Featured
-
-
News & Views |
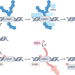 Unsprung traps keep PARP inhibitors effective
Unsprung traps keep PARP inhibitors effective
Understanding how cancers react to poly(ADP-ribose) polymerase (PARP) trapping on DNA is crucial to thwart PARP inhibitor resistance. A recent study finds that trapped PARP1 is removed via the ubiquitin-dependent segregase p97, and that perturbing this molecular cascade increases PARP inhibitor cytotoxicity.
- Peter M. Brownlee
- , Luc Provencher
- & Aaron A. Goodarzi
-
Article
| Open Access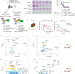 The ubiquitin-dependent ATPase p97 removes cytotoxic trapped PARP1 from chromatin
The ubiquitin-dependent ATPase p97 removes cytotoxic trapped PARP1 from chromatin
Krastev et al. report that trapped PARP1 undergoes SUMOylation, followed by ubiquitylation, resulting in the recruitment of the p97 ATPase to remove trapped PARP1 from chromatin and prevent PARP inhibitor-induced cytotoxicity.
- Dragomir B. Krastev
- , Shudong Li
- & Christopher J. Lord
-
Article
| Open Access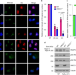 XRCC1 protects transcription from toxic PARP1 activity during DNA base excision repair
XRCC1 protects transcription from toxic PARP1 activity during DNA base excision repair
Adamowicz et al. report that toxic PARP1 activity, induced by ataxia-associated mutations in XRCC1, impairs the recovery of global transcription during DNA base excision repair by promoting aberrant recruitment and activity of the histone ubiquitin protease USP3.
- Marek Adamowicz
- , Richard Hailstone
- & Keith W. Caldecott
-
Article |
 TRIM15 and CYLD regulate ERK activation via lysine-63-linked polyubiquitination
TRIM15 and CYLD regulate ERK activation via lysine-63-linked polyubiquitination
Zhu et al. report that, in response to growth signals, ERK undergoes TRIM15-mediated lysine-63-linked ubiquitination, which facilitates ERK interaction with MEK and therefore enhances ERK activity.
- Guixin Zhu
- , Meenhard Herlyn
- & Xiaolu Yang
-
Article |
 Targeting liquid–liquid phase separation of SARS-CoV-2 nucleocapsid protein promotes innate antiviral immunity by elevating MAVS activity
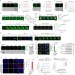
Targeting liquid–liquid phase separation of SARS-CoV-2 nucleocapsid protein promotes innate antiviral immunity by elevating MAVS activity
Wang et al. report that the nucleocapsid protein of SARS-CoV-2 forms phase-separated condensates to repress K63-linked ubiquitination and aggregation of mitochondrial antiviral-signalling protein, thus suppressing antiviral immunity.
- Shuai Wang
- , Tong Dai
- & Fangfang Zhou
-
News & Views |
 The ELOF(1)ant in the room of TCR
The ELOF(1)ant in the room of TCR
Transcription-coupled repair of DNA lesions takes place across all domains of life. A transcription-elongation factor, ELOF1, highly conserved in eukaryotes, has now been shown to play a key role in this process. Importantly, these findings also include evidence of a second transcription-coupled repair pathway in mammalian cells.
- Christopher J. Carnie
- & Stephen P. Jackson
-
Article |
 ELOF1 is a transcription-coupled DNA repair factor that directs RNA polymerase II ubiquitylation
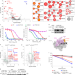
ELOF1 is a transcription-coupled DNA repair factor that directs RNA polymerase II ubiquitylation
Two side-by-side papers report that the transcription elongation factor ELOF1 drives transcription-coupled repair and prevents replication stress.
- Yana van der Weegen
- , Klaas de Lint
- & Martijn S. Luijsterburg
-
Letter |
 UFMylation maintains tumour suppressor p53 stability by antagonizing its ubiquitination
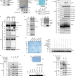
UFMylation maintains tumour suppressor p53 stability by antagonizing its ubiquitination
Liu et al. characterize a mechanism of UFL1-mediated UFMylation of p53 at multiple lysine residues and show how this dampens MDM2-induced ubiquitination of p53 to enhance its stability and tumour-suppressive function.
