Featured
-
-
Books & Arts |
 Epigenetics: Beyond face value
Epigenetics: Beyond face value
Jonathan Weitzman relishes two accounts of how environment can influence the script of our genome.
- Jonathan Weitzman
-
Letter |
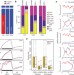 Spontaneous epigenetic variation in the Arabidopsis thaliana methylome
Spontaneous epigenetic variation in the Arabidopsis thaliana methylome
- Claude Becker
- , Jörg Hagmann
- & Detlef Weigel
-
Letter |
 The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes
The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes
- Tian-Peng Gu
- , Fan Guo
- & Guo-Liang Xu
-
News & Views |
 Imprinting in the brain
Imprinting in the brain
A gene is considered to be imprinted if only the copy inherited from the mother or from the father is expressed throughout life. But one imprinted gene, Dlk1, disobeys this rule during postnatal neurodevelopment. See Letter p.381
- Edwin C. Oh
- & Nicholas Katsanis
-
Letter |
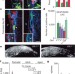 Postnatal loss of Dlk1 imprinting in stem cells and niche astrocytes regulates neurogenesis
Postnatal loss of Dlk1 imprinting in stem cells and niche astrocytes regulates neurogenesis
- Sacri R. Ferrón
- , Marika Charalambous
- & Anne C. Ferguson-Smith
-
Research Highlights |
Stressed genes are inherited
-
Letter |
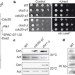 Coordination of DNA replication and histone modification by the Rik1–Dos2 complex
Coordination of DNA replication and histone modification by the Rik1–Dos2 complex
- Fei Li
- , Rob Martienssen
- & W. Zacheus Cande
-
Letter |
 XUTs are a class of Xrn1-sensitive antisense regulatory non-coding RNA in yeast
XUTs are a class of Xrn1-sensitive antisense regulatory non-coding RNA in yeast
- E. L. van Dijk
- , C. L. Chen
- & A. Morillon
-
News & Views |
Tet proteins in the limelight
Tet proteins mediate the hydroxymethylation of DNA. New work reveals their function in gene regulation and the extent of their activity throughout the genome of embryonic stem cells. See Article p.343 & Letters p.389, p.394 & p.398
- Nathalie Véron
- & Antoine H. F. M. Peters
-
Article |
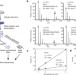 Global quantification of mammalian gene expression control
Global quantification of mammalian gene expression control
- Björn Schwanhäusser
- , Dorothea Busse
- & Matthias Selbach
-
Article |
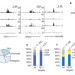 TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity
TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity
- Kristine Williams
- , Jesper Christensen
- & Kristian Helin
-
Letter |
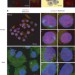 Eutherian mammals use diverse strategies to initiate X-chromosome inactivation during development
Eutherian mammals use diverse strategies to initiate X-chromosome inactivation during development
- Ikuhiro Okamoto
- , Catherine Patrat
- & Edith Heard
-
Letter |
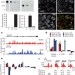 Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation
Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation
- Gabriella Ficz
- , Miguel R. Branco
- & Wolf Reik
-
Letter |
 Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells
Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells
- Hao Wu
- , Ana C. D’Alessio
- & Yi Zhang
-
Outlook |
 Epigenetics: Unravelling the cancer code
Epigenetics: Unravelling the cancer code
There's more to the genetic causes of cancer than sequence mutations. This added complexity could offer scientists an opportunity to tackle cancer even earlier.
- Vicki Brower
-
Letter |
 X chromosome dosage compensation via enhanced transcriptional elongation in Drosophila
X chromosome dosage compensation via enhanced transcriptional elongation in Drosophila
Different organisms use a variety of mechanisms to compensate for X chromosome dosage imbalance between the sexes. In Drosophila, the MSL complex increases transcription on the single X chromosome of males and is thought to regulate transcription elongation, although mechanistic details have been unclear. Here, a global run-on sequencing technique is used to reveal that the MSL complex seems to enhance transcription by facilitating the progression of RNA polymerase II across the bodies of active X linked genes. In this way, MSL can impose dosage compensation on diverse genes with a wide range of transcription levels along the X chromosome.
- Erica Larschan
- , Eric P. Bishop
- & Mitzi I. Kuroda
-
News |
 Flaw in induced-stem-cell model
Flaw in induced-stem-cell model
Adult cells do not fully convert to embryonic-like state.
