-dependent suppression of DC cytokine secretion and effector cell proliferation. Both icos and the anergy-associated gene egr-2 are concomitantly upregulated with T-bet in this model. In the absence of inflammation, transformation to an antigen-specific regulatory phenotype may be durable, suggesting that T-bet can be functionally silenced by coexpression of other genes.
The Gastrointestinal Mucosal immune System
Given the central role of T-bet in the development and maintenance of Th1 cells, it is unsurprising that T-bet has a critical role in preclinical models of gastrointestinal inflammation that are reliant on pathological Th1 T cells. Experimental colitis secondary to adoptive transfer of naive T cells, trinitrobenzene sulfonate administration, and the spontaneous colitis developing in IL-10−/− mice, are all associated with increased expression of T-bet in lamina propria T cells.5 In the naive T-cell transfer model of colitis, recipients of T-bet−/− T cells do not develop disease, whereas recipients of T cells overexpressing T-bet develop accelerated disease.5 In contrast, Th2-mediated murine models of inflammatory bowel disease (IBD), such as oxazolone-induced colitis and the TCR-α−/−μ−/− spontaneous colitis model are not associated with increased T-bet expression in lamina propria T cells. Indeed, T-bet−/− mice develop more severe oxazolone-induced colitis, in association with increased expression of Th2 cytokines, including IL-4.5 In humans, elevated expression of T-bet has also been confirmed in lamina propria T cells from patients with intestinal inflammation, most notably in Crohn's disease5, 51, 52 and celiac disease,53 in which Th1 cells have been implicated as pathological effector cells.54, 55, 56, 57 Increased expression of T-bet has also been observed in peripheral blood T cells, B cells, and monocytes of untreated patients with celiac disease.58 After implementation of gluten avoidance and disease remission, T-bet levels were also observed to fall, implying that dynamic changes in T-bet expression may be a useful biomarker to track disease activity in celiac disease.
In Crohn's disease, T-cell stimulation through the T-cell receptor with anti-CD3 antibodies induces T-bet expression in lamina propria T cells.51, 52 In the case of celiac disease, in which the offending antigen is known, stimulation of duodenal T cells with specific peptide antigen also induces T-bet expression.59 In these human Th1 diseases, blockade of IL-2152, 59 or IFN-γ53 reduces the expression of T-bet in T cells, while antagonism of the antiinflammatory cytokine TGF-β induces T-bet expression,60 suggesting that other inflammatory and regulatory pathways converge with T-bet-mediated pathways to influence the prevailing inflammatory response. Antiinflammatory therapies, such as glucocorticoids, which are effective in the management of severe exacerbations of IBD, also inhibit T-bet expression in T cells.61 The activated glucocorticoid receptor physically interacts with T-bet in the nucleus preventing it from transactivating proinflammatory genes such as the IFN-γ gene.61 Inhibiting lamina propria T-cell activation in patients with IBD by blocking Ca2+ channels also reduces T-bet expression, with attendant reduction in T-cell IFN-γ production.62 These studies imply that specifically targeting T-bet expression in T cells may represent a novel therapeutic option in IBD.
Murine models have also provided valuable insight into the role of T-bet in the innate immune system in IBD. Loss of T-bet confined to the innate immune compartment in mice lacking an adaptive immune system results in the development of a spontaneous colitis closely resembling the clinical features of human ulcerative colitis (UC).63 Similar to the human disease, TRUC (T-bet × RAG UC) begins distally, before extending proximally with a predominantly superficial colonic inflammation. Histological appearances are comparable and long-term, unabated disease results in colonic dysplasia and malignancy.64 This model is dependent on the colonic microflora, as broad-spectrum antibiotics can prevent the onset of disease. The inflammatory process is characterized by overproduction of tumor necrosis factor-α (TNFα) in the lamina propria, which predominantly maps to CD11c+ class II+ myeloid cell populations, including DCs. Selective depletion of CD11c+ cells64 or TNFα blockade prevents disease,63 and TRUC mice lacking the p55 component of the TNFα receptor (triple knockout RAG−/− T-bet−/− TNF1/p55−/− mice)63 are resistant to disease. However, it should be noted that, although TNFα contributes to the diseased process in UC, as evidenced by the clinical efficacy of anti-TNFα therapies, other cytokine-driven pathways are also thought to operate in UC, such as IL-13 secreted by NK T cells.65
There are T-bet consensus sites upstream from the transcriptional start of the TNFα gene in both mouse and human DCs,63 and chromatin immunoprecipitation studies confirm that T-bet binds to the TNFα promoter.63 In contrast to its role in T cells, T-bet acts as a transcriptional repressor of the TNFα promoter in the RAW myeloid cell line, consistent with the observed deregulated overproduction of TNFα in T-bet−/− DCs.63 It is of interest that a single transcription factor acts as a transcriptional repressor of the TNFα gene in DCs and a transactivator of the same gene in Th1 T cells.23 Clearly, transcription factor programing of cellular behavior is complex, and in primary Th1 cells alone, T-bet binds at the promoter of more than 800 protein-coding genes.23 A number of genes identified as T-bet-binding targets by chromatin immunoprecipitation studies are at loci shared with single-nucleotide polymorphisms that are significantly associated with IBD or celiac disease in genome-wide association scans. For instance, significantly associated single-nucleotide polymorphisms at loci encoding TNF superfamily 4 and chemokine receptor 566, 67 are also genes specifically bound by T-bet in chromatin immunoprecipitation experiments.23, 68 The capacity for T-bet to influence the development and differentiation of Th17 cells offers another potentially relevant mechanism whereby this transcription factor might govern critical intestinal inflammatory pathways. The Th17 response is strongly suspected to have a crucial role in intestinal inflammation, as IL-17-secreting T cells have been identified in the lamina propria of patients with IBD and celiac disease,69, 70, 71, 72 and genome-wide association studies have identified significantly associated polymorphisms at numerous loci encoding key components of the Th17 pathway, including IL-12p40, STAT3, JAK2, CCR6, IRF-4, IL-21, as well as the IL-23 receptor.66, 67, 73 T cells from mice lacking T-bet also preferentially differentiate into IL-17-producing cells.38, 40, 41 However, in experimental T-cell-mediated intestinal inflammation, T-bet-deficient T cells fail to induce disease. It is not clear whether this is because of a failure of Th17 development in vivo, or whether the pathogenicity of Th17 cells relies on lineage plasticity, and switching to an IFN-γ-secreting Th1 phenotype, as has been observed in other models of autoimmunity.44, 45
T-bet is also involved in the homeostatic homing of mast cells to the small intestine.74T-bet−/− mice have reduced numbers of intestinal mast cells and their progenitors. Mast cells from T-bet-null hosts have diminished binding to VCAM-1 and MadCam-1, whose expression on small intestinal vascular endothelium is responsible for mast cell recruitment. Although T-bet expression has not been detected in mast cells at any stage during ontogeny, adoptive transfer of T-bet-sufficient DCs restores normal mast cell homing to the small intestine, implying that T-bet indirectly influences mast cell trafficking to the gut through its actions in DCs. It is not clear whether T-bet influences the homing of mast cells to the gut in man; however, the mechanisms responsible for governing mast cell trafficking to the gut are of considerable clinical interest, as various common gastrointestinal diseases, including food allergy,75 irritable bowel syndrome,76 and postoperative ileus,77 are all associated with disordered intestinal mast cell homeostasis.
