Key Points
-
Sirtuins regulate protein function by NAD+-dependent post-translational modification, thereby representing a metabolic sensor.
-
Compounds activating sirtuins could be used to treat age-associated mitochondrial diseases.
-
Activation of SIRT1 improves metabolism and global health in organisms ranging from Caenorhabditis elegans to humans.
-
The role of sirtuins in natural longevity is debated, but it is widely accepted that sirtuins have a role in the maintenance of metabolic health.
Abstract
Since the beginning of the century, the mammalian sirtuin protein family (comprising SIRT1–SIRT7) has received much attention for its regulatory role, mainly in metabolism and ageing. Sirtuins act in different cellular compartments: they deacetylate histones and several transcriptional regulators in the nucleus, but also specific proteins in other cellular compartments, such as in the cytoplasm and in mitochondria. As a consequence, sirtuins regulate fat and glucose metabolism in response to physiological changes in energy levels, thereby acting as crucial regulators of the network that controls energy homeostasis and as such determines healthspan.
Similar content being viewed by others
Main
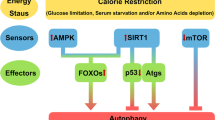
Metabolic control involves a delicate balance among energy intake, utilization and storage. When food is ample, the excess energy is stored so that it can be used in times of scarcity. A carefully tuned regulatory and evolutionarily conserved programme controls these switches in nutrient intake, use and storage, involving classical food excess signalling pathways, such as those revolving around insulin, insulin growth factor 1 (IGF1) and target of rapamycin (TOR; mTOR in mammals), and food restriction pathways involving AMP-activated protein kinase (AMPK) and sirtuins (for further reading on these pathways, see Refs 1,2,3,4).
Sirtuins have received significant attention since the discovery that the yeast sirtuin silent information regulator 2 (Sir2), which was originally described as a regulator of transcriptional silencing of mating-type loci, telomeres and ribosomal DNA (reviewed in Refs 5,6), extends yeast lifespan7. As Sir2 was soon found to be an NAD-dependent histone deacetylase (HDAC)8, it became apparent that sirtuins serve both as energy sensors and as transcriptional effectors by controlling the acetylation state of histones. What is more, sirtuins do not just deacetylate histones, but also a wide range of trascriptional regulators, thereby controlling their activity.
In mammals the sirtuin family comprises seven proteins (SIRT1–SIRT7), which vary in tissue specificity, subcellular localization, enzymatic activity and targets. Sirtuins, notably SIRT1, have been studied for their role in caloric restriction (the only physiological intervention that extends lifespan), the prevention of ageing-related diseases and the maintenance of metabolic homeostasis. As a consequence, the hunt for nutriceutical or pharmaceutical sirtuin activators was intense and led to the identification of several sirtuin activators. Of these, a polyphenol found in red grapes, berries and peanuts that is known as resveratrol received a lot of attention9. Activation of sirtuins is thought to be beneficial not only for diseases relating to metabolism, such as type 2 diabetes and obesity, but also for neurodegenerative diseases, such as Alzheimer's disease and Parkinson's disease. This is in part because sirtuins stimulate the activity of mitochondria, the powerhouses of cells, and of mitochondrial proteins, which have a key role in the above-mentioned pathologies.
In this Review, we present our current knowledge of the sirtuin family, discussing their mode of action, function and regulation. We focus primarily on SIRT1, as it is the best-described sirtuin, and place particular focus on the impact of sirtuins on metabolic homeostasis and healthspan.
The sirtuin family
Sequence-based phylogenetic analysis revealed that mammalian sirtuins can be divided into four classes: SIRT1–SIRT3 belong to class I, SIRT4 to class II, SIRT5 to class III, and SIRT6 and SIRT7 to class IV10. Below we discuss the subcellular localization of sirtuins, their mode of action and their functions in different compartments (Table 1).
Sirtuin subcellular localization. Mammalian sirtuins show a discrete pattern of subcellular localization. SIRT1 is mainly localized in the nucleus but is also present in the cytosol. Its nuclear export signal allows shuttling to the cytosol under specific circumstances, for instance when the insulin pathway is pharmacologically inhibited11. Although the physiological relevance of this shuttling is unclear, it is possible that either cytosolic targets could be deacetylated or that shuttling is another level of control on nuclear target proteins. SIRT2 is considered to be cytosolic but is also present in the nucleus in the G2 phase to M phase transition of the cell cycle12. SIRT3, SIRT4 and SIRT5 have a mitochondrial targeting sequence, and their localization to this organelle has been confirmed experimentally13. SIRT6 is predominantly nuclear14, and SIRT7 was reported to reside in the nucleolus15, but further research is necessary to confirm this localization and its physiological relevance.
Sirtuin enzymatic activity. Originally, sirtuins were described as NAD-dependent type III HDACs, as the founding member, Sir2 in yeast, silenced specific genomic loci by deacetylating histones H3 and H4 (Ref. 16). Interestingly, mammalian sirtuins, notably those in class I, not only target histones but also deacetylate a wide range of proteins in different subcellular compartments (Box 1). In addition, SIRT4 (Ref. 17) and SIRT6 (Ref. 18) were reported to function as ADP-ribosyltransferases, even though SIRT6 also can act as a deacetylase19,20. SIRT5 was initially reported to deacetylate the urea cycle enzyme carbamoyl phosphate synthetase 1 (CPS1)21 but was recently shown to primarily demalonylate and desuccinylate proteins22, including CPS1 (Ref. 23) (Box 1).
The enzymatic reaction catalysed by sirtuins requires NAD+ as a substrate, the concentration of which is determined by the nutritional state of the cell24. As such, NAD+ is well positioned to control adaptive responses to energy stress by modulating the activity of sirtuins and their downstream effectors (see below). Sirtuins convert NAD+ to nicotinamide, which at higher concentrations can non-competitively bind and thereby feedback-inhibit sirtuin activity25,26. The other by-product of the sirtuin deacetylase reaction, O-acetyl-ADP-ribose, was also reported to be a signalling molecule27,28, but similarly to nicotinamide, its exact role in metabolic control requires further investigation (for details see Refs 24,29).
Sirtuin function — SIRT1. The most-studied member of the mammalian sirtuin family is SIRT1, which was originally described to deacetylate histones but soon after was also shown to deacetylate other protein targets (see Refs 5,30,31 for more details) (Table 1). The first-described non-histone target for SIRT1 was p53, which is deacetylated and repressed upon DNA damage or oxidative stress, resulting in impaired apoptosis32,33. It was therefore hypothesized that increased SIRT1 activity could be tumorigenic, but the contrary seems to be the case (reviewed in Ref. 34). The activity of peroxisome proliferator-activated receptor-γ co-activator 1α (PGC1α), which is a transcriptional co-regulator that governs mitochondrial biogenesis and activity, is also controlled by reversible acetylation35,36. PGC1α deacetylation by SIRT1 leads to its activation and to the induction of downstream pathways that control mitochondrial gene expression37,38,39,40,41,42. Similarly, SIRT1 controls the acetylation of forkhead box O (FOXO) transcription factors, which are important regulators of lipid and glucose metabolism as well as of stress responses (see below). It is thought that SIRT1-mediated deacetylation does not just activate or inhibit FOXO but selectively directs FOXO to certain targets, thereby conferring another layer of specificity to regulation by phosphorylation43,44,45 (see below).
Sirtuin function — SIRT3. Three sirtuins localize primarily to mitochondria: SIRT3, SIRT4 and SIRT5 (for recent reviews, see Refs 46,47). Of these, SIRT3 is the major mitochondrial deacetylase48, and several of its targets have been identified, many of which have important roles in metabolic homeostasis. For example, long-chain acyl CoA dehydrogenase (LCAD), a protein involved in fatty acid oxidation, is targeted by SIRT3 during prolonged fasting, resulting in the activation of fatty acid breakdown49. As a result, deletion of Sirt3 in mice impairs fat breakdown, exacerbating diet-induced obesity and rendering them sensitive to cold upon fasting50. SIRT3 deacetylation sites have also been identified in 3-hydroxy-3-methylglutaryl CoA synthase 2 (HMGCS2), which regulates the production of ketone bodies, an important energy source for the brain when blood glucose levels are low51. In response to caloric restriction, SIRT3 deacetylates and thereby activates isocitrate dehydrogenase 2 (IDH2; which is involved in the tricarboxylic acid cycle (TCA cycle))52. SIRT3 also activates the TCA cycle enzyme glutamate dehydrogenase (GDH)48,57, although the physiological relevance of GDH deacetylation is unclear. Moreover, SIRT3 deacetylates components of complex I53, complex II54 and complex III55, which are involved in oxidative phosphorylation (OXPHOS), the final stage of mitochondrial aerobic respiration.
