Abstract
Prominin 1 (PROM1) is one of a few clinically relevant progenitor markers in human alcoholic hepatitis (AH) and hepatocellular carcinoma (HCC), and mouse liver tumor initiating stem cell-like cells (TICs). However, the origin, fate and functions of PROM1+ cells in AH and HCC are unknown. Here we show by genetic lineage tracing that PROM1+ cells are derived in part from hepatocytes in AH and become tumor cells in mice with diethyl nitrosamine (DEN)-initiated, Western alcohol diet-promoted liver tumorigenesis. Our RNA sequencing analysis of mouse PROM1+ cells, reveals transcriptomic landscapes indicative of their identities as ductular reaction progenitors (DRPs) and TICs. Indeed, single-cell RNA sequencing reveals two subpopulations of Prom1+ Afp– DRPs and Prom1+ Afp+ TICs in the DEN-WAD model. Integrated bioinformatic analysis identifies Discodin Domain Receptor 1 (DDR1) as a uniquely upregulated and patient-relevant gene in PROM1+ cells in AH and HCC. Translational relevance of DDR1 is supported by its marked elevation in HCC which is inversely associated with patient survival. Further, knockdown of Ddr1 suppresses the growth of TICs and TIC-derived tumor growth in mice. These results suggest the importance of PROM1+ cells in the evolution of liver cancer and DDR1 as a potential driver of this process.
Similar content being viewed by others
Introduction
Liver progenitor cells (LPCs) proliferate after massive parenchymal loss or/and suppressed hepatocyte proliferation in different liver diseases1. However, the origin and fate of these cells in liver injury and cancer development are unclear2. LPCs are identified by immunophenotype markers such as A6, SOX9, KRT19 (CK19), EpCAM and PROM1 and generated from biliary epithelial cells or hepatocytes through hepatocyte-to-ductal metaplasia3. Tumor-initiating stem cell-like cells (TICs) also express the overlapping progenitor genes and uniquely possess self-renewing and tumor-initiating properties due to intrinsic TLR4-Nanog oncogenic pathway4. Therefore, LPCs and TICs may share the same or similar lineage, but a definite relationship between them is currently unknown.
PROM1 is a well-known marker of TICs in many cancers and used to define cancer stemness and disease severity in patients5,6. PROM1-positive (PROM1+) cells also accumulate in mouse liver injury models and livers of alcoholic hepatitis (AH) patients in the pattern commonly characterized as ductular reaction progenitors (DRPs). The expression of PROM1 is also inversely associated with survival of AH patients7. These clinical findings underscore the potential importance of PROM1+ cells in chronic liver disease and cancer. A study using mice bearing Prom1C-L which expresses Cre recombinase and LacZ genes in the Prom1 locus, reports that PROM1+ cells are generated by proliferating ductular epithelial cells in response to chronic thioacetamide liver damage8. Another study using the same genetic mice, reports that PROM1+ cells from neonatal liver have high regenerative capacity, which is almost undetectable in the adult liver9. However, these PROM1+ cells reappear in adult mice upon feeding a diet containing 3,5-diethoxycarbonyl-1,4-dihydrocollidine (DDC), and such livers express similar stemness transcriptomes as those in neonatal liver. Further, PROM1+ cells are transformed to tumor cells and give rise to more aggressive liver tumors upon DDC feeding when they are forced to express oncogenes or mutant tumor suppressors9. Although HCC cells are not derived from cytokeratin 19 (Krt19) or osteopontin (Spp1) expressing biliary epithelial cells, they express higher levels of Prom1 compared to non-tumor tissues, suggesting the contribution of hepatocyte-derived PROM1+ cells in HCC formation10.
Alcoholic liver disease (ALD) is one of the significant health-related financial burdens in the world11. AH represents the most severe spectrum of ALD with high mortality12 and is presented as acute syndrome superimposed on chronic liver disease characterized by intense polymorphonuclear cell (PMN) inflammation, the appearance of DRPs and jaundice. One study suggests that these DRPs in AH patients reflect cholangiocyte proliferation13. But it is largely unknown how DRPs emerge in AH or if and how they contribute to AH pathogenesis. To address these questions in an animal model, we developed a mouse model which reproduces histologic AH with DRP proliferation14. The model also reproduces clinical features of AH including portal hypertension as judged by splenomegaly, hypoalbuminemia, and increased plasma bilirubin. More importantly, an expansion of PROM1+ cells expressing SPP1, NANOG, and AFP is detected in the model14.
Understanding the mechanisms underlying how PROM1+ cells emerge and expand in AH mouse liver may reveal the potential pathogenic role of these cells in AH and liver cancer susceptibility. Unfortunately, epidemiology of liver cancer incidence in AH patients is lacking due primarily to the high mortality rate of AH patients and difficulties in long-term tracking and assessment of the patients after hospital discharge. However, heavy alcohol intake and Western diet are two top-ranked risk factors for both AH and liver cancer15. In the present study, we aimed to determine the origin and molecular characteristics of PROM1+ cells in AH by genetic lineage tracing and RNA sequencing (RNAseq) analysis and to assess the fate of PROM1+ cells in liver tumor development in mice given Western alcohol diet (WAD) by genetic tracing and single-cell RNA sequencing (scRNAseq).
Results
Transcriptome of PROM1+ cells from AH resembles known DRP and HCC gene signatures
To characterize a gene expression profile of PROM1+ cells in AH, we isolated PROM1+ cells from AH mouse livers14 by FACS using antibodies against PROM1 and CD49f which were used as stem cell markers for mouse liver TICs4,16. In normal mouse liver, a PROM1+CD49f+CD45– population accounts for ~ 0.1% which increases to 1.93% in diethyl nitrosamine (DEN)-liver cancer model4. The yield of PROM1+CD49f+CD45– cells from AH mouse livers ranged from 1 to 4% of sorted cells (Fig. 1a), suggesting PROM1+ cells are increased in the liver of AH mice. To characterize PROM1+ cells, PROM1+CD49f+CD45– (R6) versus PROM1– CD49f+CD45– (R5) cells were sorted and total RNA was sequenced as shown in a flow chart in Fig. 1b. We also sorted PROM1+CD49f+ cells present in CD45+ population as hematopoietic PROM1+ cells (R8).
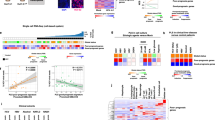
Progenitor and tumorigenic characteristic of PROM1+CD49f+ cells isolated from mouse model of alcoholic hepatitis (AH). (a) FACS plots of PROM1+CD4f+ cells isolated from normal mice, AH mice and DEN-initiated liver tumor mice. For DEN mice, 2-week old C57/BL6 mice were injected with 25 mg/kg of DEN intraperitoneally and fed with WAD for 5 months. (b) The flow chart shows how PROM1+CD49f+CD45– cells were isolated from mouse liver. (c) Hierarchical clustering of PROM1+CD49f+CD45– (R6), PROM1–CD49f+CD45– (R5), PROM1+CD49f+CD45+ (R8) cells and CD45– liver cells, is shown using a differentially expressed gene (DEG) set specific for R6 versus R5. (d) Venn diagrams show the number of upregulated and downregulated genes specific for PROM1+ liver cells (CD45–). (e) Ingenuity pathway analysis of DEG specific for PROM1+ liver cells, reveals known stemness and tumorigenic pathways highlighted in red, are significantly regulated.