- Jiang Liu
- , Di Guan
- & Yu-Sheng Cong
-
Article |
 The HOIL-1L ligase modulates immune signalling and cell death via monoubiquitination of LUBAC
The HOIL-1L ligase modulates immune signalling and cell death via monoubiquitination of LUBAC
Fuseya et al. show that HOIL-1L catalyses monoubiquitination on all three LUBAC subunits, thereby impairing the function of LUBAC and its role in infection defence and dermatitis pathogenesis.
- Yasuhiro Fuseya
- , Hiroaki Fujita
- & Kazuhiro Iwai
-
News & Views |
 PTEN isoforms with dual and opposing function
PTEN isoforms with dual and opposing function
PTEN is a phosphatase that functions as a tumour suppressor by antagonizing the PI3K–AKT pathway. However, a study now demonstrates that translational variants of PTEN enable new interactions between PTEN and the COMPASS complex, identifying a new role for PTEN in modifying gene expression via COMPASS-mediated histone H3 lysine 4 methylation.
- Justin Taylor
- & Omar Abdel-Wahab
-
Article |
 PTENα and PTENβ promote carcinogenesis through WDR5 and H3K4 trimethylation
PTENα and PTENβ promote carcinogenesis through WDR5 and H3K4 trimethylation
Shen et al. show that PTENα/β stability is regulated through a ubiquitin-dependent mechanism mediated by USP9X and FBXW11 to modulate H3K4 trimethylation through WDR5 and promote tumour development.
- Shao-Ming Shen
- , Cheng Zhang
- & Guo-Qiang Chen
-
Article |
 CaMKII-δ9 promotes cardiomyopathy through disrupting UBE2T-dependent DNA repair
CaMKII-δ9 promotes cardiomyopathy through disrupting UBE2T-dependent DNA repair
Zhang et al. report the role of CaMKII-δ9, a previously less-studied isoform, in driving cardiomyopathy. Mechanistically, CaMKII-δ9 phosphorylates UBE2T, resulting in its degradation and impairing DNA repair.
- Mao Zhang
- , Hua Gao
- & Rui-Ping Xiao
-
Article |
 Ubiquilins regulate autophagic flux through mTOR signalling and lysosomal acidification
Ubiquilins regulate autophagic flux through mTOR signalling and lysosomal acidification
Şentürk et al. show that ubiquilins bind v-ATPase to control lysosome acidity, mTOR signalling and autophagic flux in neurons, and that feeding flies with acidic nanoparticles ameliorates defective autophagy in ubiquilin mutants.
- Mümine Şentürk
- , Guang Lin
- & Hugo J. Bellen
-
News & Views |
 Oncogenic AKTivation by methylation
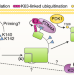
Oncogenic AKTivation by methylation
AKT, also known as protein kinase B, is one of the most frequently dysregulated serine/threonine kinases in cancer, and its hyperactivity drives tumorigenesis and chemotherapy resistance. Two studies now find that AKT methylation by the methyltransferase SETDB1 is an early step in its oncogenic activation.
- Amelia K. Luciano
- & David A. Guertin
-
Article |
 AKT methylation by SETDB1 promotes AKT kinase activity and oncogenic functions
AKT methylation by SETDB1 promotes AKT kinase activity and oncogenic functions
Guo et al. identify SETDB1 and KDM4B as the methyltransferase and demethylase, respectively, for AKT. AKT methylation promotes its kinase activity and the subsequent tumorigenesis.
- Jianping Guo
- , Xiangpeng Dai
- & Wenyi Wei
-
Article |
 SETDB1-mediated methylation of Akt promotes its
K63-linked ubiquitination and activation leading to
tumorigenesis
SETDB1-mediated methylation of Akt promotes its
K63-linked ubiquitination and activation leading to
tumorigenesis
Wang et al. show that Akt methylation by SETDB1 is recognized by demethylase JMJD2A, which then recruits E3 ligases to induce K63-linked Akt ubiquitination, leading to Akt activation and tumorigenesis.
