Abstract
Sarcomas are a rare and heterogeneous group of tumors. The last decade has witnessed extensive efforts to understand the pathobiology of many aggressive sarcoma types. In parallel, we have also begun to unravel the complex gene regulation processes mediated by microRNAs (miRNAs) in sarcomas and other cancers, discovering that microRNAs have critical roles in the majority of both oncogenic and tumor suppressor signaling networks. Expression profiles and a greater understanding of the biologic roles of microRNAs and other noncoding RNAs have considerably expanded our current knowledge and provided key pathobiological insights into many sarcomas, and helped identify novel therapeutic targets. The limited number of sarcoma patients in each sarcoma type and their heterogeneity pose distinct challenges in translating this knowledge into the clinic. It will be critical to prioritize these novel targets and choose those that have a broad applicability. A small group of microRNAs have conserved roles across many types of sarcomas and other cancers. Therapies that target these key microRNA-gene signaling and regulatory networks, in combination with standard of care treatment, may be the pivotal component in significantly improving treatment outcomes in patients with sarcoma or other cancers.
Similar content being viewed by others
Main
Sarcomas are a heterogenous group of tumors that account for ~200 000 cancers worldwide each year (~1% of all human malignant tumors); however, they represent a disproportionately high 15% of all pediatric malignant tumors.1, 2 Sarcomas comprise over 50 subtypes that can broadly be classified into bone and soft-tissue sarcomas that are generally based on the cell and/or tissue type.3 The vast majority of sarcomas fall into the soft-tissue group, primarily affecting connective tissues such as muscle (smooth and skeletal), fat, and blood vessels. Bone sarcomas are relatively rare, representing only ~20% of all diagnosed sarcomas (~0.2% of all cancers). Even within a specific subtype, sarcomas are highly heterogenous making them diagnostically and therapeutically challenging. Several sarcoma types are genetically characterized by chromosomal translocations or DNA copy number alterations, both of which are used as diagnostic markers.2, 4, 5
The four main types of bone sarcomas are defined by their histology, cell of origin (when known), clinical features, and site distribution—osteosarcoma, Ewing’s sarcoma, chondrosarcoma, and chordoma. The most common primary bone malignancy, osteosarcoma, predominantly affects children and young adults and is characterized by undifferentiated bone-forming proliferating cells.6 Ewing’s sarcoma, another aggressive pediatric malignancy, usually arises in growing bone and is genetically characterized by a fusion of EWS–FLI1 oncoproteins that act as gain-of-function transcriptional regulators.7 Chondrosarcoma is itself a heterogenous group of malignant bone tumors arising from the malignant transformation of cartilage-producing cells, frequently with mutations in IDH1/2 and COL2A1.8, 9 Chordoma is an aggressive, locally invasive cancer that typically arises from bones in the base of the skull and along the spine. It is characterized, in part, by its abnormal expression of transcription factor T, which is normally only expressed during embryonic development or in the testes.10
Soft-tissue sarcomas are also primarily defined by their histology, cell of origin, and, in some cases, by characteristic genetic translocation events. Rhabdomyosarcoma is a malignant skeletal-muscle derived tumor comprised of two main histological subtypes, embryonal and alveolar, is one of the most common childhood soft-tissue sarcomas, accounting for 6–8% of all pediatric tumors.11 Liposarcoma is the most common soft-tissue cancer overall, accounting for 20% of adult sarcoma cases. It originates in deep-tissue fat cells and is characterized primarily by amplification of the 12q chromosomal region.12 Other common soft-tissue sarcomas include angiosarcomas, fibrosarcomas, gastrointestinal stromal tumors, and synovial sarcomas, each with their own unique genetic signature.
Ever since the discovery of oncogenes, the primary emphasis in cancer research has been on understanding the role of proteins and protein-coding genes. However, the percent of the genome dedicated to coding genes is small compared with noncoding regions. The last decade has seen a surge of interest in these noncoding regions with small noncoding RNAs such as microRNAs (miRNAs) gaining particular prominence. These small RNAs have critical roles in tumor formation and progression. Understanding their roles in sarcoma will lead to new therapeutic targets and diagnostic biomarkers, opening the door to a greater understanding of the molecular mechanisms of all cancers.
miRNAs are evolutionarily conserved, small, noncoding RNA molecules of 18–24 nucleotides in length at maturity that can control gene function through mRNA degradation, translational inhibition, or chromatin-based silencing mechanisms.13 Each miRNA can potentially regulate hundreds of targets via a ‘seed’ sequence of ~5–8 nucleotides at the 5′ end of the mature miRNA. miRNAs bind to complementary sequences in the 3′-untranslated regions (3′-UTRs) of target mRNA molecules, leading to either translational repression or transcriptional degradation.14 The short seed sequence length and relatively low stringency requirement for these miRNA–3′-UTR interactions allow a single miRNA to potentially regulate hundreds of genes.15 Small changes in the expression level of a few miRNAs can therefore have a dramatic biological impact, particularly when dysregulated. miRNA expression profiles can be used to distinguish between closely related soft-tissue sarcoma subtypes and may provide a more consistent diagnosis than histological inspection.16, 17, 18
miRNAs have critical roles in the majority of canonical cellular signaling networks and their dysregulation is implicated in many cancers including breast cancer, colon cancer, gastric cancer, lung cancer, and sarcomas.19, 20 Dysregulation of miRNA expression may result from a variety of factors, including abnormal cellular stimuli, genetic mutations, epigenetic alterations, copy number variations, and chromosomal fusions. Because miRNAs act as critical regulator molecules in a variety of signaling pathways and regulatory networks, their dysregulation can be amplified across the entire signaling network.21, 22, 23, 24 Selected miRNAs and targets that have critical regulatory roles in sarcoma and other cancers are summarized in Table 1.