Total RNAseq analysis revealed a unique transcriptomic landscape of R6 PROM1+ cells as compared to R5 PROM1– or R8 hematopoietic PROM1+ cells (Fig. 1c). To identify differentially regulated genes in R6 (PROM1+) cells, we compared gene expression profiles of R6 versus R5 (PROM1–), R6 versus R8 (hematopoietic PROM1+), and R6 versus all CD45– sorted cells. As shown in the Venn diagrams in Fig. 1d, this analysis identified 613 upregulated and 708 downregulated genes specific for PROM1+ liver cells (Suppl. Table 3 in Excel format). The upregulated gene list included twofold or higher expression of genes considered as markers for progenitors (Prom1, EpCAM, Sox9), biliary epithelial cells (Krt19, Hnf1β, Spp1) and mature hepatocytes (HNF4α, Alb), suggesting hepatic PROM1+ cells in AH are bipotent progenitor cells. Ingenuity Pathway Analysis of the PROM1+ cell transcriptome, also revealed significantly regulated stemness and tumorigenesis pathways such as pluripotency, EMT, tissue factor in cancer, Ephrin receptor, HIPPO pathway, and ILK signaling, suggesting their potential involvement in liver tumorigenesis (Fig. 1e).
Lineage of PROM1+ cells in AH and their fate in liver cancer
To determine how PROM1+ cells are generated in AH, we performed lineage tracing in the AH mouse model. To test the contribution of hepatocytes to the genesis of PROM1+ cells, we utilized two approaches to label hepatocytes in Rosa26mTmG reporter mice before AH induction: the use of the conventional Albumin-Cre (Alb-Cre) and the more mature hepatocyte specific Transthyretin-Cre (Ttr-Cre). Alb-Cre; Rosa26mTmG mice showed strong GFP expression by hepatocytes. Using these cells, we were able to establish the gating for GFP positive cells.
To generate Ttr-Cre mice, we injected AAV8-Ttr-Cre virus to Rosa26mTmG mice via tail vein and allowed 1 month for recombination to occur. The labeling efficiency was 98.8% for Alb-Cre mice and 99.2% for Ttr-Cre mice according to GFP positivity (Suppl. Figure 1b and 1c). To trace GFP+ hepatocytes to PROM1+ DRPs in the AH mice, we isolated CD45– nonparenchymal liver cells by a low speed centrifugation to remove hepatocytes and magnetic depletion of CD45+ cells. The cells from C57/BL6 mice after the AH protocol were used as a negative control because the Western diet regimen contributed to high background fluorescence (Suppl. Figure 1d). The cells were sorted from Alb-Cre;Rosa26mTmG mice (Suppl. Figure 1g) and Rosa26mTmG mice injected with AAV8-Ttr-Cre (Suppl. Figure 1j) for PROM1 and GFP. From the cells from AH liver of Alb-Cre;Rosa26mTmG mice, PROM1+ /GFP+ cells were detected at 0.118% of frequency versus PROM1+ /GFP– cells at 0.172%, suggesting 40.7% of PROM1+ cells are GFP-traced cells from Alb-Cre-based labeling in this representative mouse (Suppl. Figure 1g). Using Ttr-Cre labeling method which is considered more specific for hepatocytes17, the frequency of GFP+ /PROM1+ cells was shown to be 0.195% as compared to 0.691% of total PROM1+ cells with or without GFP expression, indicating 28.2% of the PROM1+ cells are hepatocyte-derived in this particular Ttr-Cre;Rosa26mTmG mouse subjected to AH (Suppl. Figure 1j, right panel). After repeating the same analysis in 4 AH mice, the average percentage of GFP+ /PROM1+ cells was determined to be 24% (Suppl. Figure 1k). These results suggest that PROM1+ cells are generated in part from mature hepatocytes in AH and the origin appears heterogeneous. The latter notion is also consistent with the recent finding of the heterogeneity of PROM1+ cells in HCC18.
A subpopulation of PROM1+ cells observed in HCC is suggested to possess tumor initiating properties4. Indeed, the DRP markers such as Prom1, EpCAM, Krt19, and Sox9 detected in AH, were also induced in non-tumor liver tissue (NTL) of mice subjected to DEN injection and tumor promotion with WAD as compared to normal liver or tumor tissues, suggesting a more selective increase of DRPs in NTL (Fig. 2a). Indeed, immunofluorescent microscopy detected proliferating DRPs were concentrated in NTL around the peripheral border of the tumor mass as Ki67+ SOX9+ cells (purple arrows, Fig. 2b), revealing a close spatial association of DRPs with tumor cells.
Lineage tracing of PROM1+ cells in a mouse model of DEN-initiated, WAD-promoted liver cancer. (a) Expression of DRP markers in non-tumor liver (NTL) versus tumor tissues. *p < 0.05 and **p < 0.01 compared to normal livers; #p < 0.05 compared to NTL by non-parametric student’s t-test (n = 6). (b) Representative co-immunofluorescent staining for hepatocyte marker (HNF4α), proliferation marker (Ki67) and a DRP marker (SOX9) of a tumor-bearing liver section from a DEN-WAD mouse with insets of enlarged views (b1 and b2). Yellow and pink arrows indicate Ki67+HNF4α+ hepatocytes and Ki67+SOX9+ DRPs, respectively. Scale bar = 100 μm. (c) Schematic diagram of lineage tracing of PROM1+ cells using Prom1C-L;Rosa26mTmG mice in DEN-WAD model. Two-week old Prom1C-L/Rosa26mTmG mice with Prom1C-L homozygosity (Prom1KO) and its wildtype counterparts were injected with 25 mg/kg DEN intraperitoneally. The mice were then injected with two doses of Tam 3 days apart before WAD was introduced. The mice were euthanized after 5 months of WAD feeding. (d) Total tumor volumes from Prom1KO and its wildtype were shown. Width (W) and length (L) of each visible tumor was measured with a caliper. Tumor volume was calculated with the following formula: (LxW2)/2. Statistical comparison was done with student’s t-test. (e) Representative tumor images of hematoxylin and eosin sections of WT and Prom1KO DEN-WAD mouse livers. The bottom images are magnifications of the top images with tumor (T) and nontumor (NT) areas. Scale bar = 200 μm. (f) Immunofluorescent (IF) staining of Prom1C-L;Rosa26mTmG liver sections showing KRT19+ (pink) DRP cells with some with co-localized GFP staining (yellow). Scale bar = 50 μm. (g) IF section from the same mouse shown in f, depicting GFP (green)-labeled tumor cells (T) and KRT19+ (pink) DRP cells in NTL area. Scale bar = 50 μm.
To determine the fate and role of PROM1+ cells in liver tumorigenesis, we traced PROM1+ cells using the DEN-initiated and WAD-promoted liver tumor model in Prom1C-L mice, a strain engineered to have CreERT2 and Lacz genes in the Prom1 locus19. While this mouse serves to express CreERT2 under the Prom1 promoter for labeling of PROM1+ cells, the homozygous strain of Prom1C-L/C-L can also be used as Prom1 knockout because it completely ablates the Prom1 expression due to the biallelic knock-in of the CreERT2-Lacz cassette. We crossed Prom1C-L/C-L mice with Rosa26mTmG mice to generate Prom1 C-L; Rosa26mTmG (Prom1WT) and Prom1C-L/C-L;Rosa26mTmG (Prom1KO) mice, injected DEN at 2-week age and fed WAD from 5-week age for 6 months. Tamoxifen (Tam) was injected to label PROM1+ cells at 3 weeks after DEN injection (Fig. 2c). Liver tumor development in Prom1WT and Prom1KO mice was assessed via gross and histologic examination (Fig. 2d,e). As predicted, Prom1 expression in tumors of Prom1C-L/C-L;Rosa26mTmG (Prom1KO) mice were almost undetectable by qPCR analysis, validating their knockout status (Suppl. Figure 2a).