More recently, it became apparent that SIRT3 also affects oxidative stress defence by protecting cells from reactive oxygen species (ROS)52,55,56. Indeed, during caloric restriction, SIRT3 activates superoxide dismutase 2 (SOD2)56, a key mitochondrial antioxidant enzyme. In addition, caloric restriction induces SIRT3-mediated deacetylation of IDH2 in various tissues, including inner ear cells, and thereby increases the ratio of reduced to oxidized glutathione, thus attenuating ROS levels52. As a result, caloric restriction protects against age-related hearing loss in a SIRT3-dependent manner52.
Finally, it is important to note that most data regarding SIRT3 function are derived from the analysis of mice lacking Sirt3 in the germ line, and studies in somatic Sirt3−/− mice will be necessary to address tissue contributions.
Sirtuin function — other sirtuins. Compared with SIRT1 and SIRT3, not much is known about the physiology of the other sirtuins. Cytosolic SIRT2 deacetylates tubulin58, but the relevance of this is unclear. More importantly, SIRT2 also deacetylates partitioning defective 3 homologue (PAR3), which, in turn, decreases the activity of the cell polarity control protein atypical protein kinase C (aPKC), thereby changing myelin formation of Schwann cells59. In addition, SIRT2 may have roles in metabolic homeostasis by deacetylating phosphoenolpyruvate carboxykinase (PEPCK)60, which is involved in gluconeogenesis, and thereby preventing its ubiquitylation-dependent degradation. SIRT2 also deacetylates and activates FOXO1, which is involved in adipogenesis61.
The mitochondrial protein SIRT4 seems to primarily function in metabolism. SIRT4 ADP-ribosylates GDH, thereby inhibiting its activity and blocking amino acid-induced insulin secretion17. As a consequence, Sirt4−/− mice have increased plasma insulin levels, both in fed and fasted state, and when stimulated with Gln17. SIRT4 also regulates fatty acid oxidation in hepatocytes and myocytes, and short hairpin RNA (shRNA)-mediated knockdown of Sirt4 in the liver increased fatty acid oxidation62. It is interesting to note that SIRT3 and SIRT4 have apparently opposing roles in the regulation of GDH17,48,57 and fatty acid oxidation49,62, and further work is needed to define how these two NAD+-dependent enzymes integrate similar nutrient states into divergent responses.
The only target described for SIRT5 is CPS1, deacetylation of which during fasting activates ammonia detoxification through the urea cycle21. SIRT5 may not primarily act as a deacetylase21 but rather as a demalonylase and desuccinylase22, even of the described deacetylase target CPS1 (Ref. 23). Future studies will have to determine whether SIRT5 indeed has deacetylase activity in vivo and whether these different post-translational modifications, even on the same protein, coexist.
SIRT6 is involved in genomic DNA stability and repair and has a role in metabolism and ageing. Sirt6−/− mice die early in life14, have reduced IGF1 levels and are severely hypoglycaemic14, possibly mediated by hypoxia-inducible factor 1α (HIF1α)-dependent activation of glycolysis20. This switch towards glycolysis causes increased glucose uptake in muscle and brown adipose tissue, explaining the fatal hypoglycaemia that these mice develop20. Interestingly, mice with neural-specific ablation of Sirt6 are small at birth owing to reduced growth hormone and IGF1 levels, reach normal body weight at 1 year of age, and become obese later in life, effects that are accompanied by strong hyperacetylation of histone H3Lys9 and H3Lys56 (Ref. 63).
Finally, SIRT7 was reported to activate RNA polymerase I transcription, although its protein substrate is still unknown15. SIRT7 may also deacetylate p53: Sirt7−/− mice display cardiac hypertrophy64, which is linked to p53 hyperacetylation64. However, further studies will have to determine whether p53 is indeed deacetylated by both SIRT1 and SIRT7 and, if so, whether and how these two sirtuins interconnect on this target.
Regulation of sirtuin activity
Regulation of sirtuin activity occurs at various levels. As mentioned above, subcellular localization partly determines activity. However, additional regulation is required, as several sirtuins share the same compartment and also need to show specificity towards distinct substrates. Below we discuss various models of regulation, focusing primarily on the best-described sirtuin, SIRT1.
Regulation by expression. SIRT1 expression changes in various physiological conditions, resulting in induction during low energy status and repression during energy excess states. For instance, nutrient starvation increases SIRT1 expression65, whereas high-fat diet reduces it66. SIRT1 promoter analysis revealed binding sites for various transcription factors including FOXO1 (Ref. 65), CREB (cAMP response element-binding), CHREBP (carbohydrate response element-binding protein)67 and peroxisome proliferator-activated receptors (PPARs)68,69, which suggests that these transcription factors regulate SIRT1 expression in response to these stimuli. Indeed, FOXO1 (Ref. 65), PPARα68, PPARβ (also known as PPARδ)70 and CREB67 increase SIRT1 levels, whereas PPARγ69 and CHREBP67 repress SIRT1 expression (Fig. 1). In addition, HIC1 (hypermethylated in cancer 1) functions as a transcriptional repressor of SIRT1 (Ref. 71). This repression is mediated by the transcriptional repressor CTBP (carboxy-terminal-binding protein) and is enhanced by NADH72 (Fig. 1a), in line with repression of SIRT1 expression in times of energy excess. Finally, poly(ADP-ribose) polymerase 2 (PARP2), which belongs to a family of nuclear enzymes involved in DNA repair, apoptosis and transcription, binds to and represses the SIRT1 promoter (see below), although the exact mechanism is not yet understood73 (Fig. 1a). Importantly, CREB, CHREBP67 and PARP2 (Ref. 73) not only regulate sirtuin expression in vitro but also have been shown to control its expression in vivo.
a | Various transcription factors regulate sirtuin expression. Forkhead box O1 (FOXO1), peroxisome proliferator-activated receptor-α (PPARα), PPARβ and cAMP response element-binding (CREB) enhance SIRT1 expression, whereas PPARγ, carbohydrate response element-binding protein (CHREBP), poly(ADP-ribose) polymerase 2 (PARP2) and hypermethylated in cancer 1 (HIC1) repress SIRT1 expression. Only CREB, CHREBP and PARP2 were also shown to possess this activity in vivo. SIRT1 expression is also repressed by the microRNAs (miRNAs) miR-34a and miR-199a. b | Reversible post-translational modifications affect SIRT1 activity. The cyclin B–CDK1 (cyclin-dependent kinase 1) complex phosphorylates SIRT1, thereby allowing cell cycle progression. Activation of JUN N-terminal kinase (JNK) by reactive oxygen species (ROS) results in SIRT1 phosphorylation and subsequent deacetylation of histone H3. Dual specificity Tyr-phosphorylated and regulated kinase 1 (DYRK1) and DYRK3 activate SIRT1 by phosphorylation, leading to p53 deaceylation and increased cell survival. Genotoxic stress, for instance by ultraviolet (UV) light or hydrogen peroxide (H2O2) exposure, results in desumoylation of SIRT1 by sentrin-specific protease (SENP), thereby inactivating SIRT1. The enzyme that sumoylates and thus activates SIRT1 is unknown. c | Complex formation with other proteins influences SIRT1 enzymatic activity. NCoR1–SMRT (nuclear receptor co-repressor 1–silencing mediator of retinoic acid and thyroid hormone receptor) and SIRT1 block the transcriptional activity of PPARγ. Genotoxic and metabolic (high-fat diet) stress induce deleted in breast cancer 1 (DBC1)–SIRT1 complex formation, leading to SIRT1 inactivation; this is relieved by fasting. The Lys-specific demethylase 1 (LSD1)–SIRT1 complex represses Notch target gene expression by demethylation and deacetylation of specific histones, but this effect is reversed by activation of the Notch pathway. d | Controlling the levels of the cofactor NAD+ governs sirtuin function. NAD+ levels are increased in the presence of the precursors nicotinic acid (NA), nicotinamide (NAM) or nicotinamide riboside (NR) by AMP-activated protein kinase (AMPK) activation following either energy stress or treatment with the AMPK activator resveratrol or following inhibition of NAD+ breakdown by PARP or CD38 inhibitors (PARPi or CD38i, respectively). Increased NAD+ levels lead to sirtuin activation, which in turn leads to the generation of NAM from NAD+. NAM can also feedback-inhibit sirtuins. CTBP, carboxy-terminal-binding protein.
At a different level of regulation, microRNAs (miRNAs) modulate mRNA levels through the degradation of the primary mRNA transcript or by inhibition of translation. As such, mouse miR-34a represses SIRT1 expression following genotoxic stress74. Interestingly, miR-34a-mediated SIRT1 repression was increased in diet-induced obesity75, suggesting a physiological relevance for this interaction. miR-199a also represses SIRT1 expression (Fig. 1a), as well as that of the oxygen sensor HIF1α76. During hypoxia, miR-199a is repressed in cardiac myocytes, allowing SIRT1 and HIF1α expression, thereby stabilizing p53 and reducing apoptosis76.