Periodontitis
The mouth, in common with other mucosal surfaces of the gastrointestinal tract, is home to hundreds of species of mucosal bacteria living symbiotically with the mucosal immune system.78 Bacteria accumulate in complex biofilms on dental surfaces. These biofilms may progress to form plaques, which cause tissue inflammation either above or below the gum line.
Gingivitis is characterized by redness and swelling of the gingival margins with contact bleeding that is reversible with effective dental hygiene.79 Periodontitis, in contrast, is characterized by much deeper tissue inflammation below the gum line, causing destruction of the periodontal connective tissue and related alveolar bone. Destruction of the dental support structures may lead to receding gums, loosening of teeth, and eventual tooth loss. Tissue inflammation in periodontitis is initially characterized by neutrophil infiltration into the gingival sulcus. Periodontipathic bacteria such as Porphyromonas gingivalis have evolved strategies such as C3b complement cleavage to avoid neutrophil-mediated clearance from plaques. Subsequent T cell and macrophage infiltration occurs, with later B and plasma cell infiltration. This immune response either results in control of infection or chronic inflammation with tissue destruction.80 There is considerable controversy as to whether type 1 or type 2 inflammation predominates in periodontitis. This may relate to clinical and experimental heterogeneity in the literature, reviewed comprehensively by Gemmell et al.80 Górska et al.81 evaluated cytokine levels from gingival tissue supernatants and found that 60% of patients with chronic periodontitis had simultaneously high levels of IL-1β, IL-2, IFN-γ, and TNFα, with low or undetectable IL-4 and IL-10. Diseased patients had approximately fivefold greater tissue IFN-γ and TNFα levels compared with controls. Levels of both IL-4 and IL-10 were more frequently detectable in healthy controls (88 and 72%, respectively) compared with periodontitis patients (35 and 45%, respectively). Dutzan et al.82 compared IFN-γ and T-bet expression in gingival biopsies between active and inactive sites in patients with chronic progressive periodontitis. IFNG and TBX21 mRNA were 2.7- and 1.4-fold increased at active disease sites compared with inactive sites. TBX21 mRNA is also highly expressed in periodontitis-associated granulomas.83 IFN-γ+ Th1 cells are associated with alveolar bone loss in periodontitis.84 RANKL (receptor activator of nuclear factor-kappa B ligand) expression is induced by IFN-γ in Th1 cells. Indeed, coexpression of RANKL and TBX21 mRNA has been observed in periodontitis-associated granulomas.83 Enhanced alveolar bone reabsorption is associated with the presence of IFN-γ+ and RANKL+ double-positive CD4+ cells in both mice and humans,85 linking type 1 inflammation to bone loss in active disease.
Oral Graft-Versus-Host Disease
Acute graft-versus-host disease (aGvHD) after allogeneic hematopoietic stem cell transplantation is recognized as a type 1 cytokine-driven, CD8+ T cell-mediated disease, which can be abrogated by skewing donor cells to a type 2 cytokine profile.86 In contrast, chronic GvHD (cGvHD) has been variously characterized as a Th1, Th2, or Th17-mediated disease, depending on the model and species investigated.87, 88, 89 However, a recently published carefully controlled case series implicated T-bet and type 1 inflammation in the pathogenesis of oral cGvHD.90 The subepithelial infiltrate seen in patients with oral cGvHD was predominantly composed of memory T-bet+ CD8+ CD45RO+ T cells, with minor populations of T-bet+ CD4+ T cells, and CD68+ macrophage–DCs. The number of infiltrating T-bet+ CD8+ cells correlated both with keratinocyte apoptosis and clinical disease severity.90 Nearly all infiltrating T cells expressed CXCR3, whose expression has been shown to be T-bet dependent.35 In addition to its potential role in CD8+ T-cell trafficking, T-bet is also required for efficient production of IFN-γ and the cytotoxins perforin and granzyme B.91, 92 However, in certain circumstances, the related T-box family transcription factor eomesodermin can compensate for the absence of T-bet in CD8+ T cells to restore IFN-γ production and cytolytic effector function.41, 93 Similar to their CD4+ counterparts, CD8+ T cells deficient in T-bet, or indeed both T-bet and eomesodermin, preferentially secrete IL-17.41, 93, 90, 35
In murine models of aGvHD, donor-derived IL-15 is required for type 1 memory CD8+ T cell polarization, IFN-γ production, and Cxcr3 expression.94 Tissue levels of IL-15 transcripts correlated with both disease severity and Tbx21 mRNA expression. IL-15 was likely driven by type 1 IFNs produced by infiltrating plasmacytoid DCs, both of which were enhanced in severe disease compared with nondiseased controls. This implies a key role for type 1 immunity in oral cGvHD and alters the paradigm for this disease in man.