- Guihua Wang
- , Jie Long
- & Hui-Kuan Lin
-
Article |
 L3MBTL2 orchestrates ubiquitin signalling by dictating the sequential recruitment of RNF8 and RNF168 after DNA damage
L3MBTL2 orchestrates ubiquitin signalling by dictating the sequential recruitment of RNF8 and RNF168 after DNA damage
Nowsheen et al. show that after DNA damage L3MBTL2 is recruited by MDC1 to DNA lesions where it is ubiquitylated by RNF8. Ubiquitylated L3MBTL2 then recruits RNF168 to promote DNA repair.
- Somaira Nowsheen
- , Khaled Aziz
- & Zhenkun Lou
-
Article |
 A20 promotes metastasis of aggressive basal-like breast cancers through multi-monoubiquitylation of Snail1
A20 promotes metastasis of aggressive basal-like breast cancers through multi-monoubiquitylation of Snail1
Lee et al. found that the ubiquitin-editing enzyme A20 promotes TGF-β1-induced EMT and metastases of breast cancer cells via ubiquitylation-mediated nuclear stabilization of Snail1.
- Ji-Hyung Lee
- , Su Myung Jung
- & Seok Hee Park
-
Article |
 Cholesterol and fatty acids regulate cysteine ubiquitylation of ACAT2 through competitive oxidation
Cholesterol and fatty acids regulate cysteine ubiquitylation of ACAT2 through competitive oxidation
Wang et al. show that lipid-induced ROS lead to ACAT2 stabilization by oxidizing a cysteine residue, thereby preventing its ubiquitylation and ACAT2 degradation. They further show that ACAT2 stabilization improves lipotoxicity and insulin resistance.
- Yong-Jian Wang
- , Yan Bian
- & Bao-Liang Song
-
Letter |
 Receptor oligomerization guides pathway choice between proteasomal and autophagic degradation
Receptor oligomerization guides pathway choice between proteasomal and autophagic degradation
Lu et al. show that the choice between proteasomal degradation and selective autophagy is independent of the ubiquitin-binding properties of the receptors but largely determined by oligomerization potential.
- Kefeng Lu
- , Fabian den Brave
- & Stefan Jentsch
-
News & Views |
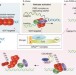 Terminating the replication helicase
Terminating the replication helicase
A feature of the cell cycle is that the events of one cycle must be reset before the next one begins. A study now shows that the replication machinery is removed from fully replicated DNA by a conserved ubiquitin- and CDC48 (also known as p97)-dependent pathway. This explains how eukaryotic chromosomes are returned to the unreplicated state.
- Vincent Gaggioli
- & Philip Zegerman
-
Article |
 CUL-2LRR-1 and UBXN-3 drive replisome disassembly during DNA replication termination and mitosis
CUL-2LRR-1 and UBXN-3 drive replisome disassembly during DNA replication termination and mitosis
The Cdc45–MCM–GINS (CMG) complex is needed for DNA replication, but how CMG dissociates from DNA in higher eukaryotes is not clear. Sonneville et al. now characterize pathways for CMG dissociation and replication termination in worm and frog.
- Remi Sonneville
- , Sara Priego Moreno
- & Karim Labib
-
Article |
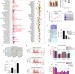 A hypoxia-responsive TRAF6–ATM–H2AX signalling axis promotes HIF1α activation, tumorigenesis and metastasis
A hypoxia-responsive TRAF6–ATM–H2AX signalling axis promotes HIF1α activation, tumorigenesis and metastasis
Lin and colleagues report that hypoxia induces TRAF6-dependent mono-ubiquitylation of histone H2AX, which promotes binding and stabilization of HIF1α. Activated HIF1α signalling in turn promotes tumorigenesis and metastasis.
- Abdol-Hossein Rezaeian
- , Chien-Feng Li
- & Hui-Kuan Lin
-
Review Article |
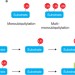 The increasing complexity of the ubiquitin code
The increasing complexity of the ubiquitin code
Yau and Rape discuss recent advances in our understanding of the many variations in ubiquitin chain topology and how these mediate ubiquitin-dependent signalling in the cell.
- Richard Yau
- & Michael Rape
-
Article |
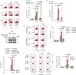 CHIP controls necroptosis through ubiquitylation- and lysosome-dependent degradation of RIPK3
CHIP controls necroptosis through ubiquitylation- and lysosome-dependent degradation of RIPK3
Receptor-interacting protein kinase 3 (RIPK3) is a key regulator of necroptosis. Seo et al. show that the E3 ligase CHIP mediates ubiquitylation and lysosomal degradation of RIPK3, thus regulating both necrosome formation and necroptosis.