Although not much is known about the transcriptional control of the other sirtuins, gain-of-function studies in mice revealed that SIRT3 expression is activated by oestrogen-related receptor-α (ERRα), a nuclear receptor that controls mitochondrial function. Together with PGC1α, ERRα binds to the Sirt3 promoter and controls the expression of genes involved in brown adipose tissue development and function77. Whether changes in physiology affect Sirt3 transcriptional regulation is not clear at present, as illustrated by the fact that Sirt3 mRNA expression in mice increased after 1 week of high-fat diet but decreased after 13 weeks of being fed the same diet50.
Regulation by post-translational modifications. Regulation of sirtuin activity by post-translational modifications is poorly understood. Several phosphorylation sites on SIRT1 have been identified78. SIRT1 is phosphorylated in vitro by the cyclin B–CDK1 (cyclin-dependent kinase 1) complex, which binds SIRT1, and mutation of the phosphorylation sites disturbs normal cell cycle progression78 (Fig. 1b). JUN N-terminal kinase (JNK) also phosphorylates SIRT1 at three residues, particularly during oxidative stress79; this results in deacetylation of histone H3, but not of p53 (Ref. 79), suggesting that phosphorylation directs SIRT1 to specific targets (Fig. 1b). In addition, the dual specificity Tyr-phosphorylated and regulated kinases DYRK1 and DYRK3 phosphorylate SIRT1 at Thr522 (Ref. 80). This activating phosphorylation event leads to enhanced SIRT1-mediated p53 deacetylation and prevents apoptosis within the context of genotoxic stress80 (Fig. 1b).
SIRT1 has also been shown to be sumoylated, which in cultured cells increases its activity81. Following genotoxic stress, for example ultraviolet light or hydrogen peroxide, the desumoylating enzyme sentrin-specific protease (SENP) inactivates SIRT1, promoting cell death81 (Fig. 1b). It is tempting to speculate that such conditions also activate the PARP enzymes, a family of major NAD+ consumers, thereby depleting NAD+ levels and inhibiting SIRT1 activity82. It would be interesting to explore whether these two events occur simultaneously.
Regulation by complex formation. Sirtuins are further regulated by forming complexes with other proteins. AROS (active regulator of SIRT1) is the only protein known to positively regulate SIRT1 following complex formation, which leads to suppression of the SIRT1 target p53 (Ref. 83).
By contrast, several negative regulators have been described. NCoR1 (nuclear receptor co-repressor 1) and SMRT (silencing mediator of retinoid and thyroid hormone receptors) form a complex with SIRT1 and PPARγ during fasting to repress the PPARγ-mediated induction of adipogenesis84 (Fig. 1c).
DBC1 (deleted in breast cancer 1) binds the SIRT1 catalytic domain and inhibits its activity in vitro during genotoxic stress85,86 (Fig. 1c). The physiological relevance of the DBC1–SIRT1 complex was confirmed, as complex formation was inhibited during fasting but increased in mice kept on a high-fat diet87. Consistent with this, deletion of Dbc1 resulted in protection against high-fat-diet-induced hepatic steatosis, even though these mice were, surprisingly, slightly heavier than control littermates87.
The histone methyltransferase LSD1 (Lys-specific demethylase 1) also interacts with the SIRT1 catalytic domain to regulate the expression of downstream Notch target genes88. SIRT1-mediated deacetylation of both H4K16 and H1K26, in conjunction with LSD1-mediated demethylation of H3K4, results in convergent repression of Notch target genes88 (Fig. 1c). As such, the abnormal wing development of Drosophila melanogaster Notch mutants can be rescued by deletion of either Sir2 or Lsd1 (Ref. 88) (the fly orthologues of SIRT1 and LSD1, respectively).
Regulation through NAD+. As mentioned above, sirtuins depend on the cofactor NAD+ for their activity, so the availability of NAD+ is another point of regulation. For example, relative NAD+ levels decrease under conditions that stimulate its conversion to its reduced form, NADH39. Specifically, NAD+ levels rise in muscle, liver and white adipose tissue (WAT) during fasting, caloric restriction and exercise38,89, accompanied by sirtuin activation, whereas high-fat diet in mice reduces the NAD+/NADH ratio90.
NAD+ availability also lies at the level of synthesis (Fig. 1d). De novo biosynthesis starts from the amino acid Trp and occurs primarily in the liver and kidney (reviewed in Ref. 24). However, NAD+ can also be synthesized from nicotinic acid (through the Preiss–Handler pathway) or nicotinamide (through the salvage pathway), both of which are present in the human diet as vitamin B3 (Ref. 91). Interestingly, nicotinamide riboside, which is found in milk and is already known to boost NAD+ synthesis in bacteria, was recently shown to act as a NAD+ precursor and to enhance NAD+ synthesis through the salvage pathway in eukaryotic cells92. In addition, treatment with another NAD+ precursor, nicotinamide mononucleotide, activates SIRT1 and improves glucose tolerance in mice93, although it should be noted that this precursor does not occur in the human diet. Future studies will have to determine whether the naturally occurring nicotinic acid, nicotinamide and/or nicotinamide riboside can activate sirtuins in vivo and to clarify the physiological importance of these precursors.
In addition to biosynthesis, manipulating the activities of NAD+-depleting enzymes would also alter NAD+ levels, thereby regulating sirtuin activity (Fig. 1d). PARPs are considered to be the major NAD+ degrading enzymes94,95. When activated by DNA damage, they catalyse the transfer of ADP-ribose units from NAD+ to substrate proteins to form branched polymers of ADP-ribose, leading to the decline of intracellular NAD+ levels24. As SIRT1 and PARPs compete for the same intracellular NAD+ pool, a functional link between PARPs and SIRT1 has been proposed. Indeed, two recent reports revealed that the deletion of Parp1 and Parp2 in mice activates SIRT1, leading to increased numbers of mitochondria, enhanced energy expenditure and protection from diet-induced obesity73,82. Parp1 and Parp2 deletions activate SIRT1 through two distinct mechanisms: deletion of Parp1 boosts intracellular NAD+ levels; and deletion of Parp2, which is a repressor of SIRT1 expression (see above), increases SIRT1 expression. Treatment of cells or mice with the pan-PARP inhibitor PJ34 also results in SIRT1 activation, similarly to what is observed after Parp deletion82. Consistent with nuclear localization of both SIRT1 and PARPs, Parp1 deletion does not increase the deacetylation activity of cytosolic SIRT2 or mitochondrial SIRT3 (Refs 73,82), suggesting that the increase in NAD+ is confined to the nucleus. Deletion of CD38, another important NAD+ consumer, also increases NAD+ levels and SIRT1 activity and mimics the metabolic phenotype expected for SIRT1 activation96. Interestingly, specific CD38 inhibitors have also been developed97, which together with PARP inhibitors open new avenues for pharmacological sirtuin activation. It will be important to determine whether these strategies are specific for SIRT1 or whether more sirtuins will be affected.
Compounds regulating sirtuin activity. Natural compounds that activate SIRT1 have been identified, providing further insights into sirtuin regulation (reviewed in Ref. 98). A small molecule screen in yeast revealed that several plant polyphenols, notably resveratrol, can induce Sir2 to deacetylate p53-based peptides in vitro, and treatment of Saccharomyces cerevisiae with these compounds increased lifespan9, although some doubt surrounds these results (Box 2). Resveratrol also increases SIRT1 activity and enhances mitochondrial function in mice41,42, protecting them from diet-induced obesity and improving exercise performance and resistance to cold41. When kept on a high-fat diet, resveratrol-treated mice also live longer42,99. Resveratrol improves mitochondrial activity and metabolic control in humans as well100. Interestingly, in humans, significantly lower resveratrol doses (∼200-fold lower than the doses given in mice) resulted in similar plasma resveratrol levels, AMP-activated protein kinase (AMPK) activation and physiological effects to those observed following resveratrol treatment in mice100. Several synthetic compounds have also been described that can activate SIRT1. The most potent of these is SRT1720 (Ref. 101), which, similarly to resveratrol, protects against diet-induced obesity by improving mitochondrial function40 and extends the lifespan of obese mice37.
Even though the physiological effects of resveratrol and SRT1720 are widely accepted, the mechanism by which they activate SIRT1 is unclear. Originally, these compounds were thought to directly activate SIRT1 by changing its enzymatic properties, lowering its Michaelis constant ( K m ) for both the protein substrate and NAD+ (Refs 9,101), but more recent reports contest these results38,39,102,103,104,105. Instead of activating SIRT1, resveratrol was shown to activate AMPK40,42, possibly through inhibition of OXPHOS106. Consistent with this, the effect of resveratrol was lost in AMPK-deficient mice38,104, although this does not necessarily demonstrate directionality of the AMPK–SIRT1 interaction. However, the same study showed that resveratrol increased the NAD+/NADH ratio in an AMPK-dependent manner, allowing downstream activation of sirtuins104 (Fig. 1). Similarly, established AMPK agonists, such as 5-aminoimidazole-4-carboxamide ribonucleotide (AICAR), increase NAD+ levels and activate SIRT1-dependent PGC1α deacetylation39, and time-course experiments revealed that AMPK activation by exercise or fasting precedes a rise in NAD+ levels and SIRT1 activation38.