Respiratory Mucosal Inflammation
Allergic inflammation of the airways is responsible for common clinical manifestations such as asthma and hayfever. It is characterized by eosinophilic inflammation, allergen-reactive immunoglobulin E antibodies, and mast cell degranulation, orchestrated by T cells secreting Th2 cytokines, including IL-4, IL-5, and IL-13.95, 96 Although, GATA-3 is an important transcriptional regulator of Th2 differentiation, the interaction between T-bet and GATA-3 is a key event in determining the fate of T-cell lineage commitment. T-bet physically interacts with GATA-3, preventing it from binding to its transcriptional targets and repressing Th2 differentiation.29 Retroviral transduction of T-bet into established Th2 cells also redirects these cells back toward the Th1 lineage.1 In addition to direct physical contact between these transcription factors, T-bet and GATA-3 also bind to a shared profile of target genes that are differentially expressed in Th1 and Th2 cells.23 Accordingly, both T-bet and GATA-3 are capable of influencing allergic inflammatory responses at mucosal surfaces. Mice lacking T-bet spontaneously develop airway changes resembling human asthma with an eosinophil and T-cell-rich inflammatory infiltrate, overexpression of Th2 cytokines, increased peribronchial collagen deposition suggestive of airway remodeling, and bronchial hyper-responsiveness.97 Restoration of T-bet in T cells reverses these changes.98 Similar changes have been observed in antigen-specific models of asthma, in which loss of T-bet in antigen-specific T cells results in enhanced disease susceptibility; however, this phenotype is dependent on STAT6 signaling, indicating that a functional Th2 response is required, rather than merely the absence of a Th1 response.42
Animal models of asthma also highlight the importance of T-bet-mediated regulation, beyond its key role in helper T cells. Although T-bet−/− mice have reduced absolute numbers of invariant NK T cells,99 these cells overproduce Th2 cytokines such as IL-4 and IL-13, with little detectable IFN-γ.100 In this model, blockade of CD1d also reverses airway hyper-responsiveness.100 T-bet is also inducible in B cells, which is associated with impaired immunoglobulin class switching toward proallergic immunoglobulin E antibodies.101 T-bet expression in DCs might also potentially influence T-cell polarization in the airways. Sensitized mice receiving antigen-pulsed DCs intranasally are unable to redirect proallergic Th2 airway responses toward Th1 responses in the absence of T-bet,102 consistent with other data showing that T-bet-deficient DCs are less capable of inducing Th1 responses in vivo compared with T-bet-sufficient DCs.103
In patients with asthma, T-bet expression is reduced in T cells from the airways97 and peripheral blood.104 Genetic analyses have also identified polymorphisms in the TBX21 gene that are associated with an increased risk of asthma and airway hyper-responsiveness.105, 106 Asthmatic children with a nonsynonymous replacement of histidine 33 with glutamine in the functional T-bet protein molecule are more responsive to inhaled corticosteroid therapy compared with patients without the polymorphism.107 Murine T cells, with a homologous mutation to this human genetic variant, have defective IFN-γ production and instead favor Th2 cytokine responses.107
Mucosal Neoplasia
At mucosal surfaces, unchecked inflammation predisposes to cancer, usually through induction of dysplastic change as a precursor to malignant transformation. Colorectal cancer (CRC) occurring in the context of long-standing colonic inflammation in UC is a well-documented example of this phenomenon.
In addition to sharing a number of important clinical features with UC, TRUC mice also develop colonic epithelial dysplasia and adenocarcinoma, due to long-standing uncontrolled colonic inflammation.64 Carcinogenesis in the TRUC model recapitulates many phenotypic and molecular features of carcinogenesis in UC, offering a potentially important preclinical model to investigate carcinogenesis in human IBD. By 3 months of age, half the TRUC colony have dysplasia, rising to 96% by 6 months of age. By that time, 42% have frank adenocarcinoma. Similar to carcinogenesis in UC, carcinoma in the TRUC mouse, arising from mucosa with the longest-standing inflammation (the rectum), is typically flat and arises from areas of dysplasia. The TRUC epithelium is rich in reactive oxygen species and DNA adducts, the levels of both of which correlate with duration of colonic inflammation. The molecular pathogenesis of TRUC CRC also more closely resembles UC-associated CRC than sporadic CRC, with early loss of function of p53, early increased epithelial COX-2 expression, and later APC mutations, leading to abnormal β-catenin localization. Furthermore, unlike the cytokine signature observed in early TRUC colitis, which is dominated by TNFα expression, the inflammatory microenvironment associated with dysplasia and adenocarcinoma is characterized by increased expression of other proinflammatory cytokines, such as IL-6, similar to the situation observed in UC-associated CRC.64, 108
TNFα is known to drive tumor metastasis.109 Targeted overexpression of T-bet in DCs in the TRUC mouse model represses DC TNF-α production, ameliorates colitis, and reduces colonic neoplasia.64 This raises the intriguing possibility that T-bet-mediated suppression of DC TNF-α production may have a role in preventing metastasis in T-bet+ CRC.
T-bet may also be a useful marker for a vigorous antitumor immune response and favorable prognosis in CRC. In both sporadic CRC and CRC with “microsatellite instability”, an acquired defect in DNA mismatch repair, an inflammatory tumor infiltrate, is associated with improved survival.110, 111 Pagès et al. used a library of 959 CRC specimens to compare inflammatory cell infiltrates in CRCs without metastasis (good prognosis) and CRCs with signs of early metastasis (poor prognosis).112 A heavy cytotoxic T-cell infiltrate, 4.9-fold greater in tumors without metastasis, conferred survival advantage. Tumors without metastasis also had higher expression of markers of both activated cytotoxic T cells (granzyme B and granulysin) and type 1 inflammation including T-bet, IRF-1, and IFN-γ. However, this antitumor effect is not uniquely attributable to cytotoxic T cells, as CD3+ CD4+ cells were also overrepresented in tumors without metastasis.
T-bet also has a role in promoting a vigorous antitumor response and preventing metastasis at other mucosal surfaces. Cervical carcinoma in situ (CIN) and cancer are associated with cervical human papilloma virus (HPV) infection in humans.113 The progression from HPV-16-associated CIN to cancer is associated with a switch in T helper response against HPV-16 epitopes from predominantly Th1 to Th2.114 In another study, although peripheral blood CD4+ T cells from patients with HPV-18-associated CIN produced a mixed Th1/Th2 response to HPV-16 E6 epitopes, greater IFN-γ production was associated with less disease relapse and less persistence of infection after surgery.115 In addition, the intensity of CIN-infiltrating CD4+ and T-bet+ cells was also associated with a favorable clinical outcome.