- Jinho Seo
- , Eun-Woo Lee
- & Jaewhan Song
-
News & Views |
 RNF138 joins the HR team
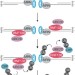
RNF138 joins the HR team
Two studies show that the E3 ubiquitin ligase RNF138 is recruited to DNA double-strand break sites, where it ubiquitylates key repair factors to promote DNA-end resection and homologous recombination. These findings add insights into the multilayered regulatory mechanisms underlying DNA double-strand break repair pathway choice in mammalian cells.
- Simon Bekker-Jensen
- & Niels Mailand
-
Article |
 The RNF138 E3 ligase displaces Ku to promote DNA end resection and regulate DNA repair pathway choice
The RNF138 E3 ligase displaces Ku to promote DNA end resection and regulate DNA repair pathway choice
Jackson and colleagues and Hendzel and colleagues reveal that the E3 ligase RNF138 functions in the repair of double-strand breaks by promoting CtIP accumulation and displacement of DNA-PK subunit Ku.
- Ismail Hassan Ismail
- , Jean-Philippe Gagné
- & Michael J. Hendzel
-
Article |
 Systematic E2 screening reveals a UBE2D–RNF138–CtIP axis promoting DNA repair
Systematic E2 screening reveals a UBE2D–RNF138–CtIP axis promoting DNA repair
Jackson and colleagues and Hendzel and colleagues reveal that the E3 ligase RNF138 functions in the repair of double-strand breaks by promoting CtIP accumulation and displacement of DNA-PK subunit Ku.
- Christine K. Schmidt
- , Yaron Galanty
- & Stephen P. Jackson
-
Resource |
 Systematic proteomics of the VCP–UBXD adaptor network identifies a role for UBXN10 in regulating ciliogenesis
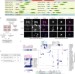
Systematic proteomics of the VCP–UBXD adaptor network identifies a role for UBXN10 in regulating ciliogenesis
Through proteomics, Harper and colleagues identify proteins interacting with UBXD adaptors for the multifunctional AAA-ATPase VCP and reveal a role for UBXN10 in ciliogenesis.
- Malavika Raman
- , Mikhail Sergeev
- & J. Wade Harper
-
Article |
 Deubiquitylase OTUD3 regulates PTEN stability and suppresses tumorigenesis
Deubiquitylase OTUD3 regulates PTEN stability and suppresses tumorigenesis
By screening 85 deubiquitylation enzymes, Zhang and colleagues identify OTUD3 as an enzyme that upregulates PTEN levels by deubiquitylation and acts as a tumour suppressor in synergy with another PTEN DUB, USP13.
- Lin Yuan
- , Yanrong Lv
- & Lingqiang Zhang
-
News & Views |
 Solving the centriole disengagement puzzle
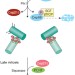
Solving the centriole disengagement puzzle
The microcephaly protein, Cep215, contributes to the engagement of duplicated centrioles in interphase. Now two distinct pools of Cep215 at centrosomes are identified, one bound to Cep68 and the other to pericentrin. Plk1-mediated degradation of Cep68 and separase-mediated cleavage of pericentrin release both pools of Cep215, thereby promoting centriole disengagement.
- Andrew M. Fry
-
Article |
 Degradation of Cep68 and PCNT cleavage mediate Cep215 removal from the PCM to allow centriole separation, disengagement and licensing
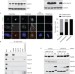
Degradation of Cep68 and PCNT cleavage mediate Cep215 removal from the PCM to allow centriole separation, disengagement and licensing
Pagano and colleagues find that Plk1 and the E3 ubiquitin ligase SCFβTrCP mediate degradation of the centrosome cohesion protein Cep68 and show this mediates removal of Cep215 from the PCM and subsequent centriole separation in late mitosis.
- Julia K. Pagan
- , Antonio Marzio
- & Michele Pagano
-
News & Views |
 Rsp5/Nedd4 clears cells of heat-damaged proteins
Rsp5/Nedd4 clears cells of heat-damaged proteins
Protein quality control systems protect cells from proteotoxicity caused by the accumulation of aberrantly folded polypeptides. The Rsp5 ubiquitin ligase (mammalian homologue Nedd4) is now identified as a major constituent of a clearance pathway that degrades misfolded cytosolic proteins after exposure to heat.