Sirtuins in glucose metabolism
The maintenance of relatively constant blood glucose concentrations is essential to provide energy to tissues, most importantly for an uninterrupted glucose supply to the brain, which almost exclusively uses glucose as an energy source. Glucose levels are regulated through various processes, including intestinal glucose uptake, hepatic glucose output and glucose uptake, utilization and storage in peripheral tissues. The main regulating hormone is insulin, which promotes glucose uptake in peripheral tissues (muscle and WAT), glycolysis and storage of glucose as glycogen in the fed state. Glucagon counteracts the effect of insulin and stimulates hepatic glucose production during fasting. Sirtuins modulate a range of cellular processes involved in maintaining glucose homeostasis in tissues such as muscle, WAT, liver and pancreas (Figs 2a,b,3).
a | SIRT1 inhibits adipogenesis and favours lipolysis in adipose tissue by repressing peroxisome proliferator-activated receptor-γ (PPARγ). In the pancreas, SIRT1 increases insulin secretion by repressing the transcription of uncoupling protein 2 (UCP2). In skeletal muscle, SIRT1 attenuates glycolysis via PPARγ co-activator 1α (PGC1α) and enhances lipid utilization by stimulating PGC1α and PPARα. In the liver, SIRT1 decreases glycolysis via hypoxia-inducible factor 1α (HIF1α) and PGC1α and lowers lipid accumulation by suppressing lipid synthesis through inhibiting sterol-response element-binding protein 1c (SREBP1c) and promoting lipid utilization through PGC1α and PPARα. In addition, SIRT1 regulates hepatic glucose production via PGC1α, CREB-regulated transcription co-activator 2 (CRTC2) and forkhead box O1 (FOXO1), although the exact role of SIRT1 in this process is under debate. b | Based on in vitro studies, SIRT3 promotes cellular respiration in brown adipocytes by activating succinate dehydrogenase (SDH). Using Sirt3 whole-body knockout mice, it has been shown that during energy limitation SIRT3 suppresses glycolysis via HIF1α, promotes fatty acid oxidation by stimulating long-chain acyl CoA dehydrogenase (LCAD), isocitrate dehydrogenase 2 (IDH2) and NADH dehydrogenase ubiquinone 1α subcomplex 9 (NDUFA9), protects from reactive oxygen species (ROS) production by stimulating superoxide dismutase 2 (SOD2) and increases ketone body formation by stimulating 3-hydroxy-3-methylglutaryl CoA synthase 2 (HMGCS2). c | In response to caloric restriction and exercise, induction of SIRT1 stimulates mitochondrial activity, leading to improved metabolism and disease prevention. During energy limitation, low ATP levels activate AMP-activated protein kinase (AMPK), which induces SIRT1 by increasing NAD+ levels. SIRT1 increases mitochondrial activity by decreasing the acetylation levels of PGC1α (acetylation is mediated by general control of amino acid synthesis 5 (GCN5)). During caloric excess and sedentary lifestyle, cellular ATP levels increase, whereas NAD+ levels decrease, thereby inhibiting SIRT1. As a result, PGC1α remains acetylated, which, together with the repressive action of nuclear receptor co-repressor 1 (NCoR1), leads to decreased mitochondrial activity, predisposing to the development of metabolic diseases. Faded circles depict inactivation. ERRα, oestrogen-related receptor-α; NRF1, nuclear respiratory factor 1.
a | The transcriptional regulation of glucose homeostasis in the nucleus. SIRT1 has a dual role in the control of gluconeogenesis: it promotes gluconeogenesis by deacetylating and activating peroxisome proliferator-activated receptor-γ co-activator 1α (PGC1α) and forkhead box O1 (FOXO1), leading to the transcriptional activation of their target genes, but also reduces gluconeogenic gene expression by promoting the degradation of CREB-regulated transcription co-activator 2 (CRTC2) upon deacetylation. SIRT1 attenuates glycolysis but activates lipid utilization and mitochondrial biogenesis by deacetylating and activating PGC1α. Glycolysis is diminished by repressing hypoxia-inducible factor 1α (HIF1α); SIRT1 deacetylates and inhibits HIF1α, and SIRT6 suppresses the transcription of HIF1α. SIRT1 increases insulin secretion by suppressing the transcription of uncoupling protein 2 (UCP2) in the pancreas. Blank ovals represent multiple transcription factors. b | Overview of glucose metabolism and the role of sirtuins; the different stages affected by sirtuins are indicated. Glucose taken up by the cell via glucose transporter (GLUT) is catabolized (glycolysis) into pyruvate, which enters the tricarboxylic acid cycle (TCA cycle) in mitochondria to generate energy. When energy supply is limited, glucose production (gluconeogenesis, dashed arrows) increases by the conversion of pyruvate to oxaloacetate and subsequently to malate. In this case, malate is transported from mitochondria to the cytoplasm, where it is converted back to oxaloacetate, which is used by phosphoenolpyruvate carboxykinase (PEPCK) to produce phosphoenolpyruvate. This can then be converted to glucose (gluconeogenesis). SIRT1 increases insulin secretion by repressing UCP2 transcription. SIRT2 activates gluconeogenesis by increasing the stability of PEPCK. SIRT3 decreases reactive oxygen species (ROS) production by stimulating superoxide dismutase 2 (SOD2), and SIRT3 also enhances cellular respiration by increasing the activities of complex I, complex II (via succinate dehydrogenase (SDH)), complex III and isocitrate dehydrogenase 2 (IDH2). SIRT3 and SIRT4 may regulate gluconeogenesis and insulin secretion by modulating the activity of glutamate dehydrogenase (GDH) through deacetylation and ADP-ribosylation, respectively. Pi, inorganic phosphate.
Gluconeogenesis and insulin sensitivity. Gluconeogenesis is a cytosolic process that increases glucose production, mainly from the liver, in situations when the supply of energy is limiting, for example during fasting, caloric restriction and exercise107. As insulin suppresses gluconeogenesis, fasting hepatic glucose production and circulating glucose levels reflect hepatic insulin sensitivity119. Intriguingly, SIRT1 has a dual and controversial role in the control of gluconeogenesis. On the one hand, SIRT1 can suppress hepatic glucose production by deacetylating CREB-regulated transcription co-activator 2 (CRTC2), leading to CRTC2 degradation and consequent reduction in transcription of gluconeogenic genes108 (Figs 2a,3a). On the other hand, SIRT1 can also stimulate the gluconeogenic transcriptional programme by deacetylating and activating FOXO1 (Refs 108,109) and PGC1α36 (Figs 2a,3a). However, it should be noted that although genetically increased PGC1α expression clearly enhances gluconeogenesis, the evidence supporting the relevance of physiological modulation of PGC1α in the control of gluconeogenesis is weak110. Given that the exact contribution of the activities of CRTC2, FOXO1 and PGC1α to gluconeogenesis is still under debate, it is unclear which of these above-mentioned SIRT1 actions is the primary and/or predominant event. This story has been made more intricate by recent findings showing that temporal regulation of SIRT1 expression and deacetylation of the SIRT1 targets are involved in controlling the appropriate rate of gluconeogenesis67,108
In contrast to SIRT1, SIRT2, SIRT3 and SIRT4 are thought to maintain gluconeogenesis, in particular during times of energy limitation. Specifically, SIRT2 deacetylates and increases the stability of the gluconeogenic enzyme PEPCK upon glucose deprivation60 (Fig. 3b). Furthermore, SIRT3 and SIRT4 may regulate gluconeogenesis from amino acids during caloric restriction through GDH, a mitochondrial enzyme that converts glutamate to α-ketoglutarate, thereby controlling glucose production via the TCA cycle17,48,57 (see below) (Fig. 3b). Further studies are needed to fully understand the role of these sirtuins in the regulation of gluconeogenesis.
Despite the conflicting findings regarding the role of SIRT1 in gluconeogenesis, it is tempting to propose that SIRT1 may function as an insulin sensitizer. This hypothesis is based on observed metabolic changes (lowered fasting glucose levels and/or hepatic glucose production) in SIRT1 transgenic mice111, after adenovirus-mediated SIRT1 overexpression137, in SIRT1 activator-treated mice40,42,101 and in resveratrol-treated obese humans100. Moreover, SIRT1 activation also protects from diet-induced and genetic insulin resistance in mice42,101, and SIRT1 mRNA expression in adipose tissue positively correlates with insulin sensitivity in humans during hyperinsulinaemia112. Mice lacking SIRT1 specifically in the liver have been shown to develop insulin resistance113 or maintain normal glucose homeostasis89,114, whereas acute adenovirus-mediated SIRT1 knockdown in the liver induces fasting hypoglycaemia115. This suggests that the effect on glucose metabolism of acute SIRT1 knockdown in the liver may be compensated for in situations of chronic SIRT1 deficiency. SIRT3 also has a role in insulin sensitization, as its absence may contribute to the development of insulin resistance in the muscle by increasing ROS production and impairing mitochondrial oxidation55. Based on these studies, pharmacological targeting of SIRT1 is a promising treatment for type 2 diabetes.