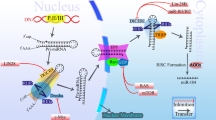
The p53 signaling pathway is one of the most highly studied cellular signaling networks. It actively induces apoptosis in response to DNA damage and oncogene activation and is therefore a key tumor suppressor pathway.25 Germline mutations in TP53 are strongly associated with the development of soft-tissue sarcomas, osteosarcoma, and are the underlying cause of Li–Fraumeni Syndrome, a familial clustering of early-onset tumors including sarcomas.26, 27 It is estimated that over 50% of human tumors harbor a TP53 mutation but over 80% of tumors have dysfunctional p53 signaling.28, 29 It is only within the last 10 years that researchers have started uncovering the roles of miRNAs in mediating p53’s activity and resulting pro-apoptotic signals (Figure 1). miRNA dysregulation could be a key factor in the ~30% of tumors with dysfunctional p53 signaling that lack an apparent TP53 mutation.
p53–miRNA interaction network. p53 interacts with the Drosha complex and promotes the processing of pri-miRNA to pre-miRNA. Although p53 directly or indirectly regulates hundreds of miRNAs, for clarity, only selected cancer-relevant miRNAs are shown. miRNAs and proteins in red are upregulated by p53. miRNAs and proteins in green are downregulated by p53. miRNAs in gray are not known to be directly regulated by p53, they are included because they target p53 regulators MDM2 and/or MDM4. miRNA, microRNA.
Like other transcription factors, p53 exerts its function primarily through transcriptional regulation of target genes that contain p53 response elements in their promoters. p53 also regulates the post-transcriptional maturation of miRNAs by interacting with the Drosha processing complex, promoting the processing of primary miRNAs to precursor miRNAs.30 In addition to protein-coding genes, many miRNA genes also contain p53 regulatory sites in their promoter regions. Large-scale screens have revealed many different miRNAs directly regulated by p53 including miR-22-3p, miR-34a, miR-125a/b, miR-182, and miR-199a-3p.31 Some of these miRNAs, such as miR-34a and miR-199a-3p, function themselves as tumor suppressors via the regulation of genes involved in cell cycle, cell proliferation, and even of itself.32, 33, 34 Although some p53-targeted miRNAs form a feedback loop, translationally and transcriptionally inhibiting the TP53 gene (e.g., miR-22-3p, miR-34a, and miR-125b), others target, or are predicted to target, p53 repressors such as MDM2 and/or MDM4 (miR-199a-3p, miR-661).31, 33, 35, 36 It is impossible to fully understand the regulation of the p53 signaling network without considering the role of these miRNAs.
miR-34a has emerged as a critical and conserved member of the p53 signaling pathway. miR-34a is downregulated in osteosarcoma tumor samples and, in conjunction with other miRNAs, regulates p53-mediated apoptosis in human osteosarcoma cell lines.32, 33, 37 The gene encoding miR-34a contains a conserved p53-binding site and is upregulated in response to cellular damage in a p53-dependent manner.37, 38 Protein-coding members of the p53 signaling pathway are well-liked targets for anticancer therapeutic development efforts and miRNAs may prove equally effective. In a preclinical model of lung cancer, therapeutic delivery of a miR-34a mimic specifically downregulated miR-34a-target genes and resulted in slower tumor growth. When combined with a siRNA targeting Kras, this small RNA combination therapy resulted in tumor regression.39 miRNAs such as miR-34a, miR-125b, and miR-199a-3p also mediate p53’s regulation of other key signaling pathways such as the IGF-1/PI3K/AKT/mTOR signaling network. Activation of the AKT network due to downregulation of PTEN (a negative regulator of AKT) by miR-21 or miR-221 or by alternate activation of AKT is a common mechanism underlying many different types of cancer.40, 41, 42, 43 The induction of cell growth, migration, invasion, and metastasis resulting from the upregulation of either miR-21 or miR-221 is seen across different tumor types.41, 44, 45, 46, 47, 48, 49, 50 Dysregulation of these miRNAs is a common factor in sarcomas and other tumors. Understanding their mechanisms of action in sarcoma could lead to broadly useful cancer therapeutics.
In prospective analyses that could be models for other sarcoma studies with sufficient numbers of patient samples, Thayanithy et al19 and Maire et al23 each analyzed collections of osteosarcoma tissues and compared them with either normal bone or osteoblasts. They each found a set of consistently downregulated miRNAs localized to the 14q32 region.19, 23 Targeting predictions performed by Thayanithy et al19 identified a subset of four miRNAs as potential regulators of cMYC. One of the many roles of cMYC is to promote the expression of the miR-17–92 family, a known oncogenic cluster that has been observed to be highly expressed in many cancer types including osteosarcoma, leiomyosarcoma, and alveolar rhabdomyosarcoma.51, 52, 53, 54, 55, 56, 57 Restoring the expression of the four 14q32 miRNAs increased apoptosis of SAOS-2 cells, an effect that was attenuated either by overexpression of a cMYC construct lacking the 3′UTR or by ectopic expression of the miR-17–92 cluster.19 Although the 14q32 region is dysregulated across many different cancer types, this pattern of dysregulation appears to be a hallmark of osteosarcoma, which is particularly interesting due to the heterogenous nature of osteosarcomas and provides an extremely attractive common therapeutic target.
