Featured
-
-
Article
| Open Access Single-cell entropy for accurate estimation of differentiation potency from a cell’s transcriptome
Single-cell entropy for accurate estimation of differentiation potency from a cell’s transcriptome
Robust quantification of the differentiation potential of single cells is a task of great importance. Here the authors integrate single-cell RNA-Seq profiles with a cellular interaction network to compute the signaling entropy, and show that it can identify normal and cancer stem-cell phenotypes.
- Andrew E. Teschendorff
- & Tariq Enver
-
Article
| Open Access R2TP/Prefoldin-like component RUVBL1/RUVBL2 directly interacts with ZNHIT2 to regulate assembly of U5 small nuclear ribonucleoprotein
R2TP/Prefoldin-like component RUVBL1/RUVBL2 directly interacts with ZNHIT2 to regulate assembly of U5 small nuclear ribonucleoprotein
The R2TP/Prefoldin-like cochaperone complex is involved in the assembly of a number of protein complexes. Here the authors provide evidence that RUVBL1/RUVBL2, subunits of that cochaperone complex, directly interact with ZNHIT2 to regulate assembly of U5 small ribonucleoprotein.
- Philippe Cloutier
- , Christian Poitras
- & Benoit Coulombe
-
Article
| Open Access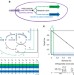 Redesigning metabolism based on orthogonality principles
Redesigning metabolism based on orthogonality principles
Growth-coupled designs for chemical production are limited by native metabolic networks’ optimality for growth. Here, the authors introduce pathway orthogonality as a measure of the independence of biomass and chemical production pathways, identify metabolic valves that allow substrate utilization to be switched between the two, and demonstrate advantages of orthogonal designs.
- Aditya Vikram Pandit
- , Shyam Srinivasan
- & Radhakrishnan Mahadevan
-
Article
| Open Access Digital logic circuits in yeast with CRISPR-dCas9 NOR gates
Digital logic circuits in yeast with CRISPR-dCas9 NOR gates
The leakiness of commonly used genetic components can make the construction of complex synthetic circuits difficult. Here the authors construct NOR gate architecture, using dCas9 fused to the chromatin remodeller Mxi1, that can be wired together into complex circuits.
- Miles W. Gander
- , Justin D. Vrana
- & Eric Klavins
-
Article
| Open Access Design of synthetic epigenetic circuits featuring memory effects and reversible switching based on DNA methylation
Design of synthetic epigenetic circuits featuring memory effects and reversible switching based on DNA methylation
Recording systems would allow synthetic organisms to store a ‘memory’ of a past event for future reference. Here the authors design an epigenetic memory system inE. colithat methylates DNA in response to exogenous and endogenous signals.
- Johannes A. H. Maier
- , Raphael Möhrle
- & Albert Jeltsch
-
Article
| Open Access An integrated model for detecting significant chromatin interactions from high-resolution Hi-C data
An integrated model for detecting significant chromatin interactions from high-resolution Hi-C data
Genome-wide chromosome conformation capture has helped us identify features of genome topology influencing biology but requires careful statistical analysis. Here the authors present HiC-DC, a bioinformatics method that can detect statistically significant regulatory interactions.
- Mark Carty
- , Lee Zamparo
- & Christina S. Leslie
-
Article
| Open Access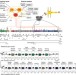 Control of type III protein secretion using a minimal genetic system
Control of type III protein secretion using a minimal genetic system
The type III secretion system is a needle-like molecular machine under tight regulatory control. Here the authors construct a synthetic type III secretion system gene cluster by deconstructing and rebuilding the wild-typeSalmonellapathogenicity island 1.
- Miryoung Song
- , David J. Sukovich
- & Christopher A. Voigt
-
Article
| Open Access Tradict enables accurate prediction of eukaryotic transcriptional states from 100 marker genes
Tradict enables accurate prediction of eukaryotic transcriptional states from 100 marker genes
Global patterns of gene transcription can be represented with reduced dimensionality. Here, the authors devise a method called Tradict that learns and uses 100 marker genes to predict transcriptome-wide pathway expression levels and patterns that reflect cell activity and state.
- Surojit Biswas
- , Konstantin Kerner
- & Philip A. Wigge
-
Article
| Open Access Subsampling scaling
Subsampling scaling
We can often observe only a small fraction of a system, which leads to biases in the inference of its global properties. Here, the authors develop a framework that enables overcoming subsampling effects, apply it to recordings from developing neural networks, and find that neural networks become critical as they mature.