The T-bet-associated antitumor response is not limited to mucosal surfaces, but also occurs in other solid-organ tumors. In experimental murine prostate cancer, T-bet prevents tumor metastasis, but not carcinogenesis, per se.116 Finally, in the murine B16F10 model of metastatic melanoma, T-bet-deficient animals were susceptible to tumor development and metastasis.117 T-cell-specific doxycycline-induced expression of T-bet restored an antitumor immune response and reduced metastasis, as did ectopic expression of T-bet in T-bet−/− Th1 cells. However, the antitumor effect of T-bet in this model is not limited to T cells. T-bet−/− in NK cells impairs both NK survival and melanoma cell killing.118 Adoptive transfer of T-bet-sufficient NK cells restores an antitumor response in T-bet−/− Rag2−/− double knockout animals, confirming that the cellular requirement for T-bet to produce a satisfactory antitumor response is not limited to T cells.
Stoicov et al.119 recently described a novel role for T-bet in the pathogenesis of Helicobacter-associated gastric cancer. H. pylori infection causes gastric mucosal inflammation, the degree and polarization of which is influenced by cytokine gene polymorphisms in humans, characterized by high expression of TNF-α, IL-1β, and low IL-10.120, 121 In murine H. felis infection, genotype has a prominent role, as gastric adenocarcinoma is associated with a mucosal Th1 response, whereas a Th2 response is protective.122 In response to H. felis infection, T-bet-sufficient mice develop mucosal inflammation with parietal cell loss and substantial TNF-α and IL-1β production with reduced IL-10 production, recapitulating the cytokine environment seen in humans who are susceptible to the carcinogenic effects of H. pylori.119, 120 T-bet-sufficient H. felis-infected mice later develop gastric mucosal dysplasia and carcinoma. T-bet−/− mice are protected from H. felis-induced gastritis and carcinoma. Although it is intriguing to think that T-bet may have a central role in the mucosal immune response to Helicobacter infection, polymorphisms in up- and downstream targets remain to be defined as potential risks for gastric adenocarcinoma.
Conclusion
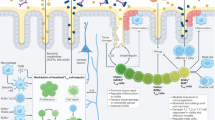
The mucosal surfaces represent the most hostile landscape that the immune system is faced with policing, and consequently complex and dynamic immune pathways exist to counter the threat of pathogen invasion, prevent the development of autoimmunity, and to maintain mucosal homeostasis, such that these highly specialized tissues are able to perform their physiological functions unhindered. The behavior of the mucosal immune system is governed by the profile of immune response proteins that are expressed at any given moment. Although the control of immune response gene expression is complex, transcription factors acting in a tissue-specific manner have a central role in this process. T-bet has emerged as one of the most important transcription factors responsible for regulating mucosal immunity. It is expressed in cells of both the adaptive and innate immune compartments, in which it participates in governing numerous immune response pathways (Figure 1). The critical role of T-bet in influencing T cell lineage commitment is particularly important at the mucosal surfaces, in which the pattern (Th1, Th2, or Th17), magnitude, and efficiency of the prevailing T-cell responses determines host susceptibility to infection, autoimmunity, allergy, or malignant change. In autoimmune diseases, therapeutic targeting of individual proinflammatory molecules, such as TNFα or IL-12p40, has provided important lessons in how improved understanding of inflammatory pathways might lead to more specific therapies. Targeting specific transcription factors might offer an even more attractive rationale for combating these diseases, as they are frequently upstream of effector cytokines, and typically target hundreds of related immune response proteins acting on the same pathway.
T-box expressed in T cells (T-bet) acts at multiple levels to regulate mucosal immune responses. T-bet is expressed in cells of both the innate and adaptive arms of the immune system, in which it regulates mucosal immune responses. (1) Loss of T-bet in CD11c+ dendritic cells (DCs) results in deregulated overexpression of tumor necrosis factor-α (TNFα), which results in colonic epithelial apoptosis and the development of spontaneous colitis. T-bet expression in DCs is also required to recruit mast cells to gastrointestinal mucosa. (2) The development and maintenance of Th1 cells is dependent on T-bet. Th1 cells support immunity to intracellular organisms (for example, mycobacteria) through activation of macrophages. (3) In the absence of T-bet, Th17 T cells preferentially expand. Th17 cells are implicated in neutrophil accumulation and immunity to extracellular bacteria, as well as in the pathogenesis of inflammatory bowel disease. (4) T-bet has been implicated in the development of regulatory T cells (Treg) that are responsible for limiting Th1 responses. (5) In the absence of T-bet, Th2 cells expand in the airways, resulting in increased expression of interleukin 4 (IL-4), IL-13 (promoting immunoglobulin (Ig) class switching to proallergic IgE antibodies and activating mast cells), and IL-5 (resulting in expansion and activation of mucosal eosinophils). Eosinophilic inflammation is implicated in immunity to helminth infection and also in the pathogenesis of allergic asthma and rhinitis. (6) T-bet is inducible in B cells, leading to impairment of IgE class switching. Cross linking of allergen-specific IgE on mast cells activates mucosal mast cells. (7) Absence of T-bet leads to overexpression of IL-13 by mucosal natural killer (NK) T cells. Ag, antigen; ECP, eosinophil cationic protein; IFN, interferon; MBP, major basic protein; MPO, myeloperoxidase.
References
Szabo, S.J., Kim, S.T., Costa, G.L., Zhang, X., Fathman, C.G. & Glimcher, L.H. A novel transcription factor, T-bet, directs Th1 lineage commitment. Cell 100, 655–669 (2000).
Esensten, J.H., Lee, M.R., Glimcher, L.H. & Bluestone, J.A. T-bet-deficient NOD mice are protected from diabetes due to defects in both T cell and innate immune system function. J. Immunol. 183, 75–82 (2009).
Bettelli, E., Sullivan, B., Szabo, S.J., Sobel, R.A., Glimcher, L.H. & Kuchroo, V.K. Loss of T-bet, but not STAT1, prevents the development of experimental autoimmune encephalomyelitis. J. Exp. Med. 200, 79–87 (2004).
Peng, S.L., Szabo, S.J. & Glimcher, L.H. T-bet regulates IgG class switching and pathogenic autoantibody production. Proc. Natl Acad. Sci. USA 99, 5545–5550 (2002).
Neurath, M.F. et al. The transcription factor T-bet regulates mucosal T cell activation in experimental colitis and Crohn's disease. J. Exp. Med. 195, 1129–1143 (2002).
Sullivan, B.M. et al. Increased susceptibility of mice lacking T-bet to infection with Mycobacterium tuberculosis correlates with increased IL-10 and decreased IFN-gamma production. J. Immunol. 175, 4593–4602 (2005).