One particular challenge with these types of expression profiling studies is that the cell-of-origin for a particular sarcoma subtype may not be definitely established. Another challenge is the scarcity of patient samples, particularly for the rare sarcoma subtypes. As a result, there have only been a limited number of studies designed to comprehensively profile miRNA expression in various sarcoma subtypes and to compare those expression profiles with the corresponding normal tissues or cell lines. These studies were reviewed recently in Drury et al20 and Subramanian and Kartha.58
Owing to the scarcity of frozen sarcoma tissue samples, it is tempting to study sarcoma cells in vitro, using either primary or immortalized cell cultures. Studies performed in culture are less expensive and more accessible; however, the cell lines used must be chosen with care and may not truly represent the tumors. Any results derived from cultured cells must be interpreted with caution and validated in vivo when possible. A tumor cell’s microenvironment has a profound effect on gene expression and cell metabolism and culturing for even short periods of time can result in large changes in gene/miRNA expression.59 Three-dimensional cultures can provide more physiological relevant in vitro models of individual tumors (eg, spheroid cultures) or multi-layered epithelial tissues (eg, organotypic cultures using extracellular matrix proteins, fibroblasts, and/or artificial matrix components) vs the previous standard two dimensional culture model.60, 61
Complicating the analysis of these miRNA expression changes is the fact that many miRNAs showing differential expression in multiple different studies do not have a consistent direction of change and/or a consistent role (tumor suppressor vs tumor promoter). This likely reflects both random chance observational differences and different tissue biology reflected in different regulatory networks. Elucidation of the regulatory roles played by miRNAs in these networks in their appropriate biological contexts may provide suitable upstream targets for more effective treatment of sarcomas. Recent advances in sequencing and downstream bioinformatics techniques provide the tools to efficiently examine these questions.
For two decades, microarray gene chips containing synthetic oligonucleotides whose sequences are designed to be representative of thousands of genes have allowed researchers to perform simultaneous expression analysis of thousands of RNA transcripts in a single reaction.62, 63, 64, 65 Gene expression profiling has been used to characterize and classify a wide range of sarcomas, in some cases providing a diagnostic resolution more accurate than histological examination.66, 67, 68, 69, 70, 71, 72 With the advent of high-throughput RNA-Seq, sarcoma researchers are now able to prospectively analyze the differential expression of small RNAs, such as miRNAs, without prior knowledge of their sequence.73, 74 RNA-Seq also allows for the prospective identification of novel genomic rearrangements resulting from gene fusions or premature truncations that may be of particular interest to cancer researchers.75, 76 These data are highly quantitative and digital in nature, allowing for a dynamic range that is theoretically only limited by the sequencing depth and approaches the estimated range within the cell itself.77 Marguerat and Bähler78 provide a basic overview of the different RNA-Seq technologies and their differences from array-based technologies.78
Several groups have taken advantage of these technologies to create miRNA expression profiles for a number of different sarcomas in an effort to find both common sarcoma oncomirs and to discover unique miRNA signatures that could be used in diagnosis, prognosis, and novel therapeutic development. Renner et al18 used a microarray-based miRNA screen, followed by qRT-PCR verification, to analyze the expression of 1146 known miRNAs across a collection of 76 primary soft-tissue sarcoma samples representing eight different subtypes and across a panel of 15 sarcoma cell lines. In addition to identifying overrepresented miRNAs synovial sarcomas (miR-200 family) and liposarcomas (miR-9) compared with other sarcomas and adipose tissue, respectively, their results revealed a high degree of co-expression of 63 miRNAs clustering in the chromosomal region 14q32.18 The most comprehensive sarcoma miRNA data set has been published by Sarver et al79 who profiled miRNA expression in over 300 sarcoma primary tumor tissue samples representing 22 different sarcoma types. These data form the basis for the web-accessible comprehensive Sarcoma microRNA Expression Database (SMED) database, which has tools that allows users to query specific sarcoma types and/or specific miRNAs.79
Integrative miRNA–mRNA analysis using a tool such as Ingenuity Pathway Analysis (Qiagen) or GeneSpring (Agilent) allows for more biologically relevant results by highlighting miRNA–mRNA pairs that are linked not only by predicted targeting interactions but whose expression levels are inversely correlated (i.e., as miRNA expression increases one would expect the target mRNA levels to decrease). For example, out of 177 differentially expressed miRNAs in osteosarcoma cell lines vs normal bone, an integrated miRNA–mRNA analysis highlighted two particularly interesting miRNA/mRNA pairs (miR-9/TGFBR2 and miR-29/p85α regulatory subunit of PI3K) that were dysregulated.44
It is important to note that the general consensus is that there is often no single ‘correct’ method to analyze miRNA expression data. Different experimental and bioinformatics techniques may reveal different aspects in the data that can be further investigated and experimentally validated. All of these experiments, whether performed at the bench or systems biology, contribute to our greater understanding of sarcoma biology and the central role of dysregulated miRNA–gene networks as drivers of tumor formation and progression.
miRNAs are part of a larger family of noncoding RNAs including long noncoding RNAs (lncRNAs) and competing endogenous RNAs (ceRNAs) that deserve to be evaluated for therapeutic potential in sarcomas with broader applicability to other cancer types. Just like miRNAs, lncRNAs are widely expressed in tissue-specific patterns that are highly disrupted in cancer.80 As their name implies, ceRNAs compete for their common miRNA targets and influence their expression, which has an indirect effect on the protein-coding genes, such as PTEN, regulated by those miRNAs.81, 82 We have just begun to unravel the role of lncRNAs and ceRNAs in cancer development and progression but recent results hint at yet another layer of complexity and genetic control in tumor biology.
The lessons learned from carcinomas, leukemias, and lymphomas will be helpful in understanding the pathobiology of sarcomas and the insights gained from sarcoma biology may form the foundation for therapeutics to treat a wide range of other cancers. Recent studies have shown miRNAs are very stable in blood serum and plasma, and extensive efforts are underway to develop circulating miRNA-based diagnostic and prognostic markers. Major technical challenges in developing circulating miRNA-based markers still need to be addressed, including standardization of pre-analytical, analytical, and post-analytical methods for effective reproducibility. For example, miR-16, which is used in the normalization of miRNA expression in serum/plasma is also found in red blood cells; thus, any hemolysis during sample collection could significantly affect the downstream expression data analysis.
