Featured
-
-
Article
| Open Access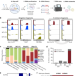 In vivo RNA interactome profiling reveals 3’UTR-processed small RNA targeting a central regulatory hub
In vivo RNA interactome profiling reveals 3’UTR-processed small RNA targeting a central regulatory hub
Here the authors report a new approach to profile RNA-RNA interactions in live bacterial cells. The charted RNA interaction networks unveil a key mRNA regulatory hub targeted by twelve small RNAs, including a novel RNA involved in fatty acid metabolism.
- Fang Liu
- , Ziying Chen
- & Yanjie Chao
-
Article
| Open Access Paired yeast one-hybrid assays to detect DNA-binding cooperativity and antagonism across transcription factors
Paired yeast one-hybrid assays to detect DNA-binding cooperativity and antagonism across transcription factors
Combinations of transcription factors (TFs) bind DNA to fine-tune gene expression. Here, the authors map cooperative and antagonistic DNA binding across hundreds of TF-pairs. TF-TF relationships vary depending on DNA targets and TF isoforms.
- Anna Berenson
- , Ryan Lane
- & Juan I. Fuxman Bass
-
Article
| Open Access Deep mutational scanning reveals the molecular determinants of RNA polymerase-mediated adaptation and tradeoffs
Deep mutational scanning reveals the molecular determinants of RNA polymerase-mediated adaptation and tradeoffs
Mutations in an RNA polymerase fragment, frequently found in lab adaptation, cluster in two modules favoring growth or maintenance via loss of interactions. Combining mutations in both modules enhances both traits, promoting compensatory evolution.
- Alaksh Choudhury
- , Benoit Gachet
- & Olivier Tenaillon
-
Article
| Open Access Genome-wide promoter responses to CRISPR perturbations of regulators reveal regulatory networks in Escherichia coli
Genome-wide promoter responses to CRISPR perturbations of regulators reveal regulatory networks in Escherichia coli
Measuring gene expression responses for every transcription factor (TF)-gene pair in living prokaryotic cells is challenging. Here the authors report pooled promoter responses to TF perturbation sequencing (PPTP-seq) using CRISPRi, which they use to address this problem in E. coli.
- Yichao Han
- , Wanji Li
- & Fuzhong Zhang
-
Article
| Open Access Sequestration of histidine kinases by non-cognate response regulators establishes a threshold level of stimulation for bacterial two-component signaling
Sequestration of histidine kinases by non-cognate response regulators establishes a threshold level of stimulation for bacterial two-component signaling
Bacterial two-component systems consist of a sensor histidine kinase (HK) that perceives a signal, and a cognate response regulator (RR) that modulates target gene expression. Here, the authors combine experiments and mathematical modelling to show that phosphorylated HKs can be sequestered by non-cognate RRs, which prevents responses to weak signals.
- Gaurav D. Sankhe
- , Rubesh Raja
- & Deepak Kumar Saini
-
Article
| Open Access Salicylic acid metabolism and signalling coordinate senescence initiation in aspen in nature
Salicylic acid metabolism and signalling coordinate senescence initiation in aspen in nature
Deciduous trees exhibit autumn senescence driven by environmental seasonality. Here, the authors show that senescence timing in aspen tree genotypes depends on environmental changes but also on the ability of each genotype to sustain stress tolerance mediated by the phytohormone salicylic acid.
- Jenna Lihavainen
- , Jan Šimura
- & Stefan Jansson
-
Article
| Open Access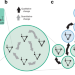 Robustness and innovation in synthetic genotype networks
Robustness and innovation in synthetic genotype networks
Genotype networks are sets of genotypes connected by small mutational changes that share the same phenotype. Here the authors combine construction of over 20 synthetic gene regulatory networks with mathematical modeling to exemplify how gene regulatory networks provide robustness in face of mutations while enabling transitions to innovative phenotypes.
- Javier Santos-Moreno
- , Eve Tasiudi
- & Yolanda Schaerli
-
Article
| Open Access Transposable elements orchestrate subgenome-convergent and -divergent transcription in common wheat
Transposable elements orchestrate subgenome-convergent and -divergent transcription in common wheat
How subgenome-divergent and -convergent transcription is mediated and harmonized in hexaploid common wheat genome remains unclear. Here, via characterizing the cistrome maps, the authors reveal that transposon elements with transcription factor binding ability have the potential to make the contribution.