- A. Levina
- & V. Priesemann
-
Article
| Open Access Automated multiplex genome-scale engineering in yeast
Automated multiplex genome-scale engineering in yeast
Genome-scale engineering is a powerful technique for understanding biology and designing microorganisms but has been limited to bacterial species. Here the authors present an automated platform for genome-scale engineering inSaccharomyces cerevisiaeusing CRISPR-Cas and RNAi.
- Tong Si
- , Ran Chao
- & Huimin Zhao
-
Article
| Open Access Biosynthesis of the antibiotic nonribosomal peptide penicillin in baker’s yeast
Biosynthesis of the antibiotic nonribosomal peptide penicillin in baker’s yeast
Filamentous fungi are a valuable source of natural therapeutic products such as antibiotics. Here the authors engineer monocellularS. cerevisiaeto perform complex secondary metabolism typical of multicellular fungi in order to demonstrate biosynthesis and secretion of bioactive penicillin.
- Ali R. Awan
- , Benjamin A. Blount
- & Tom Ellis
-
Article
| Open Access Similarity in viral and host promoters couples viral reactivation with host cell migration
Similarity in viral and host promoters couples viral reactivation with host cell migration
The coevolution of viruses and host cells can be mapped with interactomics. Here the authors identify coupling of human and viral promoters, and show that HIV-reactivation from dormancy is coincident with migration of HIV-infected cells owing to coupling of human CXCR4 and HIV LTR promoters.
- Kathrin Bohn-Wippert
- , Erin N. Tevonian
- & Roy D. Dar
-
Article
| Open Access Programming mRNA decay to modulate synthetic circuit resource allocation
Programming mRNA decay to modulate synthetic circuit resource allocation
Synthetic circuits in host cells compete with endogenous processes for limited resources. Here the authors use MazF to funnel cellular resources to a synthetic circuit to increase product production and demonstrate how resource allocation can be manipulated.
- Ophelia S. Venturelli
- , Mika Tei
- & Adam P Arkin
-
Article
| Open Access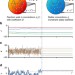 Exploratory adaptation in large random networks
Exploratory adaptation in large random networks
Recent works suggest that cellular networks may respond to novel challenges on the time-scale of cellular lifetimes through large-scale perturbation of gene expression and convergence to a new state. Here, the authors demonstrate the theoretical feasibility of exploratory adaptation in cellular networks by showing that convergence to new states depends on known features of these networks.
- Hallel I. Schreier
- , Yoav Soen
- & Naama Brenner
-
Article
| Open Access A mouse tissue transcription factor atlas
A mouse tissue transcription factor atlas
While we have abundant data for transcription factor (TF) binding sites and TF expression at the mRNA level, our knowledge of TFs at the protein level and their DNA-binding activities is sparser. Here, the authors address this by using the catTFRE approach to profile active TFs in 24 adult and 8 fetal mouse tissues, and presenting the TF networks in major mouse organs.
- Quan Zhou
- , Mingwei Liu
- & Jun Qin
-
Article
| Open Access Engineered probiotic Escherichia coli can eliminate and prevent Pseudomonas aeruginosa gut infection in animal models
Engineered probiotic Escherichia coli can eliminate and prevent Pseudomonas aeruginosa gut infection in animal models
Bacteria can be engineered to kill specific pathogens. Here, the authors modify and optimize a synthetic genetic system in a probiotic strain ofEscherichia coli, and show that the engineered probiotic can sense and kill the pathogen Pseudomonas aeruginosain two animal models of gut infection.
- In Young Hwang
- , Elvin Koh
- & Matthew Wook Chang
-
Article
| Open Access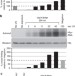 Kinetic CRAC uncovers a role for Nab3 in determining gene expression profiles during stress
Kinetic CRAC uncovers a role for Nab3 in determining gene expression profiles during stress
Protein RNA interactions are dynamic and regulated in response to environmental changes. Here the authors describe ‘kinetic CRAC’, an approach that allows time resolved analyses of protein RNA interactions with minute time point resolution and apply it to gain insight into the function of the RNA-binding protein Nab3.
- Rob van Nues
- , Gabriele Schweikert
- & Sander Granneman
-
Article
| Open Access An analytic approximation of the feasible space of metabolic networks
An analytic approximation of the feasible space of metabolic networks
Large-scale metabolic models of organisms from microbes to mammals can provide great insight into cellular function, but their analysis remains challenging. Here, the authors provide an approximate analytic method to estimate the feasible solution space for the flux vectors of metabolic networks, enabling more accurate analysis under a wide range of conditions of interest.