Cancers do not exist in isolation inside the body and extensive research has been performed on how tumor-derived proteins adapt their microenvironment to provide more favorable conditions for tumor growth and development. Recent studies have shown that miRNAs also have a major role in modulating tumor microenvironment. Although most miRNAs are found inside the cell, a significant number of miRNAs are encapsulated in exosomes that can be used as a delivery system to send miRNAs from one cell to another, allowing tumor cells to modulate gene expression in surrounding tissues.83, 84 Exosome and miRNA-mediated cross talk between sarcoma tumor cells and the surrounding stromal cells is a new and exciting avenue of research and the potential for novel therapeutics is high.
Sarcomas are a diverse collection of rare cancers with proportionally limited resources for research and development of novel treatments. It is therefore crucial that potential therapeutic targets are prioritized and novel therapeutic agents carefully selected for clinical trials to succeed. Extensive studies in preclinical models will be required; however, there are also challenges in the development of appropriate in vitro and in vivo model systems that accurately reflect the different sarcoma types. Sarcomas, such as osteosarcoma, leiomyosarcoma, and angiosarcoma are very heterogeneous in nature, making it unlikely that therapies targeting specific genomic mutations will be successful. Even if specific targets were to be identified it would still be a challenge to develop clinical trials based on the small number of patients harboring those specific mutations. Coordinated efforts such as the Cancer Genome Atlas (TCGA, http://cancergenome.nih.gov/) and its associated preclinical and clinical trial consortiums will help unravel novel miRNA–mRNA interactions and their significance as potential therapeutic targets.
Targeting common miRNA–gene oncogenic or tumor suppressor networks goes after the common denominator underlying many of these cancers. Key regulatory molecules in sarcoma are highly likely to have similar roles in leukemias and lymphomas, for instance, and vice versa. For example, oncogenic activation of STAT3 strongly promotes the expression of miR-135b in lymphoma, resulting in increased angiogenesis and tumor growth.85 miR-135b is widely overexpressed in sarcomas and STAT3 may be having a similar transcriptional regulatory role, indicating that STAT3 inhibitors could be an effective supplemental therapy in sarcomas.86 Interestingly, p53 promotes the transcription of miR-125b, which can directly target both STAT3 and p53 transcription. This finely balanced regulatory network is frequently dysregulated in osteosarcoma and Ewing’s sarcoma.87, 88 In retinoblastoma, STAT3 activation is associated with upregulation of the miR-17-92 cluster via a positive feedback loop and inhibition of STAT3-suppressed retinoblastoma proliferation, providing further evidence that STAT3 may be an attractive therapeutic target in many cancers.89 The dysregulation of key signaling molecules such as the p53 and STAT3 along with their associated signaling networks are a common feature across most cancer types implying that advances in understanding of sarcoma biology may be highly impactful in more frequently occurring solid tumors and lymphomas.
Certain miRNAs appear to be common players across many types of sarcomas and other cancers and their dysregulation contributes to the development of the hallmarks of cancer (Figure 2). miR-210, a key modulator of many downstream pathways involved in the hypoxic response, is upregulated under hypoxic conditions in most solid tumors, including soft-tissue sarcomas, osteosarcoma, renal cancer, and breast cancer.90 A recent meta-analysis demonstrated that the elevated expression of miR-210 is a prognostic indicator for disease-free, progression-free, and relapse-free survival in a variety of cancer patients.91 Perhaps the most consistently upregulated miRNA across all tumor types is the anti-apoptotic miR-21, which directly targets the tumor suppressor PDCD4.92 Levels of miR-21 correlate with cancer progression and patient prognosis.93
Therapeutics targeting miRNAs represent a largely untapped pool of potential therapies for many different diseases, including cancer. The first miRNA targeted drug to show efficacy in human clinical trials, miravirsen, is an inhibitor of miR-122, a liver-specific miRNA required by the Hepatitis C virus for replication.94 The first therapeutic targeting cancer based on a miRNA mimic entered Phase 1 clinical trials in April 2013. MRX34, a liposome-formulated double-stranded mimic of tumor suppressor miR-34, is initially being tested in patients with unresectable primary liver cancer or with liver metastasis from other cancers (NCT01829971). A double-stranded mimic of miR-16, another miRNA tumor suppressor, is also in Phase 1 clinical trials to treat patients with malignant pleural mesothelioma or non-small cell lung cancer. The miR-16 mimic is formulated in a nonliving, bacterial minicell delivery vehicle targeted to EGFR-expressing cancer cells via an anti-EGFR bispecific antibody (NCT02369198). Results from both of these studies are anticipated to be reported in 2016 and could pave the way for additional miRNA-based therapeutics.
In conclusion, incorporating miRNA–gene network-based therapies, along with the standard of care treatment, that target multiple genes/pathways may result in improved treatment outcomes in patients with sarcoma and/or other cancers. There is a critical need to develop new diagnostics and therapeutics for sarcomas, which as a group account for 15% of pediatric malignancies. It appears that a small group of miRNAs have critical conserved roles in many different sarcomas, and other cancer tumors, potentially providing broadly applicable therapeutic targets. These miRNA-gene signaling and regulatory networks hold the promise of improved sarcoma diagnosis and novel therapeutic development.
References
Taylor BS, Barretina J, Maki RG et al. Advances in sarcoma genomics and new therapeutic targets. Nat Rev Cancer 2011;11:541–557.
Dela Cruz F, Matushansky I . MicroRNAs in chromosomal translocation-associated solid tumors: learning from sarcomas. Discov Med 2011;12:307–317.
Jo VY, Fletcher CD . WHO classification of soft tissue tumours: an update based on the 2013 (4th) edition. Pathology 2014;46:95–104.
Brisset S, Schleiermacher G, Peter M et al. CGH analysis of secondary genetic changes in Ewing tumors: correlation with metastatic disease in a series of 43 cases. Cancer Genet Cytogenet 2001;130:57–61.