Surprisingly, total liver tumor volumes calculated in these two genetic groups, were not different (Fig. 2d), indicating PROM1 is not essential in liver tumorigenesis. To determine the fate of PROM1+ cells in tumor development, we stained the liver sections with visible tumors with anti-GFP and anti-KRT19 antibodies (Fig. 2f,g). While DRPs were detected as GFP+ KRT19+ cells in NTL region, tumor cells were also labeled with GFP but not for KRT19. These results suggest that PROM1+ cells give rise to liver tumor cells even though PROM1 is not functionally required for tumor development.
scRNAseq analysis reveals Prom1+ TIC and DRP subpopulations in liver tumor
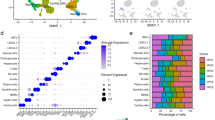
To understand the molecular characteristics of Prom1+ cells in the single cell resolution, we performed scRNAseq analysis. To enrich the LPCs and TICs, we have developed a unique approach of sorting Col1a1-expressing cells by FACS. The rationale for this approach is based on: (1) LPCs were previously reported to express high levels of Col1a1 expression3; (2) our own results confirmed 23-fold and 90-fold higher expression of Col1a1 in LPC line (PIL4) and TICs as compared to mouse primary hepatocytes, respectively (Suppl. Figure 3a); and (3) our additional data validated higher expression of Col1a1 in PROM1+ versus PROM1– cells isolated via MACS from the DEN-WAD mouse livers (Suppl. Figure 3b). We used Col1a1-driven Cre recombinase expression to label LPCs and TICs in Col1a1Cre;Rosa26mTmG mice subjected to the DEN-WAD protocol. Tumor bearing livers were perfused and digested with collagenase to collect liver cells. Mature hepatocytes were removed by low speed centrifugation, and the remaining liver cells were subjected to FACS sorting based on ultraviolet (UV) and GFP. UV-excited gating was used to exclude vitamin A-storing hepatic stellate cells (HSC), the major Col1a1-expressing cell type in the liver. The UV–GFP+ population was sorted as tumor-associated Col1a1-expressing cells and submitted for scRNAseq analysis (Fig. 3a). Barcode rank plot of the scRNAseq sample showed an excellent separation between the cell-associated barcodes and barcodes associated with empty partitions (Suppl. Figure 4). From a single cell library with the target cell number of 10,000, 8578 cells passed this quality control assessment. By this scRNAseq analysis, we indeed identified two subpopulations which included Prom1+ cells (Cluster 1 and 2) (Fig. 3b) with no or minimal expression of the HSC marker Lrat, portal fibroblast marker Thy-1, hematopoietic marker Cd45, and endothelial cell markers Cd31 and Cdh5 (Suppl. Figure 5a). As predicted, Cluster 1 and 2 cells expressed the biliary or/and progenitor markers such as Sox9, Spp1, Hnf1b, Krt19, Krt23, Epcam (Fig. 3c). To further validate these results, we assessed co-expression of these genes with Prom1 at the single cell level. As shown in Suppl Fig. 5b, Cluster 1 and 2 are enriched with cells which co-expressed Prom1 and Sox9, Spp1, Hnf1b, or Krt23 and most Prom1+ cells in both clusters did not co-express the HSC, portal fibroblast, hematopoietic, and endothelial cell markers, validating these Prom1+ Cluster 1 and 2 cells are LPCs. The cells co-expressing Prom1 and Afp, the known marker of liver tumor cells, were detected primarily in Cluster 1 (Suppl Fig. 5a and Fig. 3a). In fact, Cluster 1 contained both Prom1+ Afp+ and Prom1–Afp+ cells, suggesting TICs and tumor cells were enriched in this Cluster. Prom1+ Afp– cells enriched in Cluster 2 are DRPs as the biliary/DRP markers such as Hnf1b, Krt19, Epcam, Mmp7 and Cftr were more selectively expressed in Cluster 2 cells (Fig. 3c).
Identification of Prom1+ TICs and Prom1+ DRPs subpopulations from DEN mouse liver by scRNAseq. (a) The FACS plot depicting the sorting of UV– GFP+ cells from DEN-WAD mouse liver cells. Non-parenchymal cells were isolated by collagenase perfusion from Col1a1Cre;Rosa26mTmG mice subjected the DEN-WAD regimen. The cells were sorted by FACS to remove UV+ hepatic stellate cells and to collect UV–GFP+ cells as Col1a1 expressing and vitamin A negative cells. The scRNAseq library was prepared immediately after sorting. (b) The analysis detected Prom1+Afp+, Prom1+Afp– and Prom1–Afp+ populations. The plots were created in the t-SNE format using filters of Prom1 and Afp transcripts. (c) Co-expression analysis of Prom1+ cells with known markers of DRPs, TICs, tumor. The plots were created in the t-SNE format with gene/feature expression in Loupe browser. The scale bar represents log2-transformed normalized expression.
Novel transcript profiles specific for Prom1+Afp+, Prom1+Afp– and Prom1–Afp+ populations. (a) The graph-based clustering of DEG sets for Prom1+Afp+, Prom1+Afp– and Prom1–Afp+ cells. (b–d) volcano plots of transcript profiles from the three subpopulations. (e–g) Top representative novel transcripts identified by scRNAseq for each subpopulation are shown in t-SNE format as generated by the Loupe browser. The scale bar represents log2 transformed normalized expression.
Unique upregulation of Discoidin Domain Deceptor 1 (Ddr1) in PROM1+ cells. (a) Integrated analysis of a RNAseq profile of PROM1+ cells from mouse AH, a RNAseq profile of the liver of mouse AH, and a proteomic profile of human AH versus control liver, reveals 7 upregulated and 7 downregulated genes common in all three comparisons. (b) Quantitative PCR analysis of DEN-WAD mouse livers versus normal livers, reveals that some of the upregulated genes shown by the integrated bioinformatic analysis are also induced in the liver of the mouse liver tumor model including Ddr1. *p < 0.05 based on non-parametric student’s t-test. (c) Expression of Ddr1, Ddr2, Spint1, Ehf, Lamc3 and Pkhd1 at the single cell level are shown in the t-SNE format generated by Loupe browser. The scale bar represents log2 transformed normalized expression.
We next assessed whether the known TIC/HCC transcripts are expressed in the scRNAseq profiles of the two clusters. For this, we assessed a set of genes reported to characterize cancer progenitor cells with tumor-initiating properties which were isolated from DEN-induced tumor bearing livers20. Many of the genes upregulated in these tumor initiating cells were expressed in our Cluster 1 cells including Tff3, Tspan8, Gpc3, Ly6d, Cldn2 (Fig. 3c). These genes are involved in malignancies and cancer stemness21,22,23, further supporting the notion that Cluster 1 cells are TICs and tumor cells. We then acquired gene expression profiles unique to each of the three populations of Prom1+ Afp+ , Prom1+ Afp–, and Prom1–Afp+ cells by comparing among them by the locally distinguishing comparison feature of the Loupe Browser as depicted by heatmaps and volcano plots (Fig. 4a–d). As shown in the heatmaps, Prom1+ Afp+ and Prom1–Afp+ cells share coordinately upregulated genes most likely reflecting their common tumorigenic function yet with some discordant gene expression which may reflect genetic differences between TICs with stemness property and tumor cells which have lost the stemness. In contrast, Prom1+ Afp– cells which we consider as DRPs have a clearly distinct gene expression pattern from other two Afp+ cell populations. Expression of 4 top genes uniquely expressed in each population are shown in Fig. 4e–g, and complete lists of the genes are provided in Suppl. Tables 4–6 in Excel format.
Discoidin domain receptor 1 is a potential oncogenic driver of PROM1+ cells
To address the relevance of the transcriptomic data from PROM1+ cells to AH livers in mice and patients, we compared our PROM1+ cell RNAseq profile with a RNAseq profile of AH mouse liver and a previously reported proteomic profile of human AH liver24. This integrated analysis identified 7 upregulated and 7 downregulated genes that are common in PROM1+ cells, mouse and human AH livers (Fig. 5a). Since our results showed PROM1+ cells gave rise to liver tumor cells, we looked for potential oncogenic drivers unique to PROM1+ cells by first screening the expression of the 7 upregulated genes in NTL tissues of DEN/WAD mice compared to normal liver. This analysis revealed that Prom1, Ddr1, Ddr2, Lamc2, Lamc3 and Pkhd1 were induced significantly in DEN-WAD livers (Fig. 5b). Next, we examined the expression of these genes in cell clusters revealed by our scRNAseq analysis. Of these genes examined, Ddr1 and Pkhd1 expressions were detected in Cluster 1 and 2 (Fig. 5c). DDR1 is known as a receptor for collagen25 and implicated in oncogenesis, metastasis and chemoresistance of cancers of pancreas, lung, prostate, ovary, and colon26,27,28,29. Ddr2, in contrast, is expressed in clusters of activate Lrat+ HSC and Thy1+ portal fibroblasts (Fig. 5c and Suppl. Figure 5). Spint1 and Ehf were expressed more selectively in DRPs in Cluster 2 (Fig. 5c), suggesting their importance in the biology of Prom1+ DRPs.