Ravindran, R., Foley, J., Stoklasek, T., Glimcher, L.H. & McSorley, S.J. Expression of T-bet by CD4T cells is essential for resistance to Salmonella infection. J. Immunol. 175, 4603–4610 (2005).
Cobb, D., Guo, S., Lara, A.M., Manque, P., Buck, G. & Smeltz, R.B. T-bet-dependent regulation of CD8+ T-cell expansion during experimental Trypanosoma cruzi infection. Immunology 128, 589–599 (2009).
Rosas, L.E. et al. Cutting edge: STAT1 and T-bet play distinct roles in determining outcome of visceral leishmaniasis caused by Leishmania donovani. J. Immunol. 177, 22–25 (2006).
Matsui, M., Moriya, O., Yoshimoto, T. & Akatsuka, T. T-bet is required for protection against vaccinia virus infection. J. Virol. 79, 12798–12806 (2005).
Svensson, A., Nordstrom, I., Sun, J.B. & Eriksson, K. Protective immunity to genital herpes simplex [correction of simpex] virus type 2 infection is mediated by T-bet. J. Immunol. 174, 6266–6273 (2005).
Lighvani, A.A. et al. T-bet is rapidly induced by interferon-gamma in lymphoid and myeloid cells. Proc. Natl Acad. Sci. USA 98, 15137–15142 (2001).
Lugo-Villarino, G., Maldonado-Lopez, R., Possemato, R., Penaranda, C. & Glimcher, L.H. T-bet is required for optimal production of IFN-gamma and antigen-specific T cell activation by dendritic cells. Proc. Natl Acad. Sci. USA 100, 7749–7754 (2003).
Mosmann, T.R., Cherwinski, H., Bond, M.W., Giedlin, M.A. & Coffman, R.L. Two types of murine helper T cell clone. I. Definition according to profiles of lymphokine activities and secreted proteins. J. Immunol. 136, 2348–2357 (1986).
Hsieh, C.S., Macatonia, S.E., Tripp, C.S., Wolf, S.F., O'Garra, A. & Murphy, K.M. Development of TH1 CD4+ T cells through IL-12 produced by Listeria-induced macrophages. Science 260, 547–549 (1993).
Thierfelder, W.E. et al. Requirement for Stat4 in interleukin-12-mediated responses of natural killer and T cells. Nature 382, 171–174 (1996).
Kaplan, M.H., Sun, Y.L., Hoey, T. & Grusby, M.J. Impaired IL-12 responses and enhanced development of Th2 cells in Stat4-deficient mice. Nature 382, 174–177 (1996).
Galon, J., Sudarshan, C., Ito, S., Finbloom, D. & O'Shea, J.J. IL-12 induces IFN regulating factor-1 (IRF-1) gene expression in human NK and T cells. J. Immunol. 162, 7256–7262 (1999).
Coccia, E.M., Passini, N., Battistini, A., Pini, C., Sinigaglia, F. & Rogge, L. Interleukin-12 induces expression of interferon regulatory factor-1 via signal transducer and activator of transcription-4 in human T helper type 1 cells. J. Biol. Chem. 274, 6698–6703 (1999).
Taki, S. et al. Multistage regulation of Th1-type immune responses by the transcription factor IRF-1. Immunity 6, 673–679 (1997).
Lohoff, M. et al. Interferon regulatory factor-1 is required for a T helper 1 immune response in vivo. Immunity 6, 681–689 (1997).
Kaplan, M.H., Wurster, A.L. & Grusby, M.J. A signal transducer and activator of transcription (Stat)4-independent pathway for the development of T helper type 1 cells. J. Exp. Med. 188, 1191–1196 (1998).
Jenner, R.G. et al. The transcription factors T-bet and GATA-3 control alternative pathways of T-cell differentiation through a shared set of target genes. Proc. Natl Acad. Sci. USA 106, 17876–17881 (2009).
Cho, J.Y., Grigura, V., Murphy, T.L. & Murphy, K. Identification of cooperative monomeric Brachyury sites conferring T-bet responsiveness to the proximal IFN-gamma promoter. Int. Immunol. 15, 1149–1160 (2003).
Lee, D.U., Avni, O., Chen, L. & Rao, A. A distal enhancer in the interferon-gamma (IFN-gamma) locus revealed by genome sequence comparison. J. Biol. Chem. 279, 4802–4810 (2004).
Usui, T. et al. T-bet regulates Th1 responses through essential effects on GATA-3 function rather than on IFNG gene acetylation and transcription. J. Exp. Med. 203, 755–766 (2006).
Komine, O. et al. The Runx1 transcription factor inhibits the differentiation of naive CD4+ T cells into the Th2 lineage by repressing GATA3 expression. J. Exp. Med. 198, 51–61 (2003).
Naoe, Y. et al. Repression of interleukin-4 in T helper type 1 cells by Runx/Cbf beta binding to the Il4 silencer. J. Exp. Med. 204, 1749–1755 (2007).
Hwang, E.S., Szabo, S.J., Schwartzberg, P.L. & Glimcher, L.H. T helper cell fate specified by kinase-mediated interaction of T-bet with GATA-3. Science 307, 430–433 (2005).
Balasubramani, A., Shibata, Y., Crawford, G.E., Baldwin, A.S., Hatton, R.D. & Weaver, C.T. Modular utilization of distal cis-regulatory elements controls Ifng gene expression in T cells activated by distinct stimuli. Immunity 33, 35–47 (2010).
Fields, P.E., Kim, S.T. & Flavell, R.A. Cutting edge: changes in histone acetylation at the IL-4 and IFN-gamma loci accompany Th1/Th2 differentiation. J. Immunol. 169, 647–650 (2002).
Lewis, M.D., Miller, S.A., Miazgowicz, M.M., Beima, K.M. & Weinmann, A.S. T-bet's ability to regulate individual target genes requires the conserved T-box domain to recruit histone methyltransferase activity and a separate family member-specific transactivation domain. Mol. Cell. Biol. 27, 8510–8521 (2007).
Miller, S.A., Huang, A.C., Miazgowicz, M.M., Brassil, M.M. & Weinmann, A.S. Coordinated but physically separable interaction with H3K27-demethylase and H3K4-methyltransferase activities are required for T-box protein-mediated activation of developmental gene expression. Genes. Dev. 22, 2980–2993 (2008).
Chang, S. & Aune, T.M. Dynamic changes in histone-methylation “marks” across the locus encoding interferon-gamma during the differentiation of T helper type 2 cells. Nat. Immunol. 8, 723–731 (2007).