- Thomas Sommer
- , Annika Weber
- & Ernst Jarosch
-
Article |
 Rsp5/Nedd4 is the main ubiquitin ligase that targets cytosolic misfolded proteins following heat stress
Rsp5/Nedd4 is the main ubiquitin ligase that targets cytosolic misfolded proteins following heat stress
Mayor and colleagues identify yeast Rsp5 (mammalian homologue Nedd4) as the main ubiquitin ligase responsible for the increased ubiquitylation following heat stress when Rsp5 targets cytosolic misfolded proteins for proteasome degradation.
- Nancy N. Fang
- , Gerard T. Chan
- & Thibault Mayor
-
Resource |
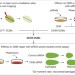 Systematic characterization of deubiquitylating enzymes for roles in maintaining genome integrity
Systematic characterization of deubiquitylating enzymes for roles in maintaining genome integrity
Systematic characterization of deubiquitylating enzymes in the DNA-damage response identifies UCHL5 as promoting DNA-end resection.
- Ryotaro Nishi
- , Paul Wijnhoven
- & Stephen P. Jackson
-
-
Article |
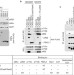 FBXL2- and PTPL1-mediated degradation of p110-free p85β regulatory subunit controls the PI(3)K signalling cascade
FBXL2- and PTPL1-mediated degradation of p110-free p85β regulatory subunit controls the PI(3)K signalling cascade
Phosphatidylinositol-3-OH kinase (PI(3)K) signalling regulates many cellular events such as cell growth and survival. Pagano and colleagues show that the E3 ligase SCF-FBXL2 promotes PI(3)K signalling and inhibits autophagy by targeting the free p85β subunit of the p110–p85 PI(3)K complex for degradation.
- Shafi Kuchay
- , Shanshan Duan
- & Michele Pagano
-
News & Views |
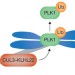 Cullin' PLK1 from kinetochores
Cullin' PLK1 from kinetochores
To ensure proper attachment of all chromosomes to the spindle, PLK1 has to associate with kinetochores during prometaphase and must be released from these sites before sister chromatid separation can begin. The monoubiquitylation of PLK1 by the ubiquitin ligase CUL3–KLHL22 is now identified as a critical step in promoting the release of PLK1 from kinetochores, pushing non-proteolytic ubiquitylation into the limelight of cell division research.
- Colleen A. McGourty
- & Michael Rape
-
Article |
 Ubiquitylation-dependent localization of PLK1 in mitosis
Ubiquitylation-dependent localization of PLK1 in mitosis
The dynamic localization of PLK1 to kinetochores is required for faithful chromosome segregation. Peter, Sumara and colleagues demonstrate that a KHL22-containing E3 ligase mediates degradation-independent removal of PLK1 from kinetochores to ensure satisfaction of the spindle assembly checkpoint and proper mitotic progression.
- Jochen Beck
- , Sarah Maerki
- & Izabela Sumara
-
-
Article |
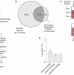 Systems-wide analysis of ubiquitylation dynamics reveals a key role for PAF15 ubiquitylation in DNA-damage bypass
Systems-wide analysis of ubiquitylation dynamics reveals a key role for PAF15 ubiquitylation in DNA-damage bypass
In a quantitative proteomics approach, Mailand, Choudhary and colleagues characterize ultraviolet-regulated ubiquitylation sites and identify a role for double mono-ubiquitylation of PCNA-associated factor PAF15 in bypassing replication-blocking lesions in DNA.
- Lou K. Povlsen
- , Petra Beli
- & Chunaram Choudhary
-
-
Article |
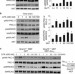 PRR5L degradation promotes mTORC2-mediated PKC-δ phosphorylation and cell migration downstream of Gα12
PRR5L degradation promotes mTORC2-mediated PKC-δ phosphorylation and cell migration downstream of Gα12
Wu and colleagues delineate an mTORC2-dependent cell migration pathway. They show that stimulation of the Gα12 protein subunit induces the ARAF/ERK-mediated expression of the RFFL E3 ubiquitin ligase. RFFL, in turn, targets the inhibitory PRR5L subunit of the mTORC2 complex for ubiquitylation and degradation, enabling mTORC2 to phosphorylate PKC-δ and promote cell migration.