- Yuyun Zhang
- , Zijuan Li
- & Yijing Zhang
-
Article
| Open Access Systematic characterization of cancer transcriptome at transcript resolution
Systematic characterization of cancer transcriptome at transcript resolution
Modification of transcribed mRNAs enables regulation of transcription but its extent in cancer cells is incompletely understood. Here, the authors analyse transcript assembly in over 1000 cancer cell lines and find unannotated transcripts are common, and are associated with drug sensitivity.
- Wei Hu
- , Yangjun Wu
- & Shengli Li
-
Article
| Open Access Proteome effects of genome-wide single gene perturbations
Proteome effects of genome-wide single gene perturbations
Protein abundance is controlled at the transcriptional, translational and posttranslational levels. Here, Öztürk et al. determine proteome changes resulting from individual knockout of 3308 nonessential genes in the yeast S. pombe, infer gene functionality, and show that protein upregulation under stable transcript expression utilizes optimal codons.
- Merve Öztürk
- , Anja Freiwald
- & Falk Butter
-
Article
| Open Access HOXA9 has the hallmarks of a biological switch with implications in blood cancers
HOXA9 has the hallmarks of a biological switch with implications in blood cancers
HOXA9 plays an important role in acute myeloid leukaemia (AML), but its relevance for other blood malignancies is unclear. Here, the authors show that HOXA9 has a binary switch function that can clinically stratify AML patients, and model how the interactions with JAK2, TET2 and NOTCH impact myeloproliferative neoplasms.
- Laure Talarmain
- , Matthew A. Clarke
- & Benjamin A. Hall
-
Article
| Open Access Artificial neural networks enable genome-scale simulations of intracellular signaling
Artificial neural networks enable genome-scale simulations of intracellular signaling
Many diseases are caused by disruptions to the network of biochemical reactions that allow cells to respond to external signals. Here Nilsson et al develop a method to simulate cellular signaling using artificial neural networks to predict cellular responses and activities of signaling molecules.
- Avlant Nilsson
- , Joshua M. Peters
- & Douglas A. Lauffenburger
-
Article
| Open Access Machine learning aided construction of the quorum sensing communication network for human gut microbiota
Machine learning aided construction of the quorum sensing communication network for human gut microbiota
Microbes communicate with each other by Quorum sensing (QS) languages. Here the authors construct a QS database and the QS communication network to decipher intricate QSbased communications and form one of the key knowledge maps for human gut microbiota.
- Shengbo Wu
- , Jie Feng
- & Jianjun Qiao
-
Article
| Open Access Characterisation of a nucleo-adhesome
Characterisation of a nucleo-adhesome
Cell adhesion proteins have been described at sites away from the cell surface, including in the nucleus. Here, the authors report the scale of nuclear localisation of adhesion proteins, establishing a nucleo-adhesome and showing that nuclear adhesion proteins can cooperate to control transcription.
- Adam Byron
- , Billie G. C. Griffith
- & Margaret C. Frame
-
Article
| Open Access Visual barcodes for clonal-multiplexing of live microscopy-based assays
Visual barcodes for clonal-multiplexing of live microscopy-based assays
Multiplex analyses of samples allow understanding complex processes in cancer initiation, progression and therapy response. Here, the authors present a fluorescence imaging-based visual barcode for livecell clonal-multiplexing which allows identifying signalling pathways clusters in response to different chemotherapy compounds.
- Tom Kaufman
- , Erez Nitzan
- & Ravid Straussman
-
Article
| Open Access Integrative network analysis of early-stage lung adenocarcinoma identifies aurora kinase inhibition as interceptor of invasion and progression
Integrative network analysis of early-stage lung adenocarcinoma identifies aurora kinase inhibition as interceptor of invasion and progression
The molecular factors that drive invasiveness and metastasis in lung adenocarcinoma (LUAD) are not completely understood. Here, the authors use an integrative network approach to identify a gene signature of invasiveness in LUAD, and reveal Aurora kinases as master regulators of invasion.