To confirm DDR1 upregulation in AH in patients in our hands, we analyzed DDR1 mRNA levels in liver tissues of AH patients versus healthy subjects received from the explant repository program of John Hopkins University. Indeed, this analysis revealed at least 20-fold higher DDR1 expression in AH livers compared to the normal liver (Fig. 6a). DDR2 level was also statistically higher in AH tissues than normal liver, but the fold induction was much lower than DDR1. To address the role of DDR1 in human liver cancer, we performed the data mining approach. This bioinformatic analysis showed that DDR1 expression was significantly elevated in both HCC30 (Fig. 6b) and cirrhotic livers31 (Fig. 6c) compared to normal livers.
DDR1 upregulation in liver cancer associated with poor patient survival. (a) DDR1 and DDR2 expression levels were measured by qPCR in normal human livers versus alcoholic hepatitis (AH) livers. *p < 0.05 based on non-parametric student’s t-test. (b,c) DDR1 expression in HCC30 and cirrhotic31 human livers were compared to normal livers using the existing microarray data from Oncomine database49. (d) DDR1 expression was compared in intrahepatic cholangiocarcinoma (ICC), combined HCC and ICC (HCC-ICC) and HCC in patients30. *p < 0.05 and **p < 0.01 based on the Oncomine database. (e–g) Kaplan–Meier analysis of TCGA HCC cohort data using UCSC Xena32, reveals an inverse relationship between the expression of DDR1, PROM1 or both with patient survival. p values were calculated by log-rank test.
Patients with combined HCC-intrahepatic cholangiocarcinoma (ICC) have poor prognosis33. ICC is characterized by higher expression of liver progenitor markers such as EpCAM and KRT19, suggestive of expanded TICs and/or DRPs. Indeed, DDR1 expression is higher in patients with ICC or combined HCC-ICC compared to those with HCC alone (Fig. 6d)34.
Next we assessed the relationship of DDR1 and PROM1 expression with the HCC patient survival, by analyzing in TCGA HCC cohort data using UCSC Xena database32. We stratified the cohort into DDR1 high and low expression groups. The Kaplan–Meier analysis showed that DDR1high patient group had lower survival than the DDR1low patients (p < 0.039) (Fig. 6e). When PROM1 was used to stratify the cohort, PROM1high patients had a significantly worse outcome in their survival compared to the PROM1low group (p < 0.021) (Fig. 6f). When PROM1 and DDR1 co-expression was analyzed, 23 patients with high expressions of both PROM1 and DDR1 had a worst outcome compared to 69 patients with both genes expressed at the low level (p < 0.014) (Fig. 6g). The difference in survival in these patients was greatest (> 1825 days). In summary, DDR1 is induced in human cirrhosis, HCC, and ICC and inversely associated with the survival of HCC patients. In addition, the expression of DDR1 is correlated with PROM1 expression and predicts the survival of HCC patients in this TCGA cohort. These data suggest that DDR1 expressing PROM1+ DRPs and TICs may determine the clinical course of HCC patients.
Functional significance of DDR1 in TIC-derived tumor development
To determine the functional significance of DDR1 expressed in PROM1+ cells in liver oncogenesis, we first measured Ddr1 expression in TICs, PIL4 liver progenitors and mouse primary hepatocytes. TICs were isolated from a mouse model of alcohol-promoted liver cancer4. PIL4 cells were previously isolated from p53-null mouse fed with choline-deficient, ethionine-supplemented diet35. TICs have self-renewing and tumor-initiating activities while PIL4 cells lack these properties but mimic DRPs. As expected, albumin is expressed exclusively by primary hepatocytes but not by TICs and PIL4 cells while Krt19 (CK19) is expressed by PIL4 cells. Tlr4 which is induced in stem cell-like cells4, is expressed moderately by PIL4 cells and most conspicuously by TICs along with Nanog, Sox2 as previously reported (Suppl. Figure 6). A pattern of Ddr1 expression followed that of Tlr4: undetectable in primary hepatocytes, moderately increased in PIL4 cells, and highest expression (~ 40-fold) in TICs. When Ddr1 but not Ddr2 was selectively knocked down 80% in TICs by using two separate shRNAs (Ddr1-sh1 and Ddr1-sh2) versus scrambled shRNA (Scr-sh) (Fig. 7c), TIC growth was significantly impaired as shown by crystal violet staining and cell count (Fig. 7a,b). Further, the treatment with the DDR1 inhibitor (DDR1in7rh), suppressed the growth of TICs at IC50 of 0.95 µM and that of the liver cancer cell line DIHXY at IC50 of 1.55 µM while PIL4 cells were four–fivefold less sensitive (Fig. 7d). These results suggested that DDR1 has a functional significance in TIC and liver cancer cell growth. To test this potential oncogenic role of DDR1 in vivo, we performed a xenograft experiment in nude mice transplanted with TICs transduced with Ddr1-sh1 versus Scr-sh. Ddr1 knockdown in TICs significantly attenuated TIC-derived tumor growth to one third of that with TICs with Scr-sh, supporting the functional role of DDR1 in tumor development (Fig. 7e).
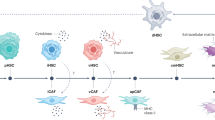
Ddr1 is required for TIC growth and TIC-derived tumor development in mice. (a,b) TICs transduced with lentivirus expressing scrambled shRNA (Scr-sh), or two Ddr1shRNAs (Ddr1-sh1 and Ddr1-sh2) were grown in 6-well plates for 4 days. The cell number was quantitated with crystal violet staining (a) and trypan blue cell counting (b). Average and SEM were calculated from three experiments. **p < 0.01 and ***p < 0.005 based on One-Way ANOVA with Bonferroni post-test. (c) Ddr1 knockdown in TICs was confirmed by qPCR. (d) TICs (circles), DIHXY (triangles) and PIL4 (diamonds) cells were treated with a DDR1 specific inhibitor at a range of concentrations and cell viability was assessed by MTT assay. Each data point represents an average of six replicates. IC50 values were calculated using Graphpad Prism. (e) TICs transduced with Scr-sh or Ddr1-sh1 were subcutaneously transplanted in nude mice for assessment of tumor development. Each data point is average and SEM of tumor volume from n = 6. *p < 0.05 versus Scr-sh TICs. (f) The proposed model of the role PROM1+ progenitors in liver cancer development. Hepatocyte damage caused by hepatotoxins, alcohol and high fat diet, stimulates proliferation of ductular reaction progenitors (DRPs) including PROM1+ liver progenitor cells (LPCs). These PROM1+ cells have potential to maintain the DRP lineage and to undergo a transition to tumor cells, possibly via PROM1+ TICs. PROM1+ DRPs and TICs have high expression of Ddr1 which drives oncogenesis in the latter cells. Inhibition of DDR1 may suppress the contribution of PROM1+ cells to liver cancer development.