- Alfredo Braunstein
- , Anna Paola Muntoni
- & Andrea Pagnani
-
Article
| Open Access Sensitive detection of rare disease-associated cell subsets via representation learning
Sensitive detection of rare disease-associated cell subsets via representation learning
While rare cell subpopulations frequently make the difference between health and disease, their detection remains a challenge. Here, the authors devise CellCnn, a representation learning approach to detecting such rare cell populations from high-dimensional single cell data, and, among other examples, demonstrate its capacity for detecting rare leukaemic blasts in minimal residual disease.
- Eirini Arvaniti
- & Manfred Claassen
-
Article
| Open Access Light-sensing via hydrogen peroxide and a peroxiredoxin
Light-sensing via hydrogen peroxide and a peroxiredoxin
While yeasts lack dedicated photoreceptors, they nonetheless possess metabolic rhythms responsive to light. Here the authors find that light signalling in budding yeast involves the production of H2O2, which in turn regulates protein kinase A through a peroxiredoxin-thioredoxin redox relay.
- Kristofer Bodvard
- , Ken Peeters
- & Mikael Molin
-
Article
| Open Access MicroRNA filters Hox temporal transcription noise to confer boundary formation in the spinal cord
MicroRNA filters Hox temporal transcription noise to confer boundary formation in the spinal cord
In the spinal cord, someHox genes are transcribed in progenitors while their proteins are only detected in differentiating postmitotic motor neurons. Here, the authors show that miRNAs (specifically mir-27) regulate post-transcriptional Hoxa5 expression in motor neurons.
- Chung-Jung Li
- , Tian Hong
- & Jun-An Chen
-
Article
| Open Access Reconstruction of the metabolic network of Pseudomonas aeruginosa to interrogate virulence factor synthesis
Reconstruction of the metabolic network of Pseudomonas aeruginosa to interrogate virulence factor synthesis
Targeting virulence rather than bacterial growth is less likely to select for antibiotic resistance, but many possible targets function in both processes. Here, the authors reconstruct a genome-scale metabolic network ofP. aeruginosastrain PA14 and update that of strain PAO1, which, together with mutant screens, enable them to identify genes uniquely critical for virulence factor production.
- Jennifer A. Bartell
- , Anna S. Blazier
- & Jason A. Papin
-
Article
| Open Access The OncoPPi network of cancer-focused protein–protein interactions to inform biological insights and therapeutic strategies
The OncoPPi network of cancer-focused protein–protein interactions to inform biological insights and therapeutic strategies
Understanding of dysregulation in cancers requires knowledge, beyond cancer genomes, of the interactions of cancer-associated proteins. Here, the authors use high-throughput, time-resolved FRET to map protein–protein interactions to establish a lung cancer protein network, and demonstrate its utility in revealing new oncogenic pathways and connectivity of tumour suppressors with druggable targets.
- Zenggang Li
- , Andrei A. Ivanov
- & Haian Fu
-
Article
| Open Access Recurrently deregulated lncRNAs in hepatocellular carcinoma
Recurrently deregulated lncRNAs in hepatocellular carcinoma
Long noncoding-RNAs have been linked to hepatocellular carcinoma (HCC) and some can be used as prognostic markers. Here the authors, by analysing RNA-seq in 60 clinical samples from 20 patients, provide a resource of functional lncRNAs and biomarkers associated with HCC tumorigenesis and metastasis.
- Yang Yang
- , Lei Chen
- & Zhi John Lu
-
Article
| Open Access Cell fate decisions emerge as phages cooperate or compete inside their host
Cell fate decisions emerge as phages cooperate or compete inside their host
The bacteriophage lambda and its hostEscherichia coli provide a model system to study cell-fate decisions. Here, Trinh et al. develop a four-colour fluorescence system at the single-cell/single-virus/single-viral-DNA level and find phages cooperate during lysogenization and compete during lysis.
- Jimmy T. Trinh
- , Tamás Székely
- & Lanying Zeng
-
Article
| Open Access Serotonin-dependent kinetics of feeding bursts underlie a graded response to food availability in C. elegans
Serotonin-dependent kinetics of feeding bursts underlie a graded response to food availability in C. elegans
Regulating food intake is an important physiological mechanism. Here, the authors use a custom microfluidic device to investigate feeding dynamics inC. elegans, and identify roles of serotonergic neurons in regulating bursts of feeding in response to food availability.