- Seungyeul Yoo
- , Abhilasha Sinha
- & Charles A. Powell
-
Article
| Open Access Human transcription factor protein interaction networks
Human transcription factor protein interaction networks
Transcription factors (TFs) interact with several other proteins in the process of transcriptional regulation. Here the authors identify 6703 and 1536 protein–protein interactions for 109 different human TFs through BioID and AP-MS analyses, respectively.
- Helka Göös
- , Matias Kinnunen
- & Markku Varjosalo
-
Article
| Open Access Network analysis reveals rare disease signatures across multiple levels of biological organization
Network analysis reveals rare disease signatures across multiple levels of biological organization
Despite the clear causal relationship between genotype and phenotype in rare diseases, identifying the pathobiological mechanisms at various levels of biological organization remains a practical and conceptual challenge. Here, the authors introduce a network approach for evaluating the impact of rare gene defects across biological scales.
- Pisanu Buphamalai
- , Tomislav Kokotovic
- & Jörg Menche
-
Article
| Open Access Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr
Single-cell normalization and association testing unifying CRISPR screen and gene co-expression analyses with Normalisr
Normalisr removes technical bias in single-cell RNA-seq and detects gene differential and coexpression accurately and efficiently. It also infers gene regulatory and co-expression networks from conventional and CRISPR screen single-cell RNA-seq datasets.
- Lingfei Wang
-
Article
| Open Access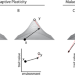 Selection on adaptive and maladaptive gene expression plasticity during thermal adaptation to urban heat islands
Selection on adaptive and maladaptive gene expression plasticity during thermal adaptation to urban heat islands
Anthropogenic change, such as urban heat islands, present challenges to biodiversity that can be overcome through phenotypic plasticity. Unlike their ancestral counterparts, urban lizards have fewer maladaptive gene expression responses to higher temperatures in a common garden experiment, suggesting the evolution of adaptive plasticity.
- Shane C. Campbell-Staton
- , Jonathan P. Velotta
- & Kristin M. Winchell
-
Article
| Open Access Integrated omics networks reveal the temporal signaling events of brassinosteroid response in Arabidopsis
Integrated omics networks reveal the temporal signaling events of brassinosteroid response in Arabidopsis
Brassinosteroids (BR) regulate plant development and stress responses. Here, by integrating multiple omics datasets and inferring networks, the authors profile BR signaling in Arabidopsis and characterize BRONTOSAURUS, a BR-regulated transcription factor that impacts cell division in roots.
- Natalie M. Clark
- , Trevor M. Nolan
- & Justin W. Walley
-
Article
| Open Access VEGA is an interpretable generative model for inferring biological network activity in single-cell transcriptomics
VEGA is an interpretable generative model for inferring biological network activity in single-cell transcriptomics
Developing interpretable models is a major challenge in single cell deep learning. Here we show that the VEGA variational autoencoder model, whose decoder wiring mirrors gene modules, can provide direct interpretability to the latent space further enabling the inference of biological module activity.
- Lucas Seninge
- , Ioannis Anastopoulos
- & Joshua Stuart
-
Article
| Open Access Rapid prototyping and design of cybergenetic single-cell controllers
Rapid prototyping and design of cybergenetic single-cell controllers
Practical implementation of genetic circuits is difficult due to low predictability and time-intensive troubleshooting. Here the authors present Cyberloop, which interfaces a computer with single cells to enable cell-in-the-loop testing and optimization of circuit designs before they are built.
- Sant Kumar
- , Marc Rullan
- & Mustafa Khammash
-
Article
| Open Access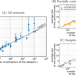 The basis of easy controllability in Boolean networks
The basis of easy controllability in Boolean networks
Boolean networks allow a simplified representation of interactions. Here, the authors systematically analyze regulation in dozens of biological Boolean networks, finding mathematical regularities that suggest biological systems could be controlled through a relatively small number of components.
- Enrico Borriello
- & Bryan C. Daniels
-
Article
| Open Access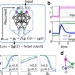 Finding gene network topologies for given biological function with recurrent neural network
Finding gene network topologies for given biological function with recurrent neural network
Networks are useful ways to describe interactions between molecules in a cell, but predicting the real topology of large networks can be challenging. Here, the authors use deep learning to predict the topology of networks that perform biologically-plausible functions.