Lord, G.M. et al. T-bet is required for optimal proinflammatory CD4+ T-cell trafficking. Blood 106, 3432–3439 (2005).
Matsuda, J.L., George, T.C., Hagman, J. & Gapin, L. Temporal dissection of T-bet functions. J. Immunol. 178, 3457–3465 (2007).
Lund, R., Ahlfors, H., Kainonen, E., Lahesmaa, A.M., Dixon, C. & Lahesmaa, R. Identification of genes involved in the initiation of human Th1 or Th2 cell commitment. Eur. J. Immunol. 35, 3307–3319 (2005).
Harrington, L.E. et al. Interleukin 17-producing CD4(+) effector T cells develop via a lineage distinct from the T helper type 1 and 2 lineages. Nat. Immunol. 6, 1123–1132 (2005).
Park, H. et al. A distinct lineage of CD4T cells regulates tissue inflammation by producing interleukin 17. Nat. Immunol. 6, 1133–1141 (2005).
Durrant, D.M., Gaffen, S.L., Riesenfeld, E.P., Irvin, C.G. & Metzger, D.W. Development of allergen-induced airway inflammation in the absence of T-bet regulation is dependent on IL-17. J. Immunol. 183, 5293–5300 (2009).
Yang, Y., Xu, J.N., Niu, Y.Y., Bromberg, J.S. & Ding, Y.Z. T-bet and eomesodermin play critical roles in directing T cell differentiation to Th1 versus Th17. J. Immunol. 181, 8700–8710 (2008).
Fujiwara, M. et al. T-bet inhibits both TH2 cell-mediated eosinophil recruitment and TH17 cell-mediated neutrophil recruitment into the airways. J. Allergy Clin. Immunol. 119, 662–670 (2007).
Yang, Y. et al. T-bet is essential for encephalitogenicity of both Th1 and Th17 cells. J. Exp. Med. 206, 1549–1564 (2009).
Bending, D. et al. Highly purified Th17 cells from BDC2.5NOD mice convert into Th1-like cells in NOD/SCID recipient mice. J. Clin. Invest. 119, 565–572 (2009).
Lee, Y.K. et al. Late developmental plasticity in the T helper 17 lineage. Immunity 30, 92–107 (2009).
Koch, M.A., Tucker-Heard, G., Perdue, N.R., Killebrew, J.R., Urdahl, K.B. & Campbell, D.J. The transcription factor T-bet controls regulatory T cell homeostasis and function during type 1 inflammation. Nat. Immunol. 10, 595–602 (2009).
Jankovic, D. et al. Conventional T-bet(+)Foxp3(−) Th1 cells are the major source of host-protective regulatory IL-10 during intracellular protozoan infection. J. Exp. Med. 204, 273–283 (2007).
Gabrysova, L. et al. Negative feedback control of the autoimmune response through antigen-induced differentiation of IL-10-secreting Th1 cells. J. Exp. Med. 206, 1755–1767 (2009).
Gazzinelli, R.T. et al. In the absence of endogenous IL-10, mice acutely infected with Toxoplasma gondii succumb to a lethal immune response dependent on CD4+ T cells and accompanied by overproduction of IL-12, IFN-gamma and TNF-alpha. J. Immunol. 157, 798–805 (1996).
Jankovic, D., Kugler, D.G. & Sher, A. IL-10 production by CD4+ effector T cells: a mechanism for self-regulation. Mucosal Immunol. 3, 239–246 (2010).
Matsuoka, K. et al. T-bet upregulation and subsequent interleukin 12 stimulation are essential for induction of Th1 mediated immunopathology in Crohn's disease. Gut 53, 1303–1308 (2004).
Monteleone, G., Monteleone, I., Fina, D., Vavassori, P., Del Vecchio Blanco, G. & Caruso, R. et al. Interleukin-21 enhances T-helper cell type I signaling and interferon-gamma production in Crohn's disease. Gastroenterology 128, 687–694 (2005).
Monteleone, I. et al. Regulation of the T helper cell type 1 transcription factor T-bet in coeliac disease mucosa. Gut 53, 1090–1095 (2004).
Fuss, I.J. et al. Disparate CD4+ lamina propria (LP) lymphokine secretion profiles in inflammatory bowel disease. Crohn's disease LP cells manifest increased secretion of IFN-gamma, whereas ulcerative colitis LP cells manifest increased secretion of IL-5. J. Immunol. 157, 1261–1270 (1996).
Parronchi, P. et al. Type 1 T-helper cell predominance and interleukin-12 expression in the gut of patients with Crohn's disease. Am. J. Pathol. 150, 823–832 (1997).
Nilsen, E.M., Lundin, K.E., Krajci, P., Scott, H., Sollid, L.M. & Brandtzaeg, P. Gluten specific HLA-DQ restricted T cells from coeliac mucosa produce cytokines with Th1 or Th0 profile dominated by interferon gamma. Gut 37, 766–776 (1995).
Nilsen, E.M. et al. Gluten induces an intestinal cytokine response strongly dominated by interferon gamma in patients with celiac disease. Gastroenterology 115, 551–563 (1998).
Frisullo, G. et al. T-bet and pSTAT-1 expression in PBMC from coeliac disease patients: new markers of disease activity. Clin. Exp. Immunol. 158, 106–114 (2009).
Fina, D. et al. Interleukin 21 contributes to the mucosal T helper cell type 1 response in coeliac disease. Gut 57, 887–892 (2008).
Di Sabatino, A. et al. Blockade of transforming growth factor beta upregulates T-box transcription factor T-bet, and increases T helper cell type 1 cytokine and matrix metalloproteinase-3 production in the human gut mucosa. Gut 57, 605–612 (2008).
Liberman, A.C. et al. The activated glucocorticoid receptor inhibits the transcription factor T-bet by direct protein-protein interaction. FASEB J. 21, 1177–1188 (2007).
Di Sabatino, A. et al. Targeting gut T cell Ca2+ release-activated Ca2+ channels inhibits T cell cytokine production and T-box transcription factor T-bet in inflammatory bowel disease. J. Immunol. 183, 3454–3462 (2009).
Garrett, W.S. et al. Communicable ulcerative colitis induced by T-bet deficiency in the innate immune system. Cell 131, 33–45 (2007).
Garrett, W.S. et al. Colitis-associated colorectal cancer driven by T-bet deficiency in dendritic cells. Cancer Cell 16, 208–219 (2009).