- Elie Dolgin
-
Letter |
 Distinct physiological and behavioural functions for parental alleles of imprinted Grb10
Distinct physiological and behavioural functions for parental alleles of imprinted Grb10
Genetic imprinting, or the preferential expression of a single parental allele, has typically been implicated as an influential factor during development, but whether unequal representation of one allele can influence social behaviour has not been studied. Here, it is demonstrated that the adaptor protein Grb10 is predominantly expressed from the paternal allele in brain and that ablating this allelic bias induces behavioural modifications of a social nature. At this time, Grb10 is unique in the sense that tissue-specific actions of each parental allele can influence distinct physiological or behavioural processes.
- Alastair S. Garfield
- , Michael Cowley
- & Andrew Ward
-
Outlook |
 Epigenetics: Tales of adversity
Epigenetics: Tales of adversity
Genetic studies of people conceived during famine reveals that prenatal malnutrition lingers long after the event.
- Farooq Ahmed
-
Letter |
 A unique chromatin signature uncovers early developmental enhancers in humans
A unique chromatin signature uncovers early developmental enhancers in humans
Identifying the genomic regulatory sequences, such as enhancers, that control early embryonic development remains a difficult challenge. Here, profiling of histone modifications and chromatin regulators in human embryonic stem cells (hESCs) reveals unique signatures that are used to identify over 2,000 putative enhancers. These enhancers are either active in the h ESCs or associated with early developmental genes.
- Alvaro Rada-Iglesias
- , Ruchi Bajpai
- & Joanna Wysocka
-
Letter |
 Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2
Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2
The TET family of enzymes convert 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC) in DNA. Mutations in the gene encoding TET2 are frequently observed in myeloid malignancies. Here it is shown that these disease-associated mutations compromise TET2 catalytic activity; bone marrow samples from patients with TET2 mutations have low levels of 5hmC in genomic DNA, and TET2 is required for normal haematopoietic differentiation. Measurement of genomic 5hmC levels may prove valuable as a diagnostic tool in myeloid cancers.
- Myunggon Ko
- , Yun Huang
- & Anjana Rao
-
Research Highlights |
 Epigenetics: What makes a queen bee?
Epigenetics: What makes a queen bee?
-
News & Views |
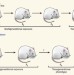 Fathers' nutritional legacy
Fathers' nutritional legacy
A female can develop a diabetes-like disease due to a high fat content in her father's diet before she was conceived. Epigenetic modifications of the father's sperm DNA might underlie this peculiar observation. See Letter p.963
- Michael K. Skinner
-
News |
 Fat fathers affect daughters' health
Fat fathers affect daughters' health
Female offspring of male rats on bad diets are more likely to develop diabetes-like disease.
- Geoff Marsh
-
-
Research Highlights |
Epigenetics: Mapping methylation
-
News & Views |
 Troublesome memories
Troublesome memories
Methods for generating embryonic-like stem cells have been established. The focus now is on finding ways to coax these cells into matching their natural counterparts as closely as possible, should this be desired.
- Thomas P. Zwaka
-
Letter |
 Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification
Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification
TET1 is an enzyme that catalyses the conversion of 5-methylcytosine of DNA to 5-hydroxymethylcytosine, raising the possibility that it is involved in mediating DNA demethylation. These authors show that Tet1 is involved in mouse embryonic stem cell maintenance and specification of the inner cell mass. It is required to maintain both the expression of Nanog in embryonic stem cells and the Nanog promoter in a hypomethylated state, supporting a role for Tet1 in regulating DNA methylation.
- Shinsuke Ito
- , Ana C. D’Alessio
- & Yi Zhang
-
Article |
 Heterochromatin silencing of p53 target genes by a small viral protein
Heterochromatin silencing of p53 target genes by a small viral protein
Adenovirus E1B-55k targets transcription factor p53 for degradation and is thought to be critical for p53 inactivation during adenovirus replication. Indeed, mutant viruses lacking E1B-55k have been tested as viral cancer therapies selective for p53-positive tumours. These authors find another adenoviral protein, E4-ORF3, to inactivate p53 independently of E1B-55k by means of a chromatin-silencing mechanism that prevents access of p53 to its DNA target sites.