Xie H, Lev D, Gong Y et al. Reduced mitochondrial DNA copy number in peripheral blood leukocytes increases the risk of soft tissue sarcoma. Carcinogenesis 2013;34:1039–1043.
Thomas DM, Johnson SA, Sims NA et al. Terminal osteoblast differentiation, mediated by runx2 and p27KIP1, is disrupted in osteosarcoma. J Cell Biol 2004;167:925–934.
de Alava E, Kawai A, Healey JH et al. EWS-FLI1 fusion transcript structure is an independent determinant of prognosis in Ewing’s sarcoma. J Clin Oncol 1998;16:1248–1255.
Amary MF, Bacsi K, Maggiani F et al. IDH1 and IDH2 mutations are frequent events in central chondrosarcoma and central and periosteal chondromas but not in other mesenchymal tumours. J Pathol 2011;224:334–343.
Tarpey PS, Behjati S, Cooke SL et al. Frequent mutation of the major cartilage collagen gene COL2A1 in chondrosarcoma. Nat Genet 2013;45:923–926.
Tirabosco R, Mangham DC, Rosenberg AE et al. Brachyury expression in extra-axial skeletal and soft tissue chordomas: a marker that distinguishes chordoma from mixed tumor/myoepithelioma/parachordoma in soft tissue. Am J Surg Pathol 2008;32:572–580.
Cieśla M, Dulak J, Józkowicz A . MicroRNAs and epigenetic mechanisms of rhabdomyosarcoma development. Int J Biochem Cell Biol 2014;53C:482–492.
Barretina J, Taylor BS, Banerji S et al. Subtype-specific genomic alterations define new targets for soft-tissue sarcoma therapy. Nat Genet 2010;42:715–721.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004;116:281–297.
Elbashir SM, Harborth J, Lendeckel W et al. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 2001;411:494–498.
Lewis BP, Burge CB, Bartel DP . Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005;120:15–20.
Gits CM, van Kuijk PF, Jonkers MB et al. MicroRNA expression profiles distinguish liposarcoma subtypes and implicate miR-145 and miR-451 as tumor suppressors. Int J Cancer 2014;135:348–361.
Armeanu-Ebinger S, Herrmann D, Bonin M et al. Differential expression of miRNAs in rhabdomyosarcoma and malignant rhabdoid tumor. Exp Cell Res 2012;318:2567–2577.
Renner M, Czwan E, Hartmann W et al. MicroRNA profiling of primary high-grade soft tissue sarcomas. Genes Chromosomes Cancer 2012;51:982–996.
Thayanithy V, Sarver AL, Kartha RV et al. Perturbation of 14q32 miRNAs-cMYC gene network in osteosarcoma. Bone 2012;50:171–181.
Drury R, Verghese ET, Hughes TA . The roles of microRNAs in sarcomas. J Pathol 2012;227:385–391.
Miao J, Wu S, Peng Z et al. MicroRNAs in osteosarcoma: diagnostic and therapeutic aspects. Tumour Biol 2013;34:2093–2098.
Kong YW, Ferland-McCollough D, Jackson TJ et al. microRNAs in cancer management. Lancet Oncol 2012;13:e249–e258.
Maire G, Martin JW, Yoshimoto M et al. Analysis of miRNA-gene expression-genomic profiles reveals complex mechanisms of microRNA deregulation in osteosarcoma. Cancer Genet 2011;204:138–146.
Missiaglia E, Shepherd CJ, Patel S et al. MicroRNA-206 expression levels correlate with clinical behaviour of rhabdomyosarcomas. Br J Cancer 2010;102:1769–1777.
Vogelstein B, Lane D, Levine AJ . Surfing the p53 network. Nature 2000;408:307–310.
Birch JM, Alston RD, McNally RJ et al. Relative frequency and morphology of cancers in carriers of germline TP53 mutations. Oncogene 2001;20:4621–4628.
Malkin D, Li FP, Strong LC et al. Germ line p53 mutations in a familial syndrome of breast cancer, sarcomas, and other neoplasms. Science 1990;250:1233–1238.
Olivier M, Hussain SP, Caron de Fromentel C et al. TP53 mutation spectra and load: a tool for generating hypotheses on the etiology of cancer. IARC Sci Publ. 2004;157:247–270.
Levine AJ, Hu W, Feng Z . The P53 pathway: what questions remain to be explored? Cell Death Differ 2006;13:1027–1036.
Suzuki HI, Yamagata K, Sugimoto K et al. Modulation of microRNA processing by p53. Nature 2009;460:529–533.
Tarasov V, Jung P, Verdoodt B et al. Differential regulation of microRNAs by p53 revealed by massively parallel sequencing: miR-34a is a p53 target that induces apoptosis and G1-arrest. Cell Cycle 2007;6:1586–1593.
He C, Xiong J, Xu X et al. Functional elucidation of MiR-34 in osteosarcoma cells and primary tumor samples. Biochem Biophys Res Commun 2009;388:35–40.
Tian Y, Zhang YZ, Chen W . MicroRNA-199a-3p and microRNA-34a regulate apoptosis in human osteosarcoma cells. Biosci Rep 2014;34:479–485.
Lima RT, Busacca S, Almeida GM et al. MicroRNA regulation of core apoptosis pathways in cancer. Eur J Cancer 2011;47:163–174.
Zhang Y, Gao JS, Tang X et al. MicroRNA 125a and its regulation of the p53 tumor suppressor gene. FEBS Lett. 2009;583:3725–3730.
Hoffman Y, Bublik DR, Pilpel Y et al. miR-661 downregulates both Mdm2 and Mdm4 to activate p53. Cell Death Differ 2014;21:302–309.
Raver-Shapira N, Marciano E, Meiri E et al. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol Cell 2007;26:731–743.
Shin S, Cha HJ, Lee EM et al. MicroRNAs are significantly influenced by p53 and radiation in HCT116 human colon carcinoma cells. Int J Oncol 2009;34:1645–1652.