- Kyung Suk Lee
- , Shachar Iwanir
- & Erel Levine
-
Article
| Open Access The impact of microRNAs on transcriptional heterogeneity and gene co-expression across single embryonic stem cells
The impact of microRNAs on transcriptional heterogeneity and gene co-expression across single embryonic stem cells
MicroRNAs can posttranscriptionally repress multiple targets in a cell population. Here the authors use single-cell sequencing to investigate the effects of an individual miRNA on transcriptional heterogeneity and gene co-expression
- Gennaro Gambardella
- , Annamaria Carissimo
- & Robert Blelloch
-
Article
| Open Access Optimality and sub-optimality in a bacterial growth law
Optimality and sub-optimality in a bacterial growth law
Organisms improve their fitness by adjusting their gene expression to the environment, for example bacteria scale the expression of metabolic enzymes near linearly to their growth rate. Here, the authors show that such linear scaling often maximizes growth rate, but that linear scaling is suboptimal under some conditions.
- Benjamin D. Towbin
- , Yael Korem
- & Uri Alon
-
Article
| Open Access A genome-scale Escherichia coli kinetic metabolic model k-ecoli457 satisfying flux data for multiple mutant strains
A genome-scale Escherichia coli kinetic metabolic model k-ecoli457 satisfying flux data for multiple mutant strains
Kinetic models of microbial metabolism have great potential to aid metabolic engineering efforts, but the challenge of parameterization has so far limited them to core metabolism. Here, the authors introduce a genome-scale metabolic model of E. colimetabolism that satisfies fluxomic data for a wild-type and 25 mutant strains in various growth conditions.
- Ali Khodayari
- & Costas D. Maranas
-
Article
| Open Access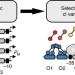 Engineering orthogonal dual transcription factors for multi-input synthetic promoters
Engineering orthogonal dual transcription factors for multi-input synthetic promoters
Genetic circuits usually employ the same set of transcription factors which can act via repression or activation of the target promoter. Here the authors present dual activator-repressor switches, designed via directed evolution, for orthogonal logic gates and multi-input circuit architectures.
- Andreas K. Brödel
- , Alfonso Jaramillo
- & Mark Isalan
-
Article
| Open Access Inferring time derivatives including cell growth rates using Gaussian processes
Inferring time derivatives including cell growth rates using Gaussian processes
High-throughput time-series data is increasingly available, yet estimating time-derivatives from such data can remain a challenge. Here, the authors provide a non-parametric method for inferring the first and second time-derivatives from multiple replicates of time-series data and for estimating errors in this inference and in any summary statistics.
- Peter S. Swain
- , Keiran Stevenson
- & Teuta Pilizota
-
Article
| Open Access Altered intestinal microbiota–host mitochondria crosstalk in new onset Crohn’s disease
Altered intestinal microbiota–host mitochondria crosstalk in new onset Crohn’s disease
Crohn’s disease is associated with altered intestinal microbiota. Here, the authors show that the microbe Atopobium parvulumis associated with Crohn’s disease patients, triggers colitis in a mouse model, and that scavenging microbe-induced hydrogen sulfide improved symptoms in mice.
- Walid Mottawea
- , Cheng-Kang Chiang
- & Alain Stintzi
-
Article
| Open Access T-cell stimuli independently sum to regulate an inherited clonal division fate
T-cell stimuli independently sum to regulate an inherited clonal division fate
Why do populations of highly similar T cells have heterogeneous division destinies in response to antigenic stimulus? Here the authors develop a multiplex-dye assay and a mathematical framework to test clonal heterogeneity and show distinction in division destiny is a result of inter-clonal variability as lineage imprinting ensures clones share similar proliferation fates.
- J. M. Marchingo
- , G. Prevedello
- & K. R. Duffy
-
Article
| Open Access Boosting functionality of synthetic DNA circuits with tailored deactivation
Boosting functionality of synthetic DNA circuits with tailored deactivation
Nonlinearity in synthetic molecular circuits is usually achieved by manipulation of network topology or of production kinetics. Here, the authors achieve bistability and other nonlinear behaviours by manipulating the individual degradation rate laws of circuit components using saturable pathways.