Glycolysis. Glycolysis is the main pathway for glucose utilization in the fed state, in which glucose is catabolized into pyruvate in the cytosol107. Under anaerobic conditions (for example, in exercising muscle), pyruvate is converted by lactate dehydrogenase into lactate. In aerobic conditions, energy from glucose is released through glucose oxidation, a process in which pyruvate and the reducing equivalents produced during glycolysis fuel mitochondrial OXPHOS to generate ATP.
Recent studies have provided insights into the role of different sirtuins in glycolysis. It is now well established that SIRT1 inhibits glycolysis by activating PGC1α, which attenuates the transcription of glycolytic genes36 (Figs 2a,3a). Furthermore, SIRT1, SIRT3 and SIRT6 all suppress the transcription factor HIF1α, and this leads to decreased glycolysis and increased oxidative metabolism116. SIRT1 suppresses HIF1α directly through deacetylation117 (Figs 2a,3), whereas SIRT3 inhibits ROS-mediated stabilization of HIF1α by activating SOD2 (Ref. 118) and possibly by increasing reduced glutathione levels52, thereby enhancing cellular antioxidant defences (Figs 2b,3b). SIRT6 acts as a co-repressor of HIF1α to diminish glycolysis during the normal nutritional state20 (Fig. 3a).
When integrating these observations, it seems that SIRT1, SIRT3 and SIRT6 inhibit glycolysis and stimulate mitochondrial oxidation of fatty acids. This finding is particularly interesting, as tumour cells have a high reliance on glycolysis, which is known as the Warburg effect. Therefore, the switch from glycolysis to oxidative metabolism induced by sirtuin activators117,118 and PARP inhibitors82, which indirectly activate SIRT1, could contribute to their antitumour activity.
Insulin secretion. The secretion of insulin from pancreatic β-cells is tightly coupled to blood glucose levels through an intricate signalling pathway119. After entering β-cells through glucose transporter 2, glucose is metabolized to generate ATP through glucose oxidation. The subsequent increase in ATP/ADP ratio closes ATP-sensitive potassium channels, leading to β-cell depolarization and calcium influx, ultimately coupling insulin release to plasma glucose levels. Although glucose is the most potent stimulator of insulin secretion, it is also induced by some amino acids119.
SIRT1 has been shown to induce glucose-stimulated insulin secretion by increasing the yield of ATP produced from glucose oxidation. This is achieved through transcriptional repression of uncoupling protein 2 (UCP2) (Figs 2a,3), which uncouples mitochondrial ATP production during OXPHOS. As such, SIRT1 deficiency results in high UCP2 levels and diminished insulin secretion120,121.
As discussed above, both SIRT3 and SIRT4 target GDH (Fig. 3b), although in an opposite manner, and thereby affect amino acid-induced insulin secretion during caloric restriction and feeding. Specifically, SIRT4 seems to inhibit insulin secretion, as SIRT4 deficiency increases GDH activity in pancreatic islet cells and increases amino acid-stimulated insulin secretion during caloric restriction17. Furthermore, SIRT4 overexpression in insulinoma cells leads to decreased insulin secretion in response to glucose122. Although SIRT3 has been proposed to activate GDH48,57, GDH activity is unchanged in Sirt3−/− mice21, so the physiological role of SIRT3 in GDH regulation and insulin secretion is unclear. SIRT1 and SIRT4 therefore seem to control insulin secretion in opposing directions in response to feeding and caloric restriction, respectively. Although the elucidation of underlying mechanisms is under way, current findings suggest the existence of a sirtuin network for maintaining proper insulin secretion.
Sirtuins in lipid metabolism
Lipid metabolism comprises lipid synthesis, uptake, storage and utilization, which requires tight control, for example during periods of fasting or prolonged exercise. Insulin promotes hepatic triglyceride synthesis and storage of triglycerides in WAT upon feeding, whereas glucagon and epinephrin stimulate lipolysis in WAT and fatty acid oxidation in other tissues when nutrients are in limited supply124. Sirtuins influence diverse aspects of lipid homeostasis in multiple tissues; here we concentrate on their effects in WAT, liver and skeletal muscle (Fig. 4). We do not discuss the role of sirtuins in cholesterol and bile acid metabolism (for a review, see Ref. 123).
a | Transcriptional regulation of lipid homeostasis in the nucleus. SIRT1 inhibits lipid synthesis by deacetylating, and thereby destabilizing, sterol-response element-binding protein 1c (SREBP1c), a downstream target of liver X receptor (LXR), which is the key transcription factor involved in lipid synthesis. SIRT1 also inhibits lipid synthesis by suppressing the activity of peroxisome proliferator-activated receptor-γ (PPARγ). SIRT1 promotes fatty acid oxidation by activating PPARα and PPARγ co-activator 1α (PGC1α) and promoting the expression of their target genes. Blank ovals represent multiple transcription factors. b | Overview of the lipid cycle showing where sirtuins act. Once taken up by the cell by fatty acid transporters, fatty acids are transported to mitochondria, where they are oxidized to produce ATP. Fat breakdown can also lead to the generation of ketone bodies through the action of 3-hydroxy-3-methylglutaryl CoA synthase 2 (HMGCS2). Fatty acids can be synthesized (lipogenesis, dashed arrows) in the cytosol from malonyl CoA by fatty acid synthase and then converted to triglycerides. During periods of high energy demand, triglycerides can be broken down to free fatty acids (lipolysis, occurring mainly in fat tissue), which are then released into the circulation. SIRT1 reduces fatty acid storage by enhancing lipolysis through the inhibition of PPARγ and by decreasing fatty acid synthesis through SREBP1c. In addition, SIRT6 may act as a repressor of genes involved in fatty acid synthesis. SIRT3 stimulates β-oxidation and ketone body formation by targeting and activating long-chain acyl CoA dehydrogenase (LCAD) and HMGCS2, respectively. Furthermore, SIRT3 decreases reactive oxygen species (ROS) production by stimulating superoxide dismutase 2 (SOD2), and SIRT3 also enhances cellular respiration by increasing the activities of complex I, complex II, complex III and isocitrate dehydrogenase 2 (IDH2). SIRT4 seems to dampen the transcription of genes governing fatty acid oxidation. NCoR1, nuclear receptor co-repressor 1; Pi, inorganic phosphate; SDH, succinate dehydrogenase; SMRT, silencing mediator of retinoid and thyroid hormone receptors.
Lipid synthesis. When whole-body energy stores are maximal, excess glucose, fatty acids and amino acids are used in the liver to synthesize fatty acids, which are exported to WAT, where they are stored as triacylglycerols124. Fatty acid synthesis occurs in the cytosol, where acetyl CoA, derived from fuel catabolism, and malonyl CoA are used as substrates for fatty acid production, catalysed by fatty acid synthase. A key transcription factor controlling the expression of genes involved in lipid synthesis is liver X receptor (LXR), which in part mediates its effect through the induction of sterol-response element-binding protein 1c (SREBP1c)125. SIRT1 deacetylates and subsequently increases the transcriptional activity of LXR126. Therefore, SIRT1 activation could theoretically increase fatty acid synthesis. However, SIRT1 also deacetylates the downstream target of LXR, SREBP1c, thereby destabilizing it and reducing its occupancy on lipogenic gene promoters, leading to suppression of fatty acid synthesis127,128 (Figs 2a,4). This notion is supported by the fact that Sirt1-overexpressing mice are protected from hepatic steatosis129, whereas Sirt1−/− mice are prone to develop it114,130 (see also below). Therefore, SIRT1-mediated LXR activation seems to target cholesterol metabolism (for a review, see Ref. 123) rather than triglyceride synthesis.
SIRT6 has also been implicated in the control of fatty acid synthesis (Fig. 4b). Indeed, genes involved in fatty acid transport and lipogenesis are induced in mice lacking SIRT6 specifically in the liver131, suggesting that SIRT6 serves as a negative regulator of triglyceride synthesis.
Lipid storage. Fatty acids can be stored as cytosolic triacylglycerol droplets in all cells, but the prime storage occurs in adipocytes of WAT132. The development of adipocytes from pre-adipocytes is controlled by a complex transcriptional programme coordinated mainly by the nuclear receptor PPARγ, the master regulator of adipocyte differentiation133.