Discussion
Our study provided several new findings on the origin, fate and functional role of PROM1+ cells in alcohol associated liver injury and tumorigenesis. Firstly, using lineage tracing, we showed hepatocytes are in part the origin of PROM1+ cells in AH. Secondly, our RNAseq analysis revealed these PROM1+ cells have a unique gene expression profile consistent with their characteristics as LPCs/DRPs or TICs/tumor cells. Thirdly, in the DEN-initiated, WAD-promoted liver tumor model, Prom1+ cells gave rise to DRPs and more importantly tumor cells. Yet, biallelic Prom1 knock-in mice which lack Prom1 expression, still developed liver tumors as WT mice, suggesting that PROM1 is a pivotal marker for DRPs/TICs which generate tumor cells while PROM1 itself is not required for tumor development. Fourthly, we have developed a unique method of enriching LPCs and TICs using Col1a1-based labeling and FACS. This allowed us to identify two Prom1+ subpopulations by scRNAseq analysis which we believe represent DRPs and TICs based on their similar yet distinct gene expression profiles. Fifthly, our integrated bioinformatic analysis of RNAseq and scRNAseq data disclosed Ddr1 as a gene uniquely upregulated in PROM1+ cells, and our in vitro and in vivo loss-of-function approaches revealed the oncogenic role of Ddr1 in TICs. Finally, the clinical relevance and importance of DDR1 was supported by DDR1 induction in patient HCC and its significant association with low patient survival.
The fate of LPCs has actively been investigated in different mouse models and generated conflicting or mixed results. Lineage tracing studies showed that cholangiocytes defined by KRT19+ or SOX9+ cells rarely gave rise to hepatocytes36,37. In line with this observation, KRT19+ or SOX9+ cells did not develop HCC in mice, further suggesting that cholangiocyte and hepatocyte lineages may not merge10,38. However, recent studies using dual lineage tracing of cholangiocytes and hepatocytes revealed that a significant portion of SOX9+ LPCs differentiated into both cholangiocytes and hepatocytes39,40. Moreover, KRT19+ cholangiocytes became hepatocytes when hepatocyte proliferation was impaired by p21 overexpression or feeding mice with choline deficient ethionine supplemented diet, methionine and choline deficient diet or DDC containing diet41. In contrast to these studies which utilized Sox9, Krt19 and Spp1 as a driver of LPC specific labeling, we used Prom1-Cre to determine the fate of LPCs/DRPs and TICs in liver tumor development based on their shared expression of Prom1. Our results revealed that PROM1+ cells which were derived in part from hepatocytes in AH, gave rise to both DRPs and liver tumor cells in the DEN-WAD mouse model. Because of TICs’ known tumor-initiating activity4, labeled TICs were presumed to have generated tumor cells while labeled DRPs must have persisted or expanded in the model. However, we did not observe labeled hepatocytes, suggesting no generation of hepatocytes from PROM1+ cells in this model. Our method could not allow selective fate mapping of DRPs versus TICs to clarify their relative contribution to generation of tumor cells. Thus, we cannot rule out a possibility of PROM1+ DRPs giving rise to tumor cells either directly or through TICs. From a list of genes shown to be uniquely expressed in Prom1+ Afp+ cells versus Prom1+ Afp– or Prom1–Afp+ cells, we may be able to devise a method of selective TIC or DRP labeling in future.
Intratumoral heterogeneity presents a unique challenge for understanding of HCC development and therapy resistance. Like other cancers, HCC comprises diverse tumor clones that develop via clonal evolution. HCC in patients with expression of tumor stem cell markers such as EpCAM, PROM1 and KRT19 tend to be aggressive and chemoresistant. Recent single cell genomics has identified an EpCAM + subpopulation with the stemness property with CTSE as an oncogene uniquely induced in this subpopulation42. In our study, DDR1 was identified as a tumor driver gene in PROM1+ subpopulation relevant to both mouse and patient liver cancer. In particular, patients with combined HCC-ICC having co-overexpression of DDR1 and PROM1 had the worst survival, suggesting the contribution of PROM1+ TICs to this clinical phenotype. We also identified a group of genes uniquely upregulated in Prom1+ Afp+ cells which obviously deserve further attention. Of interest, three of the 4 top genes expressed in this subpopulation (Fig. 4e), are metabolic genes: Gulo, beta-catenin responsive gene which encodes the limiting enzyme in the final step in ascorbic acid biosynthesis43, Cyp2c70, a key enzyme in generation of the murine specific bile acid, a-muricholic acid, or Elovl2, an enzyme involved in elongation of long chain w-3 and w-6 fatty acids which was shown to confer stemness of glioma cells44. Prom1+ Afp+ cells may constitute therapy-resistant TICs in liver cancer45 and are a logical and attractive therapeutic target. In fact, in cancer of other organs such as lung, breast, colon, and prostate, DDR1 is already considered as a therapeutic target28,29,46. A strategy to selectively direct interventions to key driver genes such as DDR1 in PROM1+ TICs may be desired for the therapy for liver cancer. Finding a molecular link between liver injury and liver carcinogenesis has been a topic of interest. Recently, the DAMP high mobility group box 1 (HMGB1) is shown to underlie hepatocyte-to-ductal metaplasia in liver injury and HCC development in mice47. However, the molecular mechanism involved in the potential transition of LPCs to TICs or tumor cells are elusive. Our findings suggest that DDR1 may at least in part facilitate a potential tumorigenic link of PROM1+ cells to liver cancer (Fig. 7f). In fact, our study is first to show that DDR1 is essential for TIC growth and TIC-derived tumor formation. Future studies are obviously needed to determine how DDR1 participates in the transition of PROM1+ cells to liver tumor cells in vivo. In summary, our study provides the evidence for hepatocyte-origin of PROM1+ cells in AH and the functional role of such cells in liver tumorigenesis (Fig. 7f).
Methods
Animals and animal models
All animal experiments were performed by the Animal Core of the Southern California Research Center for ALPD and Cirrhosis in accordance to an approved Institutional Animal Care and Use Committee (IACUC) protocol of University of Southern California (USC). C56/BL6, Prom1C-L (Prom1tm1(cre/ERT2)Gilb) and Rosa26mTmG (Gt(ROSA)26-Sortm4(ACTB-tdTomato,-EGFP)Luo) were purchased from the Jackson Laboratory and crossed to generate Prom1C-L; Rosa26mTmG(fl/fl) mice for the lineage tracing. AAV8/DJ-Ttr-Cre expressing Cre recombinase gene under hepatocyte specific Ttr promoter was purchased from Vector Biolabs and 4 × 1011 GC was injected into each mouse via tail vein. Col1a1Cre;Rosa26mTmG mice were also generated for FACS-based isolation of Col1a1 + liver cell population for scRNAseq analysis. For a model of alcoholic hepatitis, the mice were fed 40% of their daily caloric intake as ad libitum intake of Western solid diet containing 1% (w/w) cholesterol and saturated fat and the remaining 60% by intragastric feeding of alcohol-containing liquid diet. They were also superimposed with weekly binge as previously described14. For DEN-initiated, WAD-promoted liver tumorigenesis, two-week old male mice were injected with 25 mg/kg of DEN intraperitoneally (Sigma Aldrich, St Louis). For lineage tracing in Prom1C-L; Rosa26mTmG(fl/fl) mice, the mice were intraperitoneally injected with two doses of 75 mg/kg tamoxifen (Sigma Aldrich, St Louis) at the 3-day interval at 3 weeks after DEN injection. Western alcohol liquid diet (WAD) (#710362; Dyets Inc.) was introduced to the mice at 5 weeks old by incrementally increasing alcohol concentration in the diet, reaching the maximal concentration of 3.5% (v/v). After 5–6 months of WAD feeding, the mice were euthanized, the livers were carefully excised, weighed and visible tumors were counted. Each tumor length (L) and width (W) were measured by a caliper and the tumor volume was calculated using the formula of (LxW2)/2. For TIC xenograft transplantation model, NCr nude mice (Taconic Biosciences) were subcutaneously transplanted with 1 × 105 TICs transduced with shRNA against Ddr1 or scrambled shRNA, and TIC-derived tumor growth was monitored for 2 weeks by measuring the tumor length and width for assessment of tumor volume as described above.