Fuss, I.J. et al. Nonclassical CD1d-restricted NK T cells that produce IL-13 characterize an atypical Th2 response in ulcerative colitis. J. Clin. Invest. 113, 1490–1497 (2004).
Dubois, P.C. et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 42, 295–302 (2010).
Barrett, J.C. et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn's disease. Nat. Genet. 40, 955–962 (2008).
Gocke, A.R. et al. T-bet regulates the fate of Th1 and Th17 lymphocytes in autoimmunity. J. Immunol. 178, 1341–1348 (2007).
Rovedatti, L. et al. Differential regulation of interleukin 17 and interferon gamma production in inflammatory bowel disease. Gut 58, 1629–1636 (2009).
Kleinschek, M.A. et al. Circulating and gut-resident human Th17 cells express CD161 and promote intestinal inflammation. J. Exp. Med. 206, 525–534 (2009).
Kobayashi, T. et al. IL23 differentially regulates the Th1/Th17 balance in ulcerative colitis and Crohn's disease. Gut 57, 1682–1689 (2008).
Monteleone, I. et al. Characterization of IL-17A-producing cells in celiac disease mucosa. J. Immunol. 184, 2211–2218 (2010).
Franke, A. et al. Sequence variants in IL10, ARPC2 and multiple other loci contribute to ulcerative colitis susceptibility. Nat. Genet. 40, 1319–1323 (2008).
Alcaide, P. et al. Dendritic cell expression of the transcription factor T-bet regulates mast cell progenitor homing to mucosal tissue. J. Exp. Med. 204, 431–439 (2007).
Berin, M.C. & Mayer, L. Immunophysiology of experimental food allergy. Mucosal Immunol. 2, 24–32 (2009).
Ohman, L. & Simren, M. Pathogenesis of IBS: role of inflammation, immunity and neuroimmune interactions. Nat. Rev. Gastroenterol. Hepatol. 7, 163–173 (2010).
Boeckxstaens, G.E. & de Jonge, W.J. Neuroimmune mechanisms in postoperative ileus. Gut 58, 1300–1311 (2009).
Pihlstrom, B.L., Michalowicz, B.S. & Johnson, N.W. Periodontal diseases. Lancet 366, 1809–1820 (2005).
Coventry, J., Griffiths, G., Scully, C. & Tonetti, M. ABC of oral health: periodontal disease. BMJ 321, 36–39 (2000).
Gemmell, E., Yamazaki, K. & Seymour, G.J. Destructive periodontitis lesions are determined by the nature of the lymphocytic response. Crit. Rev. Oral Biol. Med. 13, 17–34 (2002).
Gorska, R., Gregorek, H., Kowalski, J., Laskus-Perendyk, A., Syczewska, M. & Madalinski, K. Relationship between clinical parameters and cytokine profiles in inflamed gingival tissue and serum samples from patients with chronic periodontitis. J. Clin. Periodontol. 30, 1046–1052 (2003).
Dutzan, N. et al. Levels of interferon-gamma and transcription factor T-bet in progressive periodontal lesions in patients with chronic periodontitis. J. Periodontol. 80, 290–296 (2009).
Fukada, S.Y., Silva, T.A., Garlet, G.P., Rosa, A.L., da Silva, J.S. & Cunha, F.Q. Factors involved in the T helper type 1 and type 2 cell commitment and osteoclast regulation in inflammatory apical diseases. Oral Microbiol. Immunol. 24, 25–31 (2009).
Baker, P.J., Dixon, M., Evans, R.T., Dufour, L., Johnson, E. & Roopenian, D.C. CD4(+) T cells and the proinflammatory cytokines gamma interferon and interleukin-6 contribute to alveolar bone loss in mice. Infect. Immun. 67, 2804–2809 (1999).
Teng, Y.T., Mahamed, D. & Singh, B. Gamma interferon positively modulates Actinobacillus actinomycetemcomitans-specific RANKL+ CD4+ Th-cell-mediated alveolar bone destruction in vivo. Infect. Immun. 73, 3453–3461 (2005).
Zeiser, R., Youssef, S., Baker, J., Kambham, N., Steinman, L. & Negrin, R.S. Preemptive HMG-CoA reductase inhibition provides graft-versus-host disease protection by Th-2 polarization while sparing graft-versus-leukemia activity. Blood 110, 4588–4598 (2007).
De Wit, D. et al. Preferential activation of Th2 cells in chronic graft-versus-host reaction. J. Immunol. 150, 361–366 (1993).
Chen, X., Vodanovic-Jankovic, S., Johnson, B., Keller, M., Komorowski, R. & Drobyski, W.R. Absence of regulatory T-cell control of TH1 and TH17 cells is responsible for the autoimmune-mediated pathology in chronic graft-versus-host disease. Blood 110, 3804–3813 (2007).
Mauermann, N. et al. Interferon-gamma regulates idiopathic pneumonia syndrome a Th17+CD4+ T-cell-mediated graft-versus-host disease. Am. J. Respir. Crit. Care Med. 178, 379–388 (2008).
Imanguli, M.M., Swaim, W.D., League, S.C., Gress, R.E., Pavletic, S.Z. & Hakim, F.T. Increased T-bet+ cytotoxic effectors and type I interferon-mediated processes in chronic graft-versus-host disease of the oral mucosa. Blood 113, 3620–3630 (2009).
Sullivan, B.M., Juedes, A., Szabo, S.J., von Herrath, M. & Glimcher, L.H. Antigen-driven effector CD8T cell function regulated by T-bet. Proc. Natl Acad. Sci. USA 100, 15818–15823 (2003).
Glimcher, L.H., Townsend, M.J., Sullivan, B.M. & Lord, G.M. Recent developments in the transcriptional regulation of cytolytic effector cells. Nat. Rev. Immunol. 4, 900–911 (2004).
Intlekofer, A.M. et al. Anomalous type 17 response to viral infection by CD8+ T cells lacking T-bet and eomesodermin. Science 321, 408–411 (2008).
Blaser, B.W. et al. Trans-presentation of donor-derived interleukin 15 is necessary for the rapid onset of acute graft-versus-host disease but not for graft-versus-tumor activity. Blood 108, 2463–2469 (2006).
Kay, A.B. Eosinophils and asthma. N. Engl. J. Med. 324, 1514–1515 (1991).
Corrigan, C.J. & Kay, A.B. T cells and eosinophils in the pathogenesis of asthma. Immunol. Today 13, 501–507 (1992).