- Jingxiang Shen
- , Feng Liu
- & Chao Tang
-
Article
| Open Access Inference and analysis of cell-cell communication using CellChat
Inference and analysis of cell-cell communication using CellChat
Single-cell methods record molecule expressions of cells in a given tissue, but understanding interactions between cells remains challenging. Here the authors show by applying systems biology and machine learning approaches that they can infer and analyze cell-cell communication networks in an easily interpretable way.
- Suoqin Jin
- , Christian F. Guerrero-Juarez
- & Qing Nie
-
Article
| Open Access An integrative multiomic network model links lipid metabolism to glucose regulation in coronary artery disease
An integrative multiomic network model links lipid metabolism to glucose regulation in coronary artery disease
Some cholesterol-lowering drugs can increase the risk of type 2 diabetes, but the mechanism behind this is not fully understood. Here the authors show that there is a single genetic regulatory module that influences both cholesterol levels and glucose levels, providing a link between cholesterol levels and diabetes.
- Ariella T. Cohain
- , William T. Barrington
- & Eric E. Schadt
-
Article
| Open Access Exploring the effect of network topology, mRNA and protein dynamics on gene regulatory network stability
Exploring the effect of network topology, mRNA and protein dynamics on gene regulatory network stability
Maintaining protein expression levels is essential to cellular homeostasis. Here, the authors investigate how transcription factors affect the stability of protein expression in a gene regulatory network, and highlight the importance of network topology.
- Yipei Guo
- & Ariel Amir
-
Article
| Open Access Robustness and lethality in multilayer biological molecular networks
Robustness and lethality in multilayer biological molecular networks
Robustness is a prominent feature of most biological systems, but most of the current efforts have been focused on studying homogeneous molecular networks. Here the authors propose a comprehensive framework for understanding how the interactions between genes, proteins, and metabolites contribute to the determinants of robustness.
- Xueming Liu
- , Enrico Maiorino
- & Amitabh Sharma
-
Article
| Open Access Stress-induced RNA–chromatin interactions promote endothelial dysfunction
Stress-induced RNA–chromatin interactions promote endothelial dysfunction
Global interaction of chromatin-associated RNAs and DNA can be identified in situ. Here the authors report the genome-wide increase of interchromosomal RNA-DNA interactions and demonstrate the importance of such RNA-DNA contacts exemplified by LINC00607 RNA and SERPINE1 gene’s super enhancer in dysfunctional endothelial cell models.
- Riccardo Calandrelli
- , Lixia Xu
- & Sheng Zhong
-
Article
| Open Access Predicting cell-to-cell communication networks using NATMI
Predicting cell-to-cell communication networks using NATMI
Single cell expression data allows for inferring cell-cell communication between cells expressing ligands and those expressing their cognate receptors. Here the authors present an updated and curated database of ligand-receptor pairs and a Python-based toolkit to construct and analyse communication networks from single cell and bulk expression data.
- Rui Hou
- , Elena Denisenko
- & Alistair R. R. Forrest
-
Article
| Open Access Reconciling qualitative, abstract, and scalable modeling of biological networks
Reconciling qualitative, abstract, and scalable modeling of biological networks
Boolean Networks are a well-established model of biological networks, but usual interpretations can preclude the prediction of behaviours observed in quantitative systems. The authors introduce Most Permissive Boolean Networks, which are shown not to miss any behaviour achievable by the corresponding quantitative model.
- Loïc Paulevé
- , Juri Kolčák
- & Stefan Haar
-
Article
| Open Access Design of a MAPK signalling cascade balances energetic cost versus accuracy of information transmission
Design of a MAPK signalling cascade balances energetic cost versus accuracy of information transmission
Cellular signalling networks provide information to the cell, but the trade-off between accuracy of information transfer and energetic cost of doing so has not been assessed. Here, the authors investigate a MAPK signalling cascade in budding yeast and find that information is maximised per unit energetic cost.