- Xiaoqing Gan
- , Jiyong Wang
- & Dianqing Wu
-
Article |
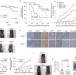 TRADD contributes to tumour suppression by regulating ULF-dependent p19Arf ubiquitylation
TRADD contributes to tumour suppression by regulating ULF-dependent p19Arf ubiquitylation
Mak and colleagues report that TRADD, an adaptor protein important in tumour necrosis factor receptor (TNFR) signalling, contributes to tumour suppression independently of TNFR. They show that nuclear TRADD binds to the tumour suppressor p19Arf and inhibits its interaction with the E3 ubiquitin ligase ULF, leading to p19Arf stability and promoting cell senescence.
- Iok In Christine Chio
- , Masato Sasaki
- & Tak Wah Mak
-
Article |
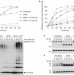 APC/C-mediated multiple monoubiquitylation provides an alternative degradation signal for cyclin B1
APC/C-mediated multiple monoubiquitylation provides an alternative degradation signal for cyclin B1
Cyclin B is targeted for proteasome-mediated degradation by the E3 ligase APC/C, which is thought to generate polyubiquitin chains for the degradation of mitotic substrates. King and colleagues now demonstrate in Xenopus laevis extracts that multiple monoubiquitylation events are sufficient to target cyclin B1 for degradation.
- Nevena V. Dimova
- , Nathaniel A. Hathaway
- & Randall W. King
-
Letter |
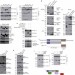 The LIMD1 protein bridges an association between the prolyl hydroxylases and VHL to repress HIF-1 activity
The LIMD1 protein bridges an association between the prolyl hydroxylases and VHL to repress HIF-1 activity
Oxygen levels regulate the stability of the transcription factor HIF-1 through the action of prolyl hydroxylases and the VHL ubiquitin ligase. Sharp and colleagues now identify a protein complex in which the Ajuba LIM-domain protein LIMD1 brings together prolyl hydroxylases and VHL to ensure efficient degradation of HIF-1.
- Daniel E. Foxler
- , Katherine S. Bridge
- & Tyson V. Sharp
-
Letter |
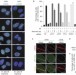 The ubiquitin-selective segregase VCP/p97 orchestrates the response to DNA double-strand breaks
The ubiquitin-selective segregase VCP/p97 orchestrates the response to DNA double-strand breaks
The p97 AAA+ ATPase (also known as VCP) functions in various ubiquitin-regulated processes. Ramadan and colleagues now find that p97 is recruited to sites of DNA damage by Lys-48-linked ubiquitin chains, which are formed in a process mediated by RNF8. p97 then removes Lys-48–ubiquitin conjugates and promotes recruitment of DNA-repair factors.
- Mayura Meerang
- , Danilo Ritz
- & Kristijan Ramadan
-
News & Views |
 Misfolded proteins driven to destruction by Hul5
Misfolded proteins driven to destruction by Hul5
Misfolded proteins are potentially toxic and are therefore subjected to highly selective degradation by the ubiquitin–proteasome system. The identification of the Hul5 ubiquitin ligase as a major mediator of such 'quality-control' ubiquitylation following heat shock raises new questions about the design of these pathways.
- Daniel Finley
-
Article |
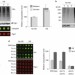 Hul5 HECT ubiquitin ligase plays a major role in the ubiquitylation and turnover of cytosolic misfolded proteins
Hul5 HECT ubiquitin ligase plays a major role in the ubiquitylation and turnover of cytosolic misfolded proteins
The ubiquitin–proteasome system clears misfolded proteins to maintain cellular homeostasis. Mayor and colleagues identify the ubiquitin ligase Hul5 as a critical component of the heat-shock response and show that it selectively targets misfolded cytosolic proteins for degradation.
- Nancy N. Fang
- , Alex H. M. Ng
- & Thibault Mayor

 CDK1–cyclin-B1-induced kindlin degradation drives focal adhesion disassembly at mitotic entry
CDK1–cyclin-B1-induced kindlin degradation drives focal adhesion disassembly at mitotic entry