Xue W, Dahlman JE, Tammela T et al. Small RNA combination therapy for lung cancer. Proc Natl Acad Sci USA 2014;111:E3553–E3561.
Song MS, Salmena L, Pandolfi PP . The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol 2012;13:283–296.
Zhao G, Cai C, Yang T et al. MicroRNA-221 induces cell survival and cisplatin resistance through PI3K/Akt pathway in human osteosarcoma. PLoS One 2013;8:e53906.
Steelman LS, Bertrand FE, McCubrey JA . The complexity of PTEN: mutation, marker and potential target for therapeutic intervention. Expert Opin Ther Targets 2004;8:537–550.
McCubrey JA, Steelman LS, Chappell WH et al. Mutations and deregulation of Ras/Raf/MEK/ERK and PI3K/PTEN/Akt/mTOR cascades which alter therapy response. Oncotarget 2012;3:954–987.
Namløs HM, Meza-Zepeda LA, Barøy T et al. Modulation of the osteosarcoma expression phenotype by microRNAs. PLoS One 2012;7:e48086.
Pan Q, Luo X, Chegini N . microRNA 21: response to hormonal therapies and regulatory function in leiomyoma, transformed leiomyoma and leiomyosarcoma cells. Mol Hum Reprod 2010;16:215–227.
Li X, Xin S, He Z et al. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor PDCD4 and promotes cell transformation, proliferation, and metastasis in renal cell carcinoma. Cell Physiol Biochem 2014;33:1631–1642.
Asangani IA, Rasheed SA, Nikolova DA et al. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 2008;27:2128–2136.
Meng F, Henson R, Wehbe-Janek H et al. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007;133:647–658.
Ziyan W, Shuhua Y, Xiufang W et al. MicroRNA-21 is involved in osteosarcoma cell invasion and migration. Med Oncol 2011;28:1469–1474.
Pan X, Wang ZX, Wang R . MicroRNA-21: a novel therapeutic target in human cancer. Cancer Biol Ther 2010;10:1224–1232.
He L, Thomson JM, Hemann MT et al. A microRNA polycistron as a potential human oncogene. Nature 2005;435:828–833.
Bogenmann E, Moghadam H, DeClerck YA et al. c-myc amplification and expression in newly established human osteosarcoma cell lines. Cancer Res 1987;47:3808–3814.
Gamberi G, Benassi MS, Bohling T et al. C-myc and c-fos in human osteosarcoma: prognostic value of mRNA and protein expression. Oncology 1998;55:556–563.
Mendell JT . miRiad roles for the miR-17-92 cluster in development and disease. Cell 2008;133:217–222.
Danielson LS, Menendez S, Attolini CS et al. A differentiation-based microRNA signature identifies leiomyosarcoma as a mesenchymal stem cell-related malignancy. Am J Pathol 2010;177:908–917.
Gordon AT, Brinkschmidt C, Anderson J et al. A novel and consistent amplicon at 13q31 associated with alveolar rhabdomyosarcoma. Genes Chromosomes Cancer 2000;28:220–226.
Williamson D, Selfe J, Gordon T et al. Role for amplification and expression of glypican-5 in rhabdomyosarcoma. Cancer Res 2007;67:57–65.
Subramanian S, Lui WO, Lee CH et al. MicroRNA expression signature of human sarcomas. Oncogene 2008;27:2015–2026.
Cairns RA, Harris IS, Mak TW . Regulation of cancer cell metabolism. Nat Rev Cancer 2011;11:85–95.
Egeblad M, Nakasone ES, Werb Z . Tumors as organs: complex tissues that interface with the entire organism. Dev Cell 2010;18:884–901.
Benton G, Kleinman HK, George J et al. Multiple uses of basement membrane-like matrix (BME/Matrigel) in vitro and in vivo with cancer cells. Int J Cancer 2011;128:1751–1757.
Shalon D, Smith SJ, Brown PO. A . DNA microarray system for analyzing complex DNA samples using two-color fluorescent probe hybridization. Genome Res 1996;6:639–645.
Schena M, Heller RA, Theriault TP et al. Microarrays: biotechnology’s discovery platform for functional genomics. Trends Biotechnol 1998;16:301–306.
Lockhart DJ, Dong H, Byrne MC et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat Biotechnol 1996;14:1675–1680.
Schena M, Shalon D, Heller R et al. Parallel human genome analysis: microarray-based expression monitoring of 1000 genes. Proc Natl Acad Sci USA 1996;93:10614–10619.
Skubitz KM, Skubitz AP . Characterization of sarcomas by means of gene expression. J Lab Clin Med 2004;144:78–91.
Segal NH, Pavlidis P, Antonescu CR et al. Classification and subtype prediction of adult soft tissue sarcoma by functional genomics. Am J Pathol 2003;163:691–700.
Lee YF, John M, Edwards S et al. Molecular classification of synovial sarcomas, leiomyosarcomas and malignant fibrous histiocytomas by gene expression profiling. Br J Cancer 2003;88:510–515.
Nielsen TO, West RB, Linn SC et al. Molecular characterisation of soft tissue tumours: a gene expression study. Lancet 2002;359:1301–1307.
Brohl AS, Solomon DA, Chang W et al. The genomic landscape of the Ewing Sarcoma family of tumors reveals recurrent STAG2 mutation. PLoS Genet 2014;10:e1004475.
Muff R, Ram Kumar RM, Botter SM et al. Genes regulated in metastatic osteosarcoma: evaluation by microarray analysis in four human and two mouse cell line systems. Sarcoma 2012;2012:937506.
Skubitz KM, Francis P, Skubitz AP et al. Gene expression identifies heterogeneity of metastatic propensity in high-grade soft tissue sarcomas. Cancer 2012;118:4235–4243.