In differentiated adipocyte cell lines, SIRT1 inhibits adipogenesis and enhances fat mobilization through lipolysis84 by suppressing the activity of PPARγ. SIRT1 achieves this by promoting the assembly of a co-repressor complex, involving NCoR1 and SMRT, on the promoters of PPARγ target genes to repress their transcription (Figs 2a,4a) and thus limit fat storage in situations of caloric restriction and fasting84. Cell-based studies have suggested that SIRT2 may also inhibit adipogenesis and promote lipolysis during nutrient deprivation by deacetylating and activating FOXO1; SIRT1 promotes the binding of FOXO1 to PPARγ, thereby repressing its transcriptional activity61,134. However, further research is required to establish the existence of a mechanistic link between these sirtuins and PPARγ and FOXO1 in vivo.
Lipid utilization and energy expenditure. Fatty acid oxidation occurs mainly in the mitochondrial matrix. Therefore, long-chain fatty acids need first to be transported from the cytoplasm into mitochondria135, where they undergo β-oxidation to generate acetyl CoAs, which can be used for ATP production via the TCA cycle and OXPHOS. As malonyl CoA, an intermediate of fatty acid synthesis, inhibits carnitine palmitoyltransferase, the enzyme controlling the mitochondrial import of fatty acids, low levels of malonyl CoA facilitate the import and oxidation of fatty acids.
Fatty acid oxidation is a major determinant of whole-body energy expenditure, as it consumes high amounts of oxygen. SIRT1 enhances energy expenditure by stimulating fatty acid oxidation, as well as OXPHOS, in response to fasting. This is achieved by activating PPARα and one of its key co-activators, PGC1α114, which stimulate the expression of genes involved in fatty acid uptake and/or β-oxidation (Figs 2a,4a). In support of this, SIRT1 activation protects mice from diet-induced obesity by increasing the rate of fatty acid oxidation40,41. In humans, SIRT1 mRNA expression positively correlates with energy expenditure during hyperinsulinaemic clamp112 (the gold standard for measuring insulin sensitivity), and three Sirt1single-nucleotide polymorphisms are associated with whole-body energy expenditure in Finnish subjects41. These observations confirm the important and conserved role of SIRT1 in modulating whole-body energy expenditure.
Fatty acids are an important source of energy in the liver, including during fasting, caloric restriction and high-fat feeding. SIRT1 has been shown to promote fatty acid oxidation in the liver. Indeed, induction of fatty acid oxidation through the activation of PPARα and PGC1α and their target genes is impaired during fasting in mice lacking SIRT1 specifically in the liver114,115. As reduced fatty acid oxidation correlates with hepatic steatosis, liver-specific and heterozygous germline Sirt1−/− mice are susceptible to hepatic steatosis under standard and high-fat diets114,130,136. Moreover, hepatic steatosis is improved in diet-induced and genetically obese mouse models following adenoviral overexpression of Sirt1 (Ref. 137) and pharmacological SIRT1 activation40,127,128,138,139, as well as in Sirt1-overexpressing mice129, and hepatic fat content is reduced by resveratrol treatment in obese humans100. Surprisingly, one strain of liver-specific Sirt1−/− mice is protected from hepatic steatosis and diet-induced obesity89. Despite this, most findings support SIRT1 stimulating lipid utilization.
Of the other sirtuins, SIRT3, SIRT4 and SIRT6 are also involved in the regulation of fatty acid oxidation in the liver and may also be potential therapeutic targets for treating liver diseases characterized by lipid accumulation. As discussed above, SIRT3 deacetylates and activates LCAD49 (Figs 2b,4b), a key enzyme involved in the oxidation of long-chain fatty acids. By contrast, SIRT4 seems to be a negative regulator of fatty acid oxidation62 (Fig. 4b), whereas SIRT6 was reported to stimulate the expression of genes involved in fatty acid oxidation131, although the exact mechanism is unclear. Importantly, both SIRT3-deficient mice and liver-specific Sirt6−/− mice are more susceptible to hepatic steatosis49,131.
Skeletal muscle uses fatty acids as fuel during fasting, prolonged exercise and high-fat feeding. The switch to oxidative metabolism includes the induction of mitochondrial biogenesis, fatty acid oxidation and OXPHOS, processes that are controlled by the 'yin and yang' between co-repressors (such as NCoR1 (Ref. 140)) and co-activators (such as PGC1α141,142), which determine the transcriptional activity of downstream targets controlling oxidative metabolism. However, what are the key events leading to the shift towards more oxidative metabolism in skeletal muscle? AMPK is a cellular energy sensor that is activated during exercise or energy demand by an increase in the AMP/ATP ratio2. Activated AMPK phosphorylates PGC1α directly143 and also stimulates SIRT1 indirectly by boosting cellular NAD+ levels39 (Fig. 2c). Nevertheless, the activation of SIRT1 may be independent of AMPK in skeletal muscle during caloric restriction144,145, although this is not a consistent finding146. Once SIRT1 is activated by NAD+, it deacetylates and locks PGC1α in an active state36, which together with the reduced activity of NCoR1 (Ref. 140) will favour oxidative metabolism. Recent data have implied that the nuclear abundance of the key PGC1α acetyltransferase, GCN5 (general control of amino-acid synthesis 5)35, is also diminished in response to exercise, which could also contribute to the decreased PGC1α acetylation147. However, further work is needed to determine the function and regulation of co-activators (such as GCN5 (Ref. 35)) and co-repressors (such as NCoR1) in the control of oxidative metabolism. In addition to the role of SIRT1, SIRT3-dependent deacetylation and activation of LCAD49 and OXPHOS complexes53,54,55, as discussed above, might affect fatty acid oxidation in skeletal muscle.
Interestingly, in states of energy excess (for instance, following high-fat feeding), these processes, which typify situations of high energy demand, are reversed. As such, cellular AMPK and SIRT1 activities are attenuated owing to high intracellular ATP levels and low NAD+ levels148. Furthermore, high-fat feeding induces the expression of GCN5 while reducing SIRT1 levels, resulting in the inhibition of PGC1α by increasing its acetylation level66 (Fig. 2c). Finally, NCoR1 is activated during energy excess, leading to decreased transcription of genes governing mitochondrial activity140.
Conclusions and future perspectives
Sirtuins constitute a protein family of metabolic sensors, translating changes in NAD+ levels into adaptive responses. Although the relevance of SIRT1 as a sensu stricto longevity gene has been disputed (Box 2), it is evident that by regulating the activity of their target enzymes and transcription factors, sirtuins affect health in a pleiotropic manner. The fact that sirtuin activation prevents diet-induced obesity and that their overexpression prevents cancer risk suggests that sirtuins are primarily involved in stress responses. This implies that activation or ectopic expression of sirtuins does not affect physiological fitness when the organism is not stressed by, for example, genotoxic or metabolic (high-fat diet) stress or ageing. However, following such external stressors, enhanced sirtuin activity activates protective pathways, and can thereby affect lifespan. As such, sirtuins should still be considered as candidate targets for preventing and/or treating age-related diseases and for increasing healthspan. In fact, in contrast to increasing lifespan, which has limited medical relevance, improving healthspan has an immediate clinical and public health impact, given the ever increasing 'greying' of the world population.
Besides the relevance of sirtuin activators within the context of common diseases of ageing, it would be worth testing sirtuin activators for the treatment of more rare, but also more insidious, inherited diseases relating to mitochondrial metabolism. This was recently exemplified by treating fibroblasts of patients with a mitochondrial fatty acid oxidation disorder with resveratrol, which resulted in restoration of fatty acid degradation158. Altogether, although sirtuins might have lost their immaculate Methuselah image, they are still likely to help those in need of a metabolic 'Samaritan'.
References
Zoncu, R., Efeyan, A. & Sabatini, D. M. mTOR: from growth signal integration to cancer, diabetes and ageing. Nature Rev. Mol. Cell Biol. 12, 21–35 (2011).
Cantó, C. & Auwerx, J. AMP-activated protein kinase and its downstream transcriptional pathways. Cell. Mol. Life Sci. 67, 3407–3423 (2010).
Houtkooper, R. H., Williams, R. W. & Auwerx, J. Metabolic networks of longevity. Cell 142, 9–14 (2010).
Fontana, L., Partridge, L. & Longo, V. D. Extending healthy life span — from yeast to humans. Science 328, 321–326 (2010).
Haigis, M. C. & Sinclair, D. A. Mammalian sirtuins: biological insights and disease relevance. Annu. Rev. Pathol. 5, 253–295 (2010).
Guarente, L. Franklin & H. Epstein lecture: sirtuins, aging, and medicine. N. Engl. J. Med. 364, 2235–2244 (2011).
Kaeberlein, M., McVey, M. & Guarente, L. The SIR2/3/4 complex and SIR2 alone promote longevity in Saccharomyces cerevisiae by two different mechanisms. Genes Dev. 13, 2570–2580 (1999).
Imai, S., Armstrong, C. M., Kaeberlein, M. & Guarente, L. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature 403, 795–800 (2000). This study describes the NAD dependence of yeast Sir2.