- Conrado Soria
- , Fanny E. Estermann
- & Clodagh C. O’Shea
-
Letter |
 Trans-acting small RNA determines dominance relationships in Brassica self-incompatibility
Trans-acting small RNA determines dominance relationships in Brassica self-incompatibility
A diploid organism has two copies of each gene, one inherited from each parent. The expression levels of the two alleles can be biased by dominant/recessive relationships. In Brassica, self-incompatibility in pollen is determined by dominance relationships between the two alleles of the gene SP11; the recessive allele is methylated and hence silenced. Here it is shown that such methylation is controlled by a small non-coding RNA encoded in the flanking region of the dominant allele.
- Yoshiaki Tarutani
- , Hiroshi Shiba
- & Seiji Takayama
-
Letter |
 Comprehensive methylome map of lineage commitment from haematopoietic progenitors
Comprehensive methylome map of lineage commitment from haematopoietic progenitors
During haematopoiesis, multipotent progenitors differentiate into progressively restricted myeloid or lymphoid progenitors. A comprehensive genome-wide DNA methylation analysis of haematopoietic cell populations with well-characterized differentiation potentials reveals remarkable epigenetic plasticity during lymphoid and myeloid restriction.
- Hong Ji
- , Lauren I. R. Ehrlich
- & Andrew P. Feinberg
-
News & Views |
A mine of imprinted genes
Some genes exclusively express only their maternal or paternal copy. Studies of the brain extend the list of such imprinted genes by an order of magnitude, highlighting their spatial and temporal regulation.
- Eric B. Keverne
-
Letter |
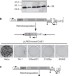 Epigenetic silencing of engineered L1 retrotransposition events in human embryonic carcinoma cells
Epigenetic silencing of engineered L1 retrotransposition events in human embryonic carcinoma cells
The ability of retrotransposons to mobilize and insert into genes presents a challenge to a cell needing to maintain its genomic integrity. These authors have studied retrotransposition in embryonic carcinoma-derived cells. On insertion into DNA, the retrotransposon is quickly silenced, but the retrotransposon-specificity of this process implies that multiple silencing mechanisms may exist. Once cells differentiate, the ability to silence newly introduced retrotransposons is lost but previously inactivated retrotransposons remain inactive.
- Jose L. Garcia-Perez
- , Maria Morell
- & John V. Moran
-
Letter |
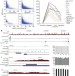 Conserved role of intragenic DNA methylation in regulating alternative promoters
Conserved role of intragenic DNA methylation in regulating alternative promoters
The methylation of DNA in 5′ promoter regions suppresses gene expression, but what is the role of DNA methylation in the bodies of genes? Here, a map of DNA methylation is generated from human brain tissue; it is found that most methylated CpG islands are within intragenic and intergenic regions, rather than within promoters. It is proposed that intragenic methylation regulates the expression of alternative gene transcripts in different tissues and cell types.
- Alika K. Maunakea
- , Raman P. Nagarajan
- & Joseph F. Costello
-
Research Highlights |
Genetics: Breaking the silence
-
-
Letter |
 Relationship between nucleosome positioning and DNA methylation
Relationship between nucleosome positioning and DNA methylation
Nucleosomes are composed of around 147 bases of DNA wrapped around an octamer of histone proteins. Here, a genome-wide analysis of nucleosome positioning in Arabidopsis thaliana has been combined with profiles of DNA methylation at single base resolution, revealing 10-base periodicities in the DNA methylation status of nucleosome-bound DNA. The results indicate that nucleosome positioning influences the pattern of DNA methylation throughout the genome.
- Ramakrishna K. Chodavarapu
- , Suhua Feng
- & Matteo Pellegrini
-
News |
 Genomics goes beyond DNA sequence
Genomics goes beyond DNA sequence
A technology that simultaneously reads a DNA sequence and its crucial modifications makes its debut.
- Alla Katsnelson
-
Letter |
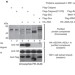 Histone H2A deubiquitinase activity of the Polycomb repressive complex PR-DUB
Histone H2A deubiquitinase activity of the Polycomb repressive complex PR-DUB
Polycomb group (PcG) proteins are transcriptional repressors that modify chromatin and regulate important developmental genes. One PcG-associated, chromatin-modifying activity is an enzyme that ubiquitinates histone H2A of chromatin. Here, a fruitfly PcG complex that is associated with H2A deubiquitination, and thereby with gene repression, is identified. PcG-mediated gene silencing might thus involve a dynamic balance between ubiquitination and deubiquitination of H2A.