Wang Z, Gerstein M, Snyder M . RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 2009;10:57–63.
Wilhelm BT, Landry JR . RNA-Seq-quantitative measurement of expression through massively parallel RNA-sequencing. Methods 2009;48:249–257.
Maher CA, Kumar-Sinha C, Cao X et al. Transcriptome sequencing to detect gene fusions in cancer. Nature 2009;458:97–101.
Zhao Q, Caballero OL, Levy S et al. Transcriptome-guided characterization of genomic rearrangements in a breast cancer cell line. Proc Natl Acad Sci USA 2009;106:1886–1891.
Mortazavi A, Williams BA, McCue K et al. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 2008;5:621–628.
Marguerat S, Bähler J . RNA-seq: from technology to biology. Cell Mol Life Sci 2010;67:569–579.
Sarver AL, Phalak R, Thayanithy V et al. S-MED: sarcoma microRNA expression database. Lab Invest 2010;90:753–761.
Gibb EA, Vucic EA, Enfield KS et al. Human cancer long non-coding RNA transcriptomes. PLoS One 2011;6:e25915.
Tay Y, Kats L, Salmena L et al. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 2011;147:344–357.
Karreth FA, Tay Y, Perna D et al. In vivo identification of tumor- suppressive PTEN ceRNAs in an oncogenic BRAF-induced mouse model of melanoma. Cell 2011;147:382–395.
Suzuki HI, Katsura A, Matsuyama H et al. MicroRNA regulons in tumor microenvironment. Oncogene 2014;34:3085–3094.
Salido-Guadarrama I, Romero-Cordoba S, Peralta-Zaragoza O et al. MicroRNAs transported by exosomes in body fluids as mediators of intercellular communication in cancer. Onco Targets Ther 2014;7:1327–1338.
Matsuyama H, Suzuki HI, Nishimori H et al. miR-135b mediates NPM-ALK-driven oncogenicity and renders IL-17-producing immunophenotype to anaplastic large cell lymphoma. Blood 2011;118:6881–6892.
Khatri R, Subramanian S . MicroRNA-135b and its circuitry networks as potential therapeutic targets in colon cancer. Front Oncol 2013;3:268.
Liu LH, Li H, Li JP et al. miR-125b suppresses the proliferation and migration of osteosarcoma cells through down-regulation of STAT3. Biochem Biophys Res Commun 2011;416:31–38.
Li J, You T, Jing J . MiR-125b inhibits cell biological progression of Ewing’s sarcoma by suppressing the PI3K/Akt signalling pathway. Cell Prolif 2014;47:152–160.
Jo DH, Kim JH, Cho CS et al. STAT3 inhibition suppresses proliferation of retinoblastoma through down-regulation of positive feedback loop of STAT3/miR-17-92 clusters. Oncotarget 2014;5:11513–11525.
Ivan M, Huang X . miR-210: fine-tuning the hypoxic response. Adv Exp Med Biol 2014;772:205–227.
Wang J, Zhao J, Shi M et al. Elevated expression of miR-210 predicts poor survival of cancer patients: a systematic review and meta-analysis. PLoS One 2014;9:e89223.
Bhatti I, Lee A, James V et al. Knockdown of microRNA-21 inhibits proliferation and increases cell death by targeting programmed cell death 4 (PDCD4) in pancreatic ductal adenocarcinoma. J Gastrointest Surg 2011;15:199–208.
Ouyang L, Liu P, Yang S et al. A three-plasma miRNA signature serves as novel biomarkers for osteosarcoma. Med Oncol 2013;30:340.
Janssen HL, Reesink HW, Lawitz EJ et al. Treatment of HCV infection by targeting microRNA. N Engl J Med 2013;368:1685–1694.
Tsang WP, Kwok TT . Let-7a microRNA suppresses therapeutics-induced cancer cell death by targeting caspase-3. Apoptosis 2008;13:1215–1222.
Zhang Z, Huang L, Yu Z et al. Let-7a functions as a tumor suppressor in Ewing’s sarcoma cell lines partly by targeting cyclin-dependent kinase 6. DNA Cell Biol 2014;33:136–147.
De Vito C, Riggi N, Suvà ML et al. Let-7a is a direct EWS-FLI-1 target implicated in Ewing’s sarcoma development. PLoS One 2011;6:e23592.
Kumar MS, Erkeland SJ, Pester RE et al. Suppression of non-small cell lung tumor development by the let-7 microRNA family. Proc Natl Acad Sci USA 2008;105:3903–3908.
Li L, Sarver AL, Alamgir S et al. Downregulation of microRNAs miR-1, -206 and -29 stabilizes PAX3 and CCND2 expression in rhabdomyosarcoma. Lab Invest 2012;92:571–583.
Chen JF, Mandel EM, Thomson JM et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat Genet 2006;38:228–233.
Yan D, Dong Xda E, Chen X et al. MicroRNA-1/206 targets c-Met and inhibits rhabdomyosarcoma development. J Biol Chem 2009;284:29596–29604.
Osaka E, Yang X, Shen JK et al. MicroRNA-1 (miR-1) inhibits chordoma cell migration and invasion by targeting slug. J Orthop Res 2014;32:1075–1082.
Cai CK, Zhao GY, Tian LY et al. miR-15a and miR-16-1 downregulate CCND1 and induce apoptosis and cell cycle arrest in osteosarcoma. Oncol Rep 2012;28:1764–1770.
Chen L, Wang Q, Wang GD et al. miR-16 inhibits cell proliferation by targeting IGF1R and the Raf1-MEK1/2-ERK1/2 pathway in osteosarcoma. FEBS Lett 2013;587:1366–1372.
Zhang X, Wan G, Mlotshwa S et al. Oncogenic Wip1 phosphatase is inhibited by miR-16 in the DNA damage signaling pathway. Cancer Res 2010;70:7176–7186.