- Alexander Anders
- , Bhaswar Ghosh
- & Victor Sourjik
-
Article
| Open Access A network of RNA-binding proteins controls translation efficiency to activate anaerobic metabolism
A network of RNA-binding proteins controls translation efficiency to activate anaerobic metabolism
mRNA translation efficiency is regulated in response to stimuli. Here the authors employ mass spectrometry analysis of ribosome fractions and show that under hypoxia, oxygen-sensitive RNA binding proteins enhance the translation efficiency of glycolysis pathway transcripts.
- J. J. David Ho
- , Nathan C. Balukoff
- & Stephen Lee
-
Article
| Open Access The Escherichia coli transcriptome mostly consists of independently regulated modules
The Escherichia coli transcriptome mostly consists of independently regulated modules
Mechanistic insight into the regulation of transcriptional modules remains scarce. Here, the authors identify statistically independent gene sets by applying independent component analysis to a high-quality E. coli RNA-seq data compendium and find that most gene sets represent the effects of specific transcriptional regulators.
- Anand V. Sastry
- , Ye Gao
- & Bernhard O. Palsson
-
Article
| Open Access The landscape of multiscale transcriptomic networks and key regulators in Parkinson’s disease
The landscape of multiscale transcriptomic networks and key regulators in Parkinson’s disease
Parkinson’s disease (PD) is characterized by neurodegeneration associated with loss of dopaminergic (DA) neurons and deposition of Lewy bodies. Here, Wang et al. use co-expression network analysis to pinpoint disease pathways and propose reduced expression of STMN2 as a cause of presynaptic function loss in PD.
- Qian Wang
- , Yuanxi Zhang
- & Bin Zhang
-
Article
| Open Access Mapping the perturbome network of cellular perturbations
Mapping the perturbome network of cellular perturbations
Our understanding of the mechanisms of drug interactions remains limited. Here the authors introduce a framework to study how complex cellular perturbations induced by different drugs affect each other in morphological feature space.
- Michael Caldera
- , Felix Müller
- & Jörg Menche
-
Article
| Open Access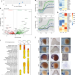 A genome-wide assessment of the ancestral neural crest gene regulatory network
A genome-wide assessment of the ancestral neural crest gene regulatory network
An understanding of the ancestral state of the neural crest (NC) gene regulatory network (GRN) gives insight into vertebrate evolution. Here, the authors use transcriptomic and chromatin accessibility analyses of the lamprey NC, as well as cross-species enhancer assays, to identify GRN elements conserved throughout vertebrates.
- Dorit Hockman
- , Vanessa Chong-Morrison
- & Tatjana Sauka-Spengler
-
Article
| Open Access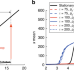 Transient hysteresis and inherent stochasticity in gene regulatory networks
Transient hysteresis and inherent stochasticity in gene regulatory networks
Cell fate commitment is understood in terms of bistable regulatory circuits with hysteresis, but inherent stochasticity in gene expression is incompatible with hysteresis. Here, the authors quantify how, under slow dynamics, the dependency of the non-stationary solutions on the initial state of the cells can lead to transient hysteresis.
- M. Pájaro
- , I. Otero-Muras
- & A. A. Alonso
-
Article
| Open Access Systematic identification of metabolites controlling gene expression in E. coli
Systematic identification of metabolites controlling gene expression in E. coli
Interactions between metabolites and transcription factors are known to control gene expression but analyzing these events at genome-scale is challenging. Here, the authors integrate dynamic metabolome and transcriptome data from E.coli to predict regulatory metabolite-transcription factor interactions.
- Martin Lempp
- , Niklas Farke
- & Hannes Link
-
Article
| Open Access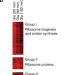 Integrated TORC1 and PKA signaling control the temporal activation of glucose-induced gene expression in yeast
Integrated TORC1 and PKA signaling control the temporal activation of glucose-induced gene expression in yeast
Yeast cells respond to nutrients by altering expression of the protein synthesis genes and thus their growth rate. Here, the authors use microarrays to show that TORC1 controls gene expression during steady state growth while PKA speeds up expression changes when nutrient levels change.
- Joseph Kunkel
- , Xiangxia Luo
- & Andrew P. Capaldi
-
Article
| Open Access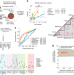 Metabolic profiling of cancer cells reveals genome-wide crosstalk between transcriptional regulators and metabolism
Metabolic profiling of cancer cells reveals genome-wide crosstalk between transcriptional regulators and metabolism
Aberrant gene expression in cancer coincides with drastic changes in metabolism. Here, the authors combined metabolome, transcriptome and proteome data in 54 cancer cell lines to uncover a genome-scale network of associations between transcriptional regulators and metabolites.