Cell culture
TICs were provided by the Integrative Liver Cell Core (ILCC) of USC and cultured in 1:1 mixture of Dulbecco's modified Eagle medium (DMEM) and Nutrient Mixture F‐12 media (Life Technologies, Camarillo, CA) with 10% fetal bovine serum (FBS), 20 ng/mL murine endothelial growth factor (EGF; Peprotech, Rocky Hill, NJ), Embryomax nucleosides (MilliporeSigma, Burlington, MA), 100 nM dexamethasone (Sigma–Aldrich, St. Louis, MO), and antibiotic‐antimycotic (Life Technologies) as described before4,45. DIHXY cells were cultured in DMEM medium with 10% fetal bovine serum (FBS) and antibiotic‐antimycotic (Life Technologies). PIL-4 cells were obtained from Dr. George Yeoh of University of Western Australia35 and cultured in Williams’ E medium supplemented with 10% FBS, EGF (Peprotech), IGF-II (Peprotech) and human Insulin (Sigma–Aldrich, St. Louis). The DDR1 inhibitor 7rh was purchased from Sigma–Aldrich for the treatment of TICs in culture. MTT assay and crystal violet assay were performed as described before45.
Alcoholic hepatitis patient and healthy liver tissues
AH patient and control human livers were obtained from the Clinical Resources for Alcoholic Hepatitis Investigators (1R24AA025017) of Johns Hopkins University as previously described24. The patient samples were obtained with informed consent in accordance with Johns Hopkins Hospital Institutional Review Board as described before41. These tissues were excised from explanted livers in patients with severe AH during liver transplantation, or wedge biopsies from the donor livers (normal control). Suppl. Table 1 summarizes basic clinical information of 4 AH patients and 4 healthy donors from which the samples we analyzed were collected.
Immunohistochemistry and immunofluorescence staining
The liver slices were fixed in 4% paraformaldehyde (PFA) at 4◦C overnight and transferred to 70% ethanol on the next day. The fixed tissues were paraffin-embedded, and paraffin sections were stained with hematoxylin & eosin (H&E). For immunofluorescence staining (IF), the liver slices were fixed in 4% PFA at 4◦C for 2 h, incubated in10% sucrose in PBS for 2 h and 30% sucrose in PBS for overnight. The tissues were then frozen in the freezing medium and sectioned with a Cryostat (CM1900; Leica, Buffalo Grove, IL) to prepare 7 µm thickness cryosections. The sections were air-dried overnight, treated with 3% H2O2 in methanol and permeabilized with proteinase K prior to blocking with the solution containing 5% donkey serum and 0.2% bovine serum albumin for 30 min. The sections were then incubated with primary antibodies against Green Fluorescent Protein (GFP; 400-fold dilution, Rockland, Limerick, PA) or KRT19 (TROMAIII; 50-fold dilution, Developmental Studies Hybridoma Bank, Iowa City, IA), subsequently incubated with secondary antibodies: Donkey anti-Goat conjugated with AlexaFluor488 and AlexaFluor 550 (400-fold dilutions, Thermo Fisher Scientific) for 30 min. The sections were counterstained with DAPI. Images were captured with a Nikon 90i microscope and DS-Fi1 digital camera (Nikon, Melville, NY). For IF shown in Fig. 3b, the liver section was stained for HNF4α (50-fold dilution, #6556, Santa Cruz Biotechnology), KI67 (50-fold dilution, #652,402, Biolegend), SOX9 (500-fold dilution, #AB5535, Millipore) as previously described before48. Images were captured with a confocal microscope.
FACS isolation
Isolation of PROM1+ cells was performed by the ILCC of USC. The liver was perfused with EMEM medium, minced on a 10 cm dish and resuspended in collagenase IV containing digestion medium (25 mg of collagenase type IV (C5138-Sigma Aldrich) in 50 mL of serum free DMEM-F12, pH 7.35–7.4). Hepatocytes were removed by low speed centrifugation. CD45+ cells were depleted with CD45 microbeads (Miltenyi Biotec). The cells were blocked with FcR blocking reagent and stained with the corresponding antibodies. CD133-APC, CD49f.-PE and CD45-efluor450 (eBioscience). The single controls and Fluorescent Minus One (FMO) were added in each experiment to gate the positive population. PI was added to separate live cells from dead cells. The cells were sorted in MoFlo Cell Sorter (Beckman Coulter). Isolation of Col1a1 expressing cells was carried out by using the protocol previously described49. A non-parenchymal liver cell fraction from Col1a1-Cre;Rosa26mTmG mice was collected by pronase-collagenase perfusion and gradient ultracentrifugation. Gating for UV and GFP positivity were set by primary hepatic stellate cells isolated from wild type mouse and Col1a1Cre;Rosa26mTmG mouse, respectively, with wild type liver myofibroblasts as a UV and GFP negative cells. UV–GFP+ fraction was collected for scRNAseq analysis.
RNA sequencing
The cells were centrifuged and total RNA was isolated with PicoPure RNA isolation kit (Thermo Fisher Scientific). Prior to library construction RNA integrity was verified by Experion analysis (Bio-Rad). PolyA selection was carried out using Illumina Truseq V2 polyA beads. Sequencing was performed on a NextSeq 500 using V2 chemistry. RNA-seq data was analyzed using the RNA-seq workflow in Partek Flow software (v3. Partek Inc., St. Louis, MO, USA). Briefly, the raw sequencing reads were first trimmed based on the quality score (Phred QC > = 20, min read length = 25 nt) before mapped to mouse genome build mm10 using Tophat version 2.0.8 with default parameter settings and using Gencode M3 annotation as guidance. Gencode M3 was then used to quantify the aligned reads to genes using Partek E/M method. The gene level aligned read counts were normalized for all samples using Upper Quartile normalization before subjected to differential expression analysis using Partek Gene Specific Analysis method (genes with fewer than 10 aligned reads in any sample among a data set were excluded). The differentially expressed gene (DEG) lists were generated using the cutoff of FDR < 0.05 and fold changes greater than 2 either direction.
Single-cell RNA sequencing
Single cell was prepared using the 10 × Single Gene Expression v3 with the 10 × Chromium technology following manufacturer protocol as described before50,51 (cat# 1000154 and 1000074, 10 × Genomics). The library was prepared immediately after cell sorting with the target cell number of 10,000. The library was validated on the Agilent TapeStation (Agilent Technologies, Palo Alto, CA, USA), and quantified by using Qubit 2.0 Fluorometer (Invitrogen, Carlsbad, CA) as well as by quantitative PCR (KAPA Biosystems, Wilmington, MA, USA). The sequencing libraries were clustered on 8 single lanes of a flowcell. After clustering, the flowcell was loaded on the Illumina HiSeq instrument according to manufacturer’s instructions. The samples were sequenced using a 2 × 150 Paired End (PE) configuration. Raw sequence data (.bcl files) generated from Illumina HiSeq was converted into fastq files and de-multiplexed using the 10 × Genomics’ cellranger version 3.1.0 by the Cell Ranger mkfastq command with default parameters. Subsequently, UMI and cell barcode de-convolution and mapping to the mm10 reference genome was performed with 10 × Genomics’ Cell Ranger software package (version 3.1.0) to generate the final digital gene expression matrices and cloupe files using the Cell Ranger count command with default parameters. The quality of data is provided as a barcode rank graph (Suppl. Figure 4). Of targeted 10,000 cells, 8578 cells were subjected for sequencing. After normalization, 38,901 mean reads per cell and 2776 median genes per cell were detected. Data analysis was performed with 10 × Genomics’ Loupe Browser software. Raw data files were uploaded to GEO database with accession number GSE152748.
Quantitative PCR
RNA extraction was performed using Quick RNA miniprep kit (Zymo Research). Tissues were homogenized in RNA lysis buffer. RNA was extracted according to the manufacturer’s protocol. 500–1000 ng of RNA was used as input to synthesize cDNA using high-capacity cDNA reverse transcription kit (ThermoFisher Scientific). Q-PCR analysis was performed using the primers shown in Suppl. Table 1 and Sybr Green reagents (ThermoFisher Scientific) in Viia7 instrument (Applied Biosystem).