Wang H, Garzon R, Sun H et al. NF-kappaB-YY1-miR-29 regulatory circuitry in skeletal myogenesis and rhabdomyosarcoma. Cancer Cell 2008;14:369–381.
Bommer GT, Gerin I, Feng Y et al. p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol 2007;17:1298–1307.
Sun F, Fu H, Liu Q et al. Downregulation of CCND1 and CDK6 by miR-34a induces cell cycle arrest. FEBS Lett 2008;582:1564–1568.
Bae Y, Yang T, Zeng HC et al. miRNA-34c regulates Notch signaling during bone development. Hum Mol Genet 2012;21:2991–3000.
He L, He X, Lim LP et al. A microRNA component of the p53 tumour suppressor network. Nature 2007;447:1130–1134.
Kim HR, Roe JS, Lee JE et al. A p53-inducible microRNA-34a downregulates Ras signaling by targeting IMPDH. Biochem Biophys Res Commun 2012;418:682–688.
Chen QR, Yu LR, Tsang P et al. Systematic proteome analysis identifies transcription factor YY1 as a direct target of miR-34a. J Proteome Res 2011;10:479–487.
Zhao A, Zeng Q, Xie X et al. MicroRNA-125b induces cancer cell apoptosis through suppression of Bcl-2 expression. J Genet Genomics 2012;39:29–35.
Scott GK, Goga A, Bhaumik D et al. Coordinate suppression of ERBB2 and ERBB3 by enforced expression of micro-RNA miR-125a or miR-125b. J Biol Chem 2007;282:1479–1486.
Balkhi MY, Iwenofu OH, Bakkar N et al. miR-29 acts as a decoy in sarcomas to protect the tumor suppressor A20 mRNA from degradation by HuR. Sci Signal 2013;6:ra63.
Pagliuca A, Valvo C, Fabrizi E et al. Analysis of the combined action of miR-143 and miR-145 on oncogenic pathways in colorectal cancer cells reveals a coordinate program of gene repression. Oncogene 2013;32:4806–4813.
Ban J, Jug G, Mestdagh P et al. Hsa-mir-145 is the top EWS-FLI1-repressed microRNA involved in a positive feedback loop in Ewing's sarcoma. Oncogene 2011;30:2173–2180.
Chen X, Guo X, Zhang H et al. Role of miR-143 targeting KRAS in colorectal tumorigenesis. Oncogene 2009;28:1385–1392.
Li E, Zhang J, Yuan T et al. MiR-145 inhibits osteosarcoma cells proliferation and invasion by targeting ROCK1. Tumour Biol 2014;35:7645–7650.
Mak IW, Singh S, Turcotte R et al. The epigenetic regulation of SOX9 by miR-145 in human chondrosarcoma. J Cell Biochem 2015;116:37–44.
Sarver AL, Li L, Subramanian S . MicroRNA miR-183 functions as an oncogene by targeting the transcription factor EGR1 and promoting tumor cell migration. Cancer Res 2010;70:9570–9580.
Zhu J, Feng Y, Ke Z et al. Down-regulation of miR-183 promotes migration and invasion of osteosarcoma by targeting Ezrin. Am J Pathol 2012;180:2440–2451.
Kundu ST, Byers LA, Peng DH et al. The miR-200 family and the miR-183~96~182 cluster target Foxf2 to inhibit invasion and metastasis in lung cancers. Oncogene 2015.
Myatt SS, Wang J, Monteiro LJ et al. Definition of microRNAs that repress expression of the tumor suppressor gene FOXO1 in endometrial cancer. Cancer Res 2010;70:367–377.
Wang J, Wang X, Li Z et al. MicroRNA-183 suppresses retinoblastoma cell growth, invasion and migration by targeting LRP6. FEBS J 2014;281:1355–1365.
Li J, Fu H, Xu C et al. miR-183 inhibits TGF-beta1-induced apoptosis by downregulation of PDCD4 expression in human hepatocellular carcinoma cells. BMC Cancer 2010;10:354.
Chiang CH, Hou MF, Hung WC . Up-regulation of miR-182 by β-catenin in breast cancer increases tumorigenicity and invasiveness by targeting the matrix metalloproteinase inhibitor RECK. Biochim Biophys Acta 2013;1830:3067–3076.
Taulli R, Foglizzo V, Morena D et al. Failure to downregulate the BAF53a subunit of the SWI/SNF chromatin remodeling complex contributes to the differentiation block in rhabdomyosarcoma. Oncogene 2014;33:2354–2362.
Coda DM, Lingua MF, Morena D et al. Smyd1 And G6Pd modulation are critical events for Mir-206-mediated differentiation of rhabdomyosarcoma. Cell Cycle 2015;14:1389–1402.
Papp G, Krausz T, Stricker TP et al. SMARCB1 expression in epithelioid sarcoma is regulated by miR-206, miR-381, and miR-671-5p on Both mRNA and protein levels. Genes Chromosomes Cancer 2014;53:168–176.
Acknowledgements
We thank the Wykoff Sarcoma Foundation, the Children’s Cancer Research Fund, and the American Cancer Society (RSG-13-381-01) for grant funding to SS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Sarcomas are a rare and heterogeneous group of tumors. This minireview outlines critical roles played by microRNAs in regulating and mediating conserved oncogenic and tumor suppressor signaling networks. Understanding and exploiting these critical interactions may result in new therapeutics for sarcomas and the many other cancers with similarly conserved mechanisms.
Rights and permissions
About this article
Cite this article
Sarver, A., Subramanian, S. MicroRNAs in the pathobiology of sarcomas. Lab Invest 95, 987–994 (2015). https://doi.org/10.1038/labinvest.2015.81
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2015.81
This article is cited by
-
A new treatment for sarcoma extracted from combination of miRNA deregulation and gene association rules
Signal Transduction and Targeted Therapy (2023)