Howitz, K. T. et al. Small molecule activators of sirtuins extend Saccharomyces cerevisiae lifespan. Nature 425, 191–196 (2003). This report describes the first small molecule screen for sirtuin activators, identifying resveratrol as a caloric restriction mimetic.
Frye, R. A. Phylogenetic classification of prokaryotic and eukaryotic Sir2-like proteins. Biochem. Biophys. Res. Commun. 273, 793–798 (2000).
Tanno, M., Sakamoto, J., Miura, T., Shimamoto, K. & Horio, Y. Nucleocytoplasmic shuttling of the NAD+-dependent histone deacetylase SIRT1. J. Biol. Chem. 282, 6823–6832 (2007).
Vaquero, A. et al. SirT2 is a histone deacetylase with preference for histone H4 Lys 16 during mitosis. Genes Dev. 20, 1256–1261 (2006).
Huang, J. Y., Hirschey, M. D., Shimazu, T., Ho, L. & Verdin, E. Mitochondrial sirtuins. Biochim. Biophys. Acta 1804, 1645–1651 (2010).
Mostoslavsky, R. et al. Genomic instability and aging-like phenotype in the absence of mammalian SIRT6. Cell 124, 315–329 (2006).
Ford, E. et al. Mammalian Sir2 homolog SIRT7 is an activator of RNA polymerase I transcription. Genes Dev. 20, 1075–1080 (2006).
Braunstein, M., Sobel, R. E., Allis, C. D., Turner, B. M. & Broach, J. R. Efficient transcriptional silencing in Saccharomyces cerevisiae requires a heterochromatin histone acetylation pattern. Mol. Cell. Biol. 16, 4349–4356 (1996).
Haigis, M. C. et al. SIRT4 inhibits glutamate dehydrogenase and opposes the effects of calorie restriction in pancreatic β-cells. Cell 126, 941–954 (2006).
Liszt, G., Ford, E., Kurtev, M. & Guarente, L. Mouse Sir2 homolog SIRT6 is a nuclear ADP-ribosyltransferase. J. Biol. Chem. 280, 21313–21320 (2005).
Michishita, E. et al. SIRT6 is a histone H3 lysine 9 deacetylase that modulates telomeric chromatin. Nature 452, 492–496 (2008).
Zhong, L. et al. The histone deacetylase Sirt6 regulates glucose homeostasis via Hif1α. Cell 140, 280–293 (2010).
Nakagawa, T., Lomb, D. J., Haigis, M. C. & Guarente, L. SIRT5 deacetylates carbamoyl phosphate synthetase 1 and regulates the urea cycle. Cell 137, 560–570 (2009).
Peng, C. et al. The first identification of lysine malonylation substrates and its regulatory enzyme. Mol. Cell. Proteomics 9 Sep 2011 (doi:10.1074/mcp.M111.012658).
Du, J. et al. Sirt5 is an NAD-dependent protein lysine demalonylase and desuccinylase. Science 334, 806–809 (2011). References 22 and 23 describe desuccinylation and demalonylation as a novel function for SIRT5. Reference 23 identifies CPS1 as a desuccinylated target.
Houtkooper, R. H., Cantó, C., Wanders, R. J. & Auwerx, J. The secret life of NAD+: an old metabolite controlling new metabolic signaling pathways. Endocr. Rev. 31, 194–223 (2010).
Bitterman, K. J., Anderson, R. M., Cohen, H. Y., Latorre-Esteves, M. & Sinclair, D. A. Inhibition of silencing and accelerated aging by nicotinamide, a putative negative regulator of yeast sir2 and human SIRT1. J. Biol. Chem. 277, 45099–45107 (2002).
Anderson, R. M., Bitterman, K. J., Wood, J. G., Medvedik, O. & Sinclair, D. A. Nicotinamide and PNC1 govern lifespan extension by calorie restriction in Saccharomyces cerevisiae. Nature 423, 181–185 (2003).
Kustatscher, G., Hothorn, M., Pugieux, C., Scheffzek, K. & Ladurner, A. G. Splicing regulates NAD metabolite binding to histone macroH2A. Nature Struct. Mol. Biol. 12, 624–625 (2005).
Liou, G. G., Tanny, J. C., Kruger, R. G., Walz, T. & Moazed, D. Assembly of the SIR complex and its regulation by O-acetyl-ADP-ribose, a product of NAD-dependent histone deacetylation. Cell 121, 515–527 (2005).
Tong, L. & Denu, J. M. Function and metabolism of sirtuin metabolite O-acetyl-ADP-ribose. Biochim. Biophys. Acta 1804, 1617–1625 (2010).
Feige, J. N. & Auwerx, J. Transcriptional targets of sirtuins in the coordination of mammalian physiology. Curr. Opin. Cell Biol. 20, 303–309 (2008).
Cantó, C. & Auwerx, J. Targeting sirtuin 1 to improve metabolism: all you need is NAD+? Pharmacol. Rev. 64, 166–187 (2011).
Vaziri, H. et al. hSIR2(SIRT1) functions as an NAD-dependent p53 deacetylase. Cell 107, 149–159 (2001).
Luo, J. et al. Negative control of p53 by Sir2α promotes cell survival under stress. Cell 107, 137–148 (2001).
Herranz, D. & Serrano, M. SIRT1: recent lessons from mouse models. Nature Rev. Cancer 10, 819–823 (2010).
Lerin, C. et al. GCN5 acetyltransferase complex controls glucose metabolism through transcriptional repression of PGC-1α. Cell Metab. 3, 429–438 (2006).
Rodgers, J. T. et al. Nutrient control of glucose homeostasis through a complex of PGC-1α and SIRT1. Nature 434, 113–118 (2005). This article provides the mechanistic link between SIRT1 activity and PGC1α acetylation.
Minor, R. K. et al. SRT1720 improves survival and healthspan of obese mice. Sci. Rep. 1, 70 (2011).
Cantó, C. et al. Interdependence of AMPK and SIRT1 for metabolic adaptation to fasting and exercise in skeletal muscle. Cell Metab. 11, 213–219 (2010).
Cantó, C. et al. AMPK regulates energy expenditure by modulating NAD+ metabolism and SIRT1 activity. Nature 458, 1056–1060 (2009). This paper describes how NAD+ levels link AMPK and SIRT1 activity.
Feige, J. N. et al. Specific SIRT1 activation mimics low energy levels and protects against diet-induced metabolic disorders by enhancing fat oxidation. Cell Metab. 8, 347–358 (2008).
Lagouge, M. et al. Resveratrol improves mitochondrial function and protects against metabolic disease by activating SIRT1 and PGC-1α. Cell 127, 1109–1122 (2006).
Baur, J. A. et al. Resveratrol improves health and survival of mice on a high-calorie diet. Nature 444, 337–342 (2006). References 41 and 42 describe the beneficial effects of resveratrol treatment in mice, showing improved metabolic profile.
Brunet, A. et al. Stress-dependent regulation of FOXO transcription factors by the SIRT1 deacetylase. Science 303, 2011–2015 (2004).
Motta, M. C. et al. Mammalian SIRT1 represses forkhead transcription factors. Cell 116, 551–563 (2004).
van der Horst, A. et al. FOXO4 is acetylated upon peroxide stress and deacetylated by the longevity protein hSir2(SIRT1). J. Biol. Chem. 279, 28873–28879 (2004).
Zhong, L. & Mostoslavsky, R. Fine tuning our cellular factories: sirtuins in mitochondrial biology. Cell Metab. 13, 621–626 (2011).
Verdin, E., Hirschey, M. D., Finley, L. W. & Haigis, M. C. Sirtuin regulation of mitochondria: energy production, apoptosis, and signaling. Trends Biochem. Sci. 35, 669–675 (2010).
Lombard, D. B. et al. Mammalian Sir2 homolog SIRT3 regulates global mitochondrial lysine acetylation. Mol. Cell. Biol. 27, 8807–8814 (2007).
Hirschey, M. D. et al. SIRT3 regulates mitochondrial fatty-acid oxidation by reversible enzyme deacetylation. Nature 464, 121–125 (2010). This study describes the fatty acid oxidation enzyme LCAD as a SIRT3 target and its role during fasting.
Hirschey, M. D. et al. SIRT3 deficiency and mitochondrial protein hyperacetylation accelerate the development of the metabolic syndrome. Mol. Cell 21, 177–190 (2011).
Shimazu, T. et al. SIRT3 deacetylates mitochondrial 3-hydroxy-3-methylglutaryl CoA synthase 2 and regulates ketone body production. Cell Metab. 12, 654–661 (2010).
Someya, S. et al. Sirt3 mediates reduction of oxidative damage and prevention of age-related hearing loss under caloric restriction. Cell 143, 802–812 (2010). This article shows how SIRT3 regulates IDH2 activity and oxidative stress defence and thereby mediates the effects of caloric restriction on age-related hearing loss.