Finotto, S. et al. Development of spontaneous airway changes consistent with human asthma in mice lacking T-bet. Science 295, 336–338 (2002).
Park, J.W. et al. Restoration of T-box-containing protein expressed in T cells protects against allergen-induced asthma. J. Allergy Clin. Immunol. 123, 479–485 (2009).
Townsend, M.J. et al. T-bet regulates the terminal maturation and homeostasis of NK and Valpha14i NKT cells. Immunity 20, 477–494 (2004).
Kim, H.Y. et al. The development of airway hyperreactivity in T-bet-deficient mice requires CD1d-restricted NKT cells. J. Immunol. 182, 3252–3261 (2009).
Liu, N., Ohnishi, N., Ni, L., Akira, S. & Bacon, K.B. CpG directly induces T-bet expression and inhibits IgG1 and IgE switching in B cells. Nat. Immunol. 4, 687–693 (2003).
Heckman, K.L. et al. T-bet expression by dendritic cells is required for the repolarization of allergic airway inflammation. Eur. J. Immunol. 38, 2464–2474 (2008).
Wang, J. et al. Transcription factor T-bet regulates inflammatory arthritis through its function in dendritic cells. J. Clin. Invest. 116, 414–421 (2006).
Ko, F.W. et al. Decreased T-bet expression and changes in chemokine levels in adults with asthma. Clin. Exp. Immunol. 147, 526–532 (2007).
Munthe-Kaas, M.C. et al. T cell-specific T-box transcription factor haplotype is associated with allergic asthma in children. J. Allergy Clin. Immunol. 121, 51–56 (2008).
Raby, B.A. et al. T-bet polymorphisms are associated with asthma and airway hyperresponsiveness. Am. J. Respir. Crit. Care Med. 173, 64–70 (2006).
Tantisira, K.G. et al. TBX21: a functional variant predicts improvement in asthma with the use of inhaled corticosteroids. Proc. Natl Acad. Sci. USA 101, 18099–18104 (2004).
Danese, S. & Mantovani, A. Inflammatory bowel disease and intestinal cancer: a paradigm of the Yin-Yang interplay between inflammation and cancer. Oncogene 29, 3313–3323 (2010).
Wu, Y., Deng, J., Rychahou, P.G., Qiu, S., Evers, B.M. & Zhou, B.P. Stabilization of snail by NF-kappaB is required for inflammation-induced cell migration and invasion. Cancer Cell 15, 416–428 (2009).
Naito, Y. et al. CD8+ T cells infiltrated within cancer cell nests as a prognostic factor in human colorectal cancer. Cancer Res. 58,3491–3494 (1998).
Guidoboni, M. et al. Microsatellite instability and high content of activated cytotoxic lymphocytes identify colon cancer patients with a favorable prognosis. Am. J. Pathol. 159, 297–304 (2001).
Pages, F. et al. Effector memory T cells, early metastasis, and survival in colorectal cancer. N. Engl. J. Med. 353, 2654–2666 (2005).
Schiffman, M., Castle, P.E., Jeronimo, J., Rodriguez, A.C. & Wacholder, S. Human papillomavirus and cervical cancer. Lancet 370, 890–907 (2007).
Warrino, D.E. et al. Disease-stage variance in functional CD4(+) T-cell responses against novel pan-human leukocyte antigen-D region presented human papillomavirus-16 E7 epitopes. Clin. Cancer Res. 10, 3301–3308 (2004).
Seresini, S. et al. IFN-gamma produced by human papilloma virus-18 E6-specific CD4+ T cells predicts the clinical outcome after surgery in patients with high-grade cervical lesions. J. Immunol. 179, 7176–7183 (2007).
Peng, S.L., Townsend, M.J., Hecht, J.L., White, I.A. & Glimcher, L.H. T-bet regulates metastasis rate in a murine model of primary prostate cancer. Cancer Res. 64, 452–455 (2004).
Lee, K., Min, H.J., Jang, E.J., Hong, J.H. & Hwang, E.S. In vivo tumor suppression activity by T cell-specific T-bet restoration. Int. J. Cancer (2010) (e-pub ahead of print).
Werneck, M.B., Lugo-Villarino, G., Hwang, E.S., Cantor, H. & Glimcher, L.H. T-bet plays a key role in NK-mediated control of melanoma metastatic disease. J. Immunol. 180, 8004–8010 (2008).
Stoicov, C. et al. T-bet knockout prevents Helicobacter felis-induced gastric cancer. J. Immunol. 183, 642–649 (2009).
El-Omar, E.M. et al. Interleukin-1 polymorphisms associated with increased risk of gastric cancer. Nature 404, 398–402 (2000).
El-Omar, E.M. et al. Increased risk of noncardia gastric cancer associated with proinflammatory cytokine gene polymorphisms. Gastroenterology 124, 1193–1201 (2003).
Eaton, K.A., Mefford, M. & Thevenot, T. The role of T cell subsets and cytokines in the pathogenesis of Helicobacter pylori gastritis in mice. J. Immunol. 166, 7456–7461 (2001).
Svensson, A.A. et al. A 3′-untranslated region polymorphism in the TBX21 gene encoding T-bet is a risk factor for genital herpes simplex virus type 2 infection in humans. J. Gen. Virol. 89, 2262–2268.
Acknowledgements
This work was supported by grants from the Wellcome Trust (N.P., GML; 088747/z/09/Z), the National Institute for Health Research (J.B.C., GML; DRF/2009/02/22), and the Medical Research Council, UK (T.T.M., GML; G0802068). We also acknowledge financial support from the Department of Health via the National Institute for Health Research (NIHR) comprehensive Biomedical Research Centre award to Guy's & St Thomas’ NHS Foundation Trust in partnership with King's College London and King's College Hospital NHS Foundation Trust.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
PowerPoint slides
Rights and permissions
About this article
Cite this article
Powell, N., Canavan, J., MacDonald, T. et al. Transcriptional regulation of the mucosal immune system mediated by T-bet. Mucosal Immunol 3, 567–577 (2010). https://doi.org/10.1038/mi.2010.53
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2010.53
This article is cited by
-
BioTarget: A Computational Framework Identifying Cancer Type Specific Transcriptional Targets of Immune Response Pathways
Scientific Reports (2019)
-
T-bet: a bridge between innate and adaptive immunity
Nature Reviews Immunology (2013)