Lentiviral mediated Ddr1 knockdown
Ddr1 mission shRNA were purchased from Sigma Aldrich. Two shRNAs with the following sequences were chosen: Ddr1shRNA1 (CCGGGATTCCACTTACGATGGA-TATCTCGAGATATCCATCGTAAGTGGAATCTTTTT) and Ddr1shRNA2 (CCGGGC-CACGCTGAACTTTGTGCATCTCGAGATGCACAAAGTTCAGCGTGGCTTTTT). The shRNAs were packaged into lentivirus with psPAX2 and pMD2.G in 293FT cell line. The lentiviruses were purified using speedy lentivirus purification kit (LV999, Applied Biological Materials Inc., Canada) and titrated using qPCR lentivirus titration kit (LV900, Applied Biological Materials Inc., Canada). The cells were infected with 10 MOI of virus and cultured in 10 µM of puromycin (Invitrogen) after 48 h of virus infection.
Bioinformatic analysis
Bioinformatic analysis was performed by using University of California Santa Cruz (UCSC) Xena program. TCGA liver cancer cohort was selected with the filters: primary tumors and hepatocellular carcinoma as primary disease. DDR1 and PROM1 expression were extracted from the gene expression generated by IlluminaHiseq.
Data analysis
RNAseq data was analyzed by Partek Flow (St. Louis, MO) and performed by USC libraries Bioinformatics Services. Pathway analysis was performed by Ingenuity Pathway Analysis (Qiagen, Redwood City, CA). Statistical Analysis was performed by Graphpad Prism (San Diego, CA) and Microsoft Excel.
References
Miyajima, A., Tanaka, M. & Itoh, T. Stem/progenitor cells in liver development, homeostasis, regeneration, and reprogramming. Cell Stem Cell 14, 561–574. https://doi.org/10.1016/j.stem.2014.04.010 (2014).
Michalopoulos, G. K. & Khan, Z. Liver stem cells: experimental findings and implications for human liver disease. Gastroenterology 149, 876–882. https://doi.org/10.1053/j.gastro.2015.08.004 (2015).
Tarlow, B. D. et al. Bipotential adult liver progenitors are derived from chronically injured mature hepatocytes. Cell Stem Cell 15, 605–618. https://doi.org/10.1016/j.stem.2014.09.008 (2014).
Chen, C. L. et al. Reciprocal regulation by TLR4 and TGF-β in tumor-initiating stem-like cells. J. Clin. Invest. 123, 2832–2849. https://doi.org/10.1172/JCI65859 (2013).
Ma, S. et al. miR-130b Promotes CD133(+) liver tumor-initiating cell growth and self-renewal via tumor protein 53-induced nuclear protein 1. Cell Stem Cell 7, 694–707. https://doi.org/10.1016/j.stem.2010.11.010 (2010).
Zhao, Q. et al. Prognostic value of the expression of cancer stem cell-related markers CD133 and CD44 in hepatocellular carcinoma: from patients to patient-derived tumor xenograft models. Oncotarget 7, 47431–47443. https://doi.org/10.18632/oncotarget.10164 (2016).
Sancho-Bru, P. et al. Liver progenitor cell markers correlate with liver damage and predict short-term mortality in patients with alcoholic hepatitis. Hepatology 55, 1931–1941. https://doi.org/10.1002/hep.25614 (2012).
Kamimoto, K. et al. Heterogeneity and stochastic growth regulation of biliary epithelial cells dictate dynamic epithelial tissue remodeling. Elife. https://doi.org/10.7554/eLife.15034 (2016).
Zhu, L. et al. Multi-organ mapping of cancer risk. Cell 166, 1132–1146.e1137. https://doi.org/10.1016/j.cell.2016.07.045 (2016).
Mu, X. et al. Hepatocellular carcinoma originates from hepatocytes and not from the progenitor/biliary compartment. J. Clin. Invest. 125, 3891–3903. https://doi.org/10.1172/JCI77995 (2015).
Seitz, H. K. et al. Alcoholic liver disease. Nat. Rev. Dis. Primers 4, 16. https://doi.org/10.1038/s41572-018-0014-7 (2018).
Lucey, M. R., Mathurin, P. & Morgan, T. R. Alcoholic hepatitis. N. Engl. J. Med. 360, 2758–2769. https://doi.org/10.1056/NEJMra0805786 (2009).
Dubuquoy, L. et al. Progenitor cell expansion and impaired hepatocyte regeneration in explanted livers from alcoholic hepatitis. Gut 64, 1949–1960. https://doi.org/10.1136/gutjnl-2014-308410 (2015).
Lazaro, R. et al. Osteopontin deficiency does not prevent but promotes alcoholic neutrophilic hepatitis in mice. Hepatology 61, 129–140. https://doi.org/10.1002/hep.27383 (2015).
Grewal, P. & Viswanathen, V. A. Liver cancer and alcohol. Clin. Liver Dis. 16, 839–850. https://doi.org/10.1016/j.cld.2012.08.011 (2012).
Machida, K. et al. Toll-like receptor 4 mediates synergism between alcohol and HCV in hepatic oncogenesis involving stem cell marker Nanog. Proc. Natl. Acad. Sci U S A 106, 1548–1553. https://doi.org/10.1073/pnas.0807390106 (2009).
Malato, Y. et al. Fate tracing of mature hepatocytes in mouse liver homeostasis and regeneration. J. Clin. Invest. 121, 4850–4860. https://doi.org/10.1172/JCI59261 (2011).
Zheng, H. et al. Single-cell analysis reveals cancer stem cell heterogeneity in hepatocellular carcinoma. Hepatology 68, 127–140. https://doi.org/10.1002/hep.29778 (2018).
Zhu, L. et al. Prominin 1 marks intestinal stem cells that are susceptible to neoplastic transformation. Nature 457, 603–607. https://doi.org/10.1038/nature07589 (2009).
He, G. et al. Identification of liver cancer progenitors whose malignant progression depends on autocrine IL-6 signaling. Cell 155, 384–396. https://doi.org/10.1016/j.cell.2013.09.031 (2013).
Zhu, R. et al. TSPAN8 promotes cancer cell stemness via activation of sonic Hedgehog signaling. Nat. Commun. 10(1), 2863. https://doi.org/10.1038/s41467-019-10739-3 (2019).
Barros-Silva, J. D. et al. Single-cell analysis identifies LY6D as a marker linking castration-resistant prostate luminal cells to prostate progenitors and cancer. Cell Rep. 25(12), 3504–3518.e6. https://doi.org/10.1016/j.celrep.2018.11.069 (2018).
Upadhyay, G. Emerging role of lymphocyte Antigen-6 family of genes in cancer and immune cells. Front. Immunol. 10, 819. https://doi.org/10.3389/fimmu.2019.00819 (2019).
Khanova, E. et al. Pyroptosis by caspase11/4-gasdermin-D pathway in alcoholic hepatitis in mice and patients. Hepatology 67, 1737–1753. https://doi.org/10.1002/hep.29645 (2018).
Vogel, W., Gish, G. D., Alves, F. & Pawson, T. The discoidin domain receptor tyrosine kinases are activated by collagen. Mol. Cell. 1, 13–23. https://doi.org/10.1016/s1097-2765(00)80003-9 (1997).
Yang, J. C. et al. TM4SF1 promotes metastasis of pancreatic cancer via regulating the expression of DDR1. Sci. Rep. 7, 45895. https://doi.org/10.1038/srep45895 (2017).
Rikova, K. et al. Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell 131(6), 1190–1203. https://doi.org/10.1016/j.cell.2007.11.025 (2007).
Azizi, R., Salemi, Z., Fallahian, F. & Aghaei, M. Inhibition of discoidin domain receptor 1 reduces epithelial-mesenchymal transition and induce cell-cycle arrest and apoptosis in prostate cancer cell lines. J. Cell Physiol. 234(11), 19539–19552. https://doi.org/10.1002/jcp.28552 (2019).