- Johanna C. Scheuermann
- , Andrés Gaytán de Ayala Alonso
- & Jürg Müller
-
Article |
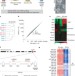 Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells
Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells
Induced pluripotent stem cells (iPSCs) are generated by the enforced expression of particular transcription factors in somatic cells. The extent to which such cells are equivalent to embryonic stem (ES) cells is an open question. Here, genetically identical mouse ES cells and iPSCs have been compared; the overall expression patterns of messenger RNAs and microRNAs are the same, with the exception of a few transcripts encoded within an imprinted gene cluster on chromosome 12qF1.
- Matthias Stadtfeld
- , Effie Apostolou
- & Konrad Hochedlinger
-
-
Letter |
 An RNA polymerase II- and AGO4-associated protein acts in RNA-directed DNA methylation
An RNA polymerase II- and AGO4-associated protein acts in RNA-directed DNA methylation
DNA methylation is an important epigenetic mark in many eukaryotes. In Arabidopsis plants, small interfering RNAs bound to the Argonaute 4 (AGO4) protein can direct de novo DNA methylation and consequent gene silencing. Here, a new regulator of RNA-directed DNA methylation has been discovered. This protein, RDM1, is proposed to bind to methylated DNA and to function in the AGO4 effector complex.
- Zhihuan Gao
- , Hai-Liang Liu
- & Jian-Kang Zhu
-
News |
 Mapping methylation's mysterious background
Mapping methylation's mysterious background
Analysis of 17 species fills in evolutionary history of DNA modification process.
- Alla Katsnelson
-
News |
 Cancer genes silenced in humans
Cancer genes silenced in humans
Tiny particles carrying short strands of RNA can interfere with protein production in tumours.
- Janet Fang
-
Letter |
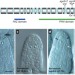 Control of female gamete formation by a small RNA pathway in Arabidopsis
Control of female gamete formation by a small RNA pathway in Arabidopsis
Female gametes in flowering plants develop from a meiotic division of a precursor cell followed by mitotic divisions of one of the resulting haploid cells to yield the gametophyte. Here, ARGONAUTE 9 (AGO9) — a protein involved in RNA interference — is identified as a factor required for specification of the gametophyte. AGO9 is found not in the cell destined to be the gametophyte, but in the neighbouring companion cells, suggesting that it functions in a non-cell-autonomous manner.
- Vianey Olmedo-Monfil
- , Noé Durán-Figueroa
- & Jean-Philippe Vielle-Calzada
-
Letter |
 Proviral silencing in embryonic stem cells requires the histone methyltransferase ESET
Proviral silencing in embryonic stem cells requires the histone methyltransferase ESET
Endogenous retroviruses (ERVs) are widely dispersed in mammalian genomes, and are silenced in somatic cells by DNA methylation. Here, an ERV silencing pathway independent of DNA methylation is shown to operate in embryonic stem cells. The pathway involves the histone H3K9 methyltransferase ESET and might be important for ERV silencing during the stages in embryogenesis when DNA methylation is reprogrammed.
- Toshiyuki Matsui
- , Danny Leung
- & Yoichi Shinkai
-
-
Letter |
 Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency
Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency
The extent of epigenetic reprogramming in mammalian primordial germ cells (PGCs) and in early embryos, and its molecular mechanisms, are poorly understood. DNA methylation profiling in PGCs now reveals a genome–wide erasure of methylation, with female PGCs being less methylated than male ones. A deficiency of the cytidine deaminase AID interferes with the genome–wide erasure of DNA methylation, indicating that AID has a critical function in epigenetic reprogramming.
- Christian Popp
- , Wendy Dean
- & Wolf Reik
-
Letter |
 DNMT1 maintains progenitor function in self-renewing somatic tissue
DNMT1 maintains progenitor function in self-renewing somatic tissue
Progenitor cells sustain the capacity of self-renewing tissues for proliferation while suppressing cell cycle exit and terminal differentiation. DNA methylation is one potential epigenetic mechanism for the cellular memory needed to preserve the somatic progenitor state through cell divisions. The DNA methyltransferase 1 and other regulators of DNA methylation are now shown to be essential for epidermal progenitor cell function.
- George L. Sen
- , Jason A. Reuter
- & Paul A. Khavari

 Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia
Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia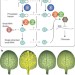 Small RNAs are on the move
Small RNAs are on the move Brain function and chromatin plasticity
Brain function and chromatin plasticity Why twins age differently
Why twins age differently