- Kevin Montagne
- , Guillaume Gines
- & Yannick Rondelez
-
Article
| Open Access Glucose-regulated and drug-perturbed phosphoproteome reveals molecular mechanisms controlling insulin secretion
Glucose-regulated and drug-perturbed phosphoproteome reveals molecular mechanisms controlling insulin secretion
Dysfunction in insulin secretion is a main driver of type 2 diabetes development. Here the authors monitor phosphoproteome modulation in cells stimulated with glucose and treated with drugs affecting glucose-mediated insulin secretion to reveal phosphorylation sites implicated in insulin secretion control and gene expression regulation.
- Francesca Sacco
- , Sean J. Humphrey
- & Matthias Mann
-
Article
| Open Access Multi-omic data integration enables discovery of hidden biological regularities
Multi-omic data integration enables discovery of hidden biological regularities
Translating omics data sets into biological insight is one of the great challenges of our time. Here, the authors make headway by synchronising pairs of omics data types via invariants across conditions and by integrating datasets into a genome-scale model of E. coli metabolism and gene expression.
- Ali Ebrahim
- , Elizabeth Brunk
- & Bernhard O. Palsson
-
Article
| Open Access A network property necessary for concentration robustness
A network property necessary for concentration robustness
Absolute concentration robustness (ACR), independence of the steady-state concentration of a molecule from the environment, is difficult to predict. Here, the authors derive a network structure-based necessary condition for ACR, and suggest that metabolites satisfying the condition are prevalent.
- Jeanne M. O. Eloundou-Mbebi
- , Anika Küken
- & Zoran Nikoloski
-
Article
| Open Access In vivo continuous evolution of genes and pathways in yeast
In vivo continuous evolution of genes and pathways in yeast
Directed evolution is a powerful technique for generating improved biological systems through repeated rounds of mutagenesis and selection. Here the authors engineer the yeast retrotransposon Ty1 to enable the creation of large mutant libraries in vivoand use this system to generate improved variants of single enzymes and multigene pathways.
- Nathan Crook
- , Joseph Abatemarco
- & Hal S. Alper
-
Article
| Open Access Multi-omics integration accurately predicts cellular state in unexplored conditions for Escherichia coli
Multi-omics integration accurately predicts cellular state in unexplored conditions for Escherichia coli
Multi-omics data integration is a great challenge. Here, the authors compile a database of E. coliproteomics, transcriptomics, metabolomics and fluxomics data to train models of recurrent neural network and constrained regression, enabling prediction of bacterial responses to perturbations.
- Minseung Kim
- , Navneet Rai
- & Ilias Tagkopoulos
-
Article
| Open Access A modular platform for one-step assembly of multi-component membrane systems by fusion of charged proteoliposomes
A modular platform for one-step assembly of multi-component membrane systems by fusion of charged proteoliposomes
Assembling multiple biological components into synthetic lipid vesicles is a limiting step in the manufacture of biomimetic cell-like structures. Here the authors use fusogenic proteoliposomes of opposite charge for fast assembly of a minimal electron transport chain consisting of F1F0 ATP-synthase and the proton pump bo3-oxidase.
- Robert R. Ishmukhametov
- , Aidan N. Russell
- & Richard M. Berry
-
Article
| Open Access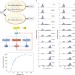 Noise reduction facilitated by dosage compensation in gene networks
Noise reduction facilitated by dosage compensation in gene networks
Cells must function despite the noisiness of their processes by tolerating or reducing such variability. Here, the authors combine experiment and modelling to show that a network motif that mediates network-dosage compensation also reduces noise in network output, suggesting that noise is tuneable.
- Weilin Peng
- , Ruijie Song
- & Murat Acar
-
Article
| Open Access Dichotomy of cellular inhibition by small-molecule inhibitors revealed by single-cell analysis
Dichotomy of cellular inhibition by small-molecule inhibitors revealed by single-cell analysis
Many drugs are small molecule inhibitors of cell signalling. Through single cell analysis and mathematical modelling here the authors show that cell-to-cell variability diversifies inhibition response into digital and analogue, and that the two translate into distinct long-term functional responses.
- Robert M. Vogel
- , Amir Erez
- & Grégoire Altan-Bonnet
-
Article
| Open Access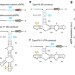 Twister ribozymes as highly versatile expression platforms for artificial riboswitches
Twister ribozymes as highly versatile expression platforms for artificial riboswitches
Twister ribozymes are small endonucleolytic RNA motifs. Here the authors develop twister ribozymes into RNA logic gates and cross-species synthetic genetic regulators.