Jeitany, M. et al. Inhibition of DDR1-BCR signaling by nilotinib as a new therapeutic strategy for metastatic colorectal cancer. EMBO Mol. Med. 10(4), e7918. https://doi.org/10.15252/emmm.201707918 (2018).
Chen, X. et al. Gene expression patterns in human liver cancers. Mol. Biol. Cell. 13, 1929–1939. https://doi.org/10.1091/mbc.02-02-0023 (2002).
Mas, V. R. et al. Genes involved in viral carcinogenesis and tumor initiation in hepatitis C virus-induced hepatocellular carcinoma. Mol. Med. 15, 85–94. https://doi.org/10.2119/molmed.2008.00110 (2009).
Goldman, M. et al. (The Cold Spring Harbor Laboratory, BioRxiv, 2018).
Xue, R. et al. Genomic and transcriptomic profiling of combined hepatocellular and intrahepatic cholangiocarcinoma reveals distinct molecular subtypes. Cancer Cell 35, 932–947.e938. https://doi.org/10.1016/j.ccell.2019.04.007 (2019).
Woo, H. G. et al. Identification of a cholangiocarcinoma-like gene expression trait in hepatocellular carcinoma. Cancer Res 70, 3034–3041. https://doi.org/10.1158/0008-5472.CAN-09-2823 (2010).
Dumble, M. L., Croager, E. J., Yeoh, G. C. & Quail, E. A. Generation and characterization of p53 null transformed hepatic progenitor cells: oval cells give rise to hepatocellular carcinoma. Carcinogenesis 23, 435–445 (2002).
Tarlow, B. D., Finegold, M. J. & Grompe, M. Clonal tracing of Sox9+ liver progenitors in mouse oval cell injury. Hepatology 60, 278–289. https://doi.org/10.1002/hep.27084 (2014).
Español-Suñer, R. et al. Liver progenitor cells yield functional hepatocytes in response to chronic liver injury in mice. Gastroenterology 143, 1564–1575.e1567. https://doi.org/10.1053/j.gastro.2012.08.024 (2012).
Font-Burgada, J. et al. Hybrid periportal hepatocytes regenerate the injured liver without giving rise to cancer. Cell 162, 766–779. https://doi.org/10.1016/j.cell.2015.07.026 (2015).
Han, X. et al. Lineage tracing reveals the bipotency of SOX9. Stem Cell Rep. 12, 624–638. https://doi.org/10.1016/j.stemcr.2019.01.010 (2019).
Deng, X. et al. Chronic liver injury induces conversion of biliary epithelial cells into hepatocytes. Cell Stem Cell 23, 114–122.e113. https://doi.org/10.1016/j.stem.2018.05.022 (2018).
Raven, A. et al. Cholangiocytes act as facultative liver stem cells during impaired hepatocyte regeneration. Nature 547, 350–354. https://doi.org/10.1038/nature23015 (2017).
Ho, D. W. et al. Single-cell transcriptomics reveals the landscape of intra-tumoral heterogeneity and stemness-related subpopulations in liver cancer. Cancer Lett. 459, 176–185. https://doi.org/10.1016/j.canlet.2019.06.002 (2019).
Nejak-Bowen, K. N., Zeng, G., Tan, X., Cieply, B. & Monga, S. P. Beta-catenin regulates vitamin C biosynthesis and cell survival in murine liver. J. Biol. Chem. 284(41), 28115–28127. https://doi.org/10.1074/jbc.M109.047258 (2009).
Gimple, R. C. et al. Glioma stem cell-specific superenhancer promotes polyunsaturated fatty-acid synthesis to support EGFR signaling. Cancer Discov. 9(9), 1248–1267. https://doi.org/10.1158/2159-8290.CD-19-0061 (2019).
Wu, R. et al. Baicalein targets GTPase-mediated autophagy to eliminate liver tumor-initiating stem cell-like cells resistant to mTORC1 inhibition. Hepatology 68(5), 1726–1740. https://doi.org/10.1002/hep.30071 (2018).
Ambrogio, C. et al. Combined inhibition of DDR1 and Notch signaling is a therapeutic strategy for KRAS-driven lung adenocarcinoma. Nat. Med. 22(3), 270–277. https://doi.org/10.1038/nm.4041 (2016).
Hernandez, C. et al. HMGB1 links chronic liver injury to progenitor responses and hepatocarcinogenesis. J. Clin. Invest. 128, 2436–2451. https://doi.org/10.1172/JCI91786 (2018).
Nakano, Y. et al. Identification of a novel alpha-fetoprotein-expressing cell population induced by the Jagged1/Notch2 signal in murine fibrotic liver. Hepatol. Commun. 1, 215–229. https://doi.org/10.1002/hep4.1026 (2017).
Page, A. et al. Alcohol directly stimulates epigenetic modifications in hepatic stellate cells. J. Hepatol. 62, 388–397. https://doi.org/10.1016/j.jhep.2014.09.033 (2015).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049. https://doi.org/10.1038/ncomms14049 (2017).
Sinha, N. et al. Developmental programming: prenatal testosterone-induced epigenetic modulation and its effect on gene expression in sheep ovary†. Biol. Reprod. 102, 1045–1054. https://doi.org/10.1093/biolre/ioaa007 (2020).
Acknowledgements
The authors thank for the services rendered by the Animal Core of the Southern California Research Center for ALPD and Cirrhosis funded by the NIAAA/NIH (P50AA011999). We also thank Sonal Sinha for DEN and Tamoxifen injections. Paraffin embedding and H&E staining were performed by the Liver Histology Core of USC Liver Center (P30DK048522). Flow cytometry was performed in the USC Flow Cytometry Facility that is supported in part by the National Cancer Institute Cancer Center Shared Grant Award (P30CA014089) and the USC Office of the Provost, Dean’s Development Funds, Keck School of Medicine of USC. PROM1+ AH mRNA-seq was performed by USC Norris Molecular Genomics Core. The bioinformatics software and computing resources used in the analysis of PROM1+ AH mRNA-seq are funded by the USC Office of Research and the Norris Medical Library. 10× Single cell RNAseq libraries were also prepared by USC Norris Molecular Genomics Core. Sequencing of single cell RNA-seq libraries and cell ranger analysis were performed by Genewiz. The study was supported by Grants: U01AA027681, U01AA018663, P50AA011999 (pilot project award to R.W.), and R24 AA012885 (Integrative Liver Cell Core) from NIAAA/NIH, and IK6BX004205 (BLR&D Research Career Scientist Award) and I01 BX001991 (VA Merit Review) from Department of Veterans Affairs.
Author information
Authors and Affiliations
Contributions
The project was originally and conceptually developed and supervised until its completion by H.T. R.W. performed the experiments and analyses except those described below. Y.C. performed Partek flow analysis of PROM1+ AH RNA-seq data in Fig. 1. Y.N. performed IF shown in Fig. 2b. S.P. isolated liver cells used in Figs. 1, 3, 4 and 5. H.T. and S.P. generated the TIC transplantation data in Fig. 7e. M.L. provided bioinformatic consultation for Fig. 6b-g. S.B. generated Suppl. Figure 3a-b and Col1a1Cre;Rosa-26mTmG mice. R.W. wrote the manuscript draft which was edited, revised and submitted by H.T.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, R., Pan, S., Chen, Y. et al. Fate and functional roles of Prominin 1+ cells in liver injury and cancer. Sci Rep 10, 19412 (2020). https://doi.org/10.1038/s41598-020-76458-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-76458-8
This article is cited by
-
NOTCH localizes to mitochondria through the TBC1D15-FIS1 interaction and is stabilized via blockade of E3 ligase and CDK8 recruitment to reprogram tumor-initiating cells
Experimental & Molecular Medicine (2024)
-
Hepatic stellate cell stearoyl co-A desaturase activates leukotriene B4 receptor 2 - β-catenin cascade to promote liver tumorigenesis
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.