- Karin Ortmayr
- , Sébastien Dubuis
- & Mattia Zampieri
-
Article
| Open Access Comparative transcriptomics of social insect queen pheromones
Comparative transcriptomics of social insect queen pheromones
Queen pheromones are used by eusocial insects to regulate all aspects of colony life. Here, Holman et al. compare the effects of queen pheromone on gene expression and splicing in four eusocial insect species, giving insight into the mechanism and evolution of division of reproductive labour.
- Luke Holman
- , Heikki Helanterä
- & Alexander S. Mikheyev
-
Article
| Open Access Network Walking charts transcriptional dynamics of nitrogen signaling by integrating validated and predicted genome-wide interactions
Network Walking charts transcriptional dynamics of nitrogen signaling by integrating validated and predicted genome-wide interactions
Temporal control of transcriptional networks enables organisms to adapt to changing environment. Here, the authors use a scaled-up cell-based assay to identify direct targets of nitrogen-early responsive transcription factors and validate a network path mediating dynamic nitrogen signaling in Arabidopsis.
- Matthew D. Brooks
- , Jacopo Cirrone
- & Gloria M. Coruzzi
-
Article
| Open Access Optogenetic dissection of Rac1 and Cdc42 gradient shaping
Optogenetic dissection of Rac1 and Cdc42 gradient shaping
A steep gradient of Cdc42 is at the front of migrating cells, whereas the active Rac1 gradient is graded. Here the authors show that Cdc42 gradients follow the distribution of GEFs and govern direction of migration, while Rac1 gradients require the activity of the GAP β2-chimaerin and control cell speed.
- S. de Beco
- , K. Vaidžiulytė
- & M. Coppey
-
Article
| Open Access Systems biology approach reveals a link between mTORC1 and G2/M DNA damage checkpoint recovery
Systems biology approach reveals a link between mTORC1 and G2/M DNA damage checkpoint recovery
DNA damage induces checkpoints to ensure that damage is not transferred to the next generation, but the molecular pathways responsible for checkpoint recovery are not clear. Here the authors show that the nutrient sensor mTORC1 is a determinant for G2/M checkpoint recovery through regulation of cyclin B1 and PLK1 expression.
- Hui-Ju Hsieh
- , Wei Zhang
- & Guang Peng
-
Article
| Open Access Temporal genetic association and temporal genetic causality methods for dissecting complex networks
Temporal genetic association and temporal genetic causality methods for dissecting complex networks
Temporal omics data have the potential to dissect complex biological networks. Here the authors develop methods for detecting temporal genetic loci (teQTLs) of quantitative traits monitored over time and inferring causal relationships between traits linked to the locus.
- Luan Lin
- , Quan Chen
- & Jun Zhu
-
Article
| Open Access Linear mapping approximation of gene regulatory networks with stochastic dynamics
Linear mapping approximation of gene regulatory networks with stochastic dynamics
The intractability of most stochastic models of gene regulatory networks (GRNs) limits their utility. Here, the authors present a linear-mapping approximation mapping models onto simpler ones, giving approximate but accurate analytic or semi- analytic solutions for a wide range of model GRNs.
- Zhixing Cao
- & Ramon Grima
-
Article
| Open Access Lineage marker synchrony in hematopoietic genealogies refutes the PU.1/GATA1 toggle switch paradigm
Lineage marker synchrony in hematopoietic genealogies refutes the PU.1/GATA1 toggle switch paradigm
The timing of cell fate choices is usually unknown, because we have to rely on indirect evidence of their molecular basis. Here, the authors introduce a method to infer decision times from marker onset in cell genealogies, and find evidence refuting the paradigmatic PU.1/GATA1 cell fate switch.
- Michael K. Strasser
- , Philipp S. Hoppe
- & Carsten Marr

 Deciphering driver regulators of cell fate decisions from single-cell transcriptomics data with CEFCON
Deciphering driver regulators of cell fate decisions from single-cell transcriptomics data with CEFCON