- Michele Felletti
- , Julia Stifel
- & Jörg S. Hartig
-
Article
| Open Access Proteome-wide association studies identify biochemical modules associated with a wing-size phenotype in Drosophila melanogaster
Proteome-wide association studies identify biochemical modules associated with a wing-size phenotype in Drosophila melanogaster
How genetic diversity generates complex phenotypes along a continuum remains a fundamental question of biology. Here—applying the emerging SWATH proteomics technology—the authors describe a proteome wide association study (PWAS) of Drosophila wing size and identify functional protein clusters associated with this trait.
- Hirokazu Okada
- , H. Alexander Ebhardt
- & Ernst Hafen
-
Article
| Open Access Integrative proteomic profiling of ovarian cancer cell lines reveals precursor cell associated proteins and functional status
Integrative proteomic profiling of ovarian cancer cell lines reveals precursor cell associated proteins and functional status
High-grade serous ovarian cancer is the most common and aggressive ovarian cancer, with uncertain cell of origin. Here, the authors undertake a mass spectrometric analysis of 26 cancer cell lines and identify a protein signature that classifies ovarian cancer tissues into epithelial and mesenchymal groups.
- F. Coscia
- , K. M. Watters
- & M. Mann
-
Article
| Open Access A rheostat mechanism governs the bifurcation of carbon flux in mycobacteria
A rheostat mechanism governs the bifurcation of carbon flux in mycobacteria
Microbes survive in dynamic environments by modulating their intracellular metabolism. Here, the authors reveal that mycobacteria employ a rheostat-like mechanism to regulate carbon flux between the oxidative TCA cycle and the glyoxylate shunt during glucose-acetate diauxic shift.
- Paul Murima
- , Michael Zimmermann
- & John D. McKinney
-
Article
| Open Access Intrinsic limits to gene regulation by global crosstalk
Intrinsic limits to gene regulation by global crosstalk
Limited specificity of transcription factor-DNA interactions leads to crosstalk in gene regulation. Here the authors consider global crosstalk in regulatory networks of growing size and complexity, and show that it imposes constraints on gene regulation and on the evolution of regulatory networks.
- Tamar Friedlander
- , Roshan Prizak
- & Gašper Tkačik
-
Article
| Open Access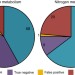 Metabolic modelling reveals the specialization of secondary replicons for niche adaptation in Sinorhizobium meliloti
Metabolic modelling reveals the specialization of secondary replicons for niche adaptation in Sinorhizobium meliloti
The genome of some bacteria consists of two or more chromosomes or replicons. Here, diCenzo et al. integrate genome-scale metabolic modelling and growth data from a collection of mutants of the plant symbiont Sinorhizobium melilotito estimate the fitness contribution of each replicon in three environments.
- George C. diCenzo
- , Alice Checcucci
- & Marco Fondi
-
Article
| Open Access Glycolytic regulation of cell rearrangement in angiogenesis
Glycolytic regulation of cell rearrangement in angiogenesis
Glycolytic regulator PFKFB3 is a key player in vessel sprouting. Here the authors develop a computational model predicting that PFKFB3 drives endothelial cell rearrangement during vessel sprouting by promoting filopodia formation and reducing intercellular adhesion, and empirically validate this prediction.
- Bert Cruys
- , Brian W. Wong
- & Peter Carmeliet
Browse broader subjects
Browse narrower subjects
- Bayesian inference
- Biobricks
- Biochemical networks
- Bioenergetics
- Cellular noise
- Complexity
- Computer modelling
- Computer science
- Control theory
- Criticality
- Differential equations
- DNA computing and cryptography
- Dynamic networks
- Dynamical systems
- Emergence
- Evolvability
- Genetic circuit engineering
- Genetic interaction
- Genomic engineering
- Information theory
- Logic gates
- Metabolic engineering
- Modularity
- Molecular engineering
- Molecular fluctuations
- Multicellular systems
- Multistability
- Nonlinear dynamics
- Numerical simulations
- Oscillators
- Population dynamics
- Programming language
- Protein engineering
- Regulatory networks
- Reverse engineering
- Robustness
- Signal processing
- Single-cell imaging
- Software
- Standardization
- Stochastic modelling
- Stochastic networks
- Synthetic biology
- Systems analysis
- Time series

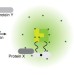 Split-BioID a conditional proteomics approach to monitor the composition of spatiotemporally defined protein complexes
Split-BioID a conditional proteomics approach to monitor the composition of spatiotemporally defined protein complexes