Ahn, B. H. et al. A role for the mitochondrial deacetylase Sirt3 in regulating energy homeostasis. Proc. Natl Acad. Sci. USA 105, 14447–14452 (2008).
Finley, L. W. et al. Succinate dehydrogenase is a direct target of sirtuin 3 deacetylase activity. PLoS ONE 6, e23295 (2011).
Jing, E. et al. Sirtuin-3 (Sirt3) regulates skeletal muscle metabolism and insulin signaling via altered mitochondrial oxidation and reactive oxygen species production. Proc. Natl Acad. Sci. USA 108, 14608–14613 (2011).
Qiu, X., Brown, K., Hirschey, M. D., Verdin, E. & Chen, D. Calorie restriction reduces oxidative stress by SIRT3-mediated SOD2 activation. Cell Metab. 12, 662–667 (2010).
Schlicker, C. et al. Substrates and regulation mechanisms for the human mitochondrial sirtuins Sirt3 and Sirt5. J. Mol. Biol. 382, 790–801 (2008).
North, B. J., Marshall, B. L., Borra, M. T., Denu, J. M. & Verdin, E. The human Sir2 ortholog, SIRT2, is an NAD+-dependent tubulin deacetylase. Mol. Cell 11, 437–444 (2003).
Beirowski, B. et al. Sir-two-homolog 2 (Sirt2) modulates peripheral myelination through polarity protein Par-3/atypical protein kinase C (aPKC) signaling. Proc. Natl Acad. Sci. USA 108, E952–E961 (2011).
Jiang, W. et al. Acetylation regulates gluconeogenesis by promoting PEPCK1 degradation via recruiting the UBR5 ubiquitin ligase. Mol. Cell 43, 33–44 (2011).
Jing, E., Gesta, S. & Kahn, C. R. SIRT2 regulates adipocyte differentiation through FoxO1 acetylation/deacetylation. Cell Metab. 6, 105–114 (2007).
Nasrin, N. et al. SIRT4 regulates fatty acid oxidation and mitochondrial gene expression in liver and muscle cells. J. Biol. Chem. 285, 31995–32002 (2010).
Schwer, B. et al. Neural sirtuin 6 (Sirt6) ablation attenuates somatic growth and causes obesity. Proc. Natl Acad. Sci. USA 107, 21790–21794 (2010).
Vakhrusheva, O. et al. Sirt7 increases stress resistance of cardiomyocytes and prevents apoptosis and inflammatory cardiomyopathy in mice. Circ. Res. 102, 703–710 (2008).
Nemoto, S., Fergusson, M. M. & Finkel, T. Nutrient availability regulates SIRT1 through a forkhead-dependent pathway. Science 306, 2105–2108 (2004).
Coste, A. et al. The genetic ablation of SRC-3 protects against obesity and improves insulin sensitivity by reducing the acetylation of PGC-1α. Proc. Natl Acad. Sci. USA 105, 17187–17192 (2008).
Noriega, L. G. et al. CREB and ChREBP oppositely regulate SIRT1 expression in response to energy availability. EMBO Rep. 12, 1069–1076 (2011).
Hayashida, S. et al. Fasting promotes the expression of SIRT1, an NAD+-dependent protein deacetylase, via activation of PPARα in mice. Mol. Cell. Biochem. 339, 285–292 (2010).
Han, L. et al. SIRT1 is regulated by a PPARγ–SIRT1 negative feedback loop associated with senescence. Nucleic Acids Res. 38, 7458–7471 (2010).
Okazaki, M. et al. PPARβ/δ regulates the human SIRT1 gene transcription via Sp1. Endocr. J. 57, 403–413 (2010).
Chen, W. Y. et al. Tumor suppressor HIC1 directly regulates SIRT1 to modulate p53-dependent DNA-damage responses. Cell 123, 437–448 (2005).
Zhang, Q. et al. Metabolic regulation of SIRT1 transcription via a HIC1:CtBP corepressor complex. Proc. Natl Acad. Sci. USA 104, 829–833 (2007).
Bai, P. et al. PARP-2 regulates SIRT1 expression and whole-body energy expenditure. Cell Metab. 13, 450–460 (2011).
Yamakuchi, M., Ferlito, M. & Lowenstein, C. J. miR-34a repression of SIRT1 regulates apoptosis. Proc. Natl Acad. Sci. USA 105, 13421–13426, (2008).
Lee, J. et al. A pathway involving farnesoid X receptor and small heterodimer partner positively regulates hepatic sirtuin 1 levels via microRNA-34a inhibition. J. Biol. Chem. 285, 12604–12611 (2010).
Rane, S. et al. Downregulation of miR-199a derepresses hypoxia-inducible factor-1α and sirtuin 1 and recapitulates hypoxia preconditioning in cardiac myocytes. Circ. Res. 104, 879–886 (2009).
Giralt, A. et al. Peroxisome proliferator-activated receptor-γ coactivator-1α controls transcription of the Sirt3 gene, an essential component of the thermogenic brown adipocyte phenotype. J. Biol. Chem. 286, 16958–16966 (2011).
Sasaki, T. et al. Phosphorylation regulates SIRT1 function. PLoS ONE 3, e4020, (2008).
Nasrin, N. et al. JNK1 phosphorylates SIRT1 and promotes its enzymatic activity. PLoS ONE 4, e8414 (2009).
Guo, X., Williams, J. G., Schug, T. T. & Li, X. DYRK1A and DYRK3 promote cell survival through phosphorylation and activation of SIRT1. J. Biol. Chem. 285, 13223–13232 (2010).
Yang, Y. et al. SIRT1 sumoylation regulates its deacetylase activity and cellular response to genotoxic stress. Nature Cell Biol. 9, 1253–1262 (2007).
Bai, P. et al. PARP-1 inhibition increases mitochondrial metabolism through SIRT1 activation. Cell Metab. 13, 461–468 (2011). This report demonstrates how the interplay between different NAD+ consumers can regulate SIRT1 activity.
Kim, E. J., Kho, J. H., Kang, M. R. & Um, S. J. Active regulator of SIRT1 cooperates with SIRT1 and facilitates suppression of p53 activity. Mol. Cell 28, 277–290 (2007).
Picard, F. et al. Sirt1 promotes fat mobilization in white adipocytes by repressing PPAR-γ. Nature 429, 771–776 (2004).
Kim, J. E., Chen, J. & Lou, Z. DBC1 is a negative regulator of SIRT1. Nature 451, 583–586 (2008).
Zhao, W. et al. Negative regulation of the deacetylase SIRT1 by DBC1. Nature 451, 587–590 (2008).
Escande, C. et al. Deleted in breast cancer-1 regulates SIRT1 activity and contributes to high-fat diet-induced liver steatosis in mice. J. Clin. Invest. 120, 545–558 (2010).
Mulligan, P. et al. A SIRT1–LSD1 corepressor complex regulates Notch target gene expression and development. Mol. Cell 42, 689–699 (2011).
Chen, D. et al. Tissue-specific regulation of SIRT1 by calorie restriction. Genes Dev. 22, 1753–1757 (2008).
Kim, H. J. et al. Metabolomic analysis of livers and serum from high-fat diet induced obese mice. J. Proteome Res. 10, 722–731 (2011).
Collins, P. B. & Chaykin, S. The management of nicotinamide and nicotinic acid in the mouse. J. Biol. Chem. 247, 778–783 (1972).
Bieganowski, P. & Brenner, C. Discoveries of nicotinamide riboside as a nutrient and conserved NRK genes establish a Preiss–Handler independent route to NAD+ in fungi and humans. Cell 117, 495–502 (2004).
Yoshino, J., Mills, K. F., Yoon, M. J. & Imai, S. Nicotinamide mononucleotide, a key NAD+ intermediate, treats the pathophysiology of diet- and age-induced diabetes in mice. Cell Metab. 14, 528–536 (2011).
Schreiber, V., Dantzer, F., Ame, J. C. & de Murcia, G. Poly(ADP-ribose): novel functions for an old molecule. Nature Rev. Mol. Cell Biol. 7, 517–528 (2006).
Krishnakumar, R. & Kraus, W. L. The PARP side of the nucleus: molecular actions, physiological outcomes, and clinical targets. Mol. Cell 39, 8–24 (2010).
Barbosa, M. T. et al. The enzyme CD38 (a NAD glycohydrolase, EC 3.2.2.5) is necessary for the development of diet-induced obesity. FASEB J. 21, 3629–3639 (2007).
Dong, M. et al. Design, synthesis and biological characterization of novel inhibitors of CD38. Org. Biomol. Chem. 9, 3246–3257 (2011).
Lavu, S., Boss, O., Elliott, P. J. & Lambert, P. D. Sirtuins — novel therapeutic targets to treat age-associated diseases. Nature Rev. Drug Discov. 7, 841–853 (2008).
Pearson, K. J. et al. Resveratrol delays age-related deterioration and mimics transcriptional aspects of dietary restriction without extending life span. Cell Metab. 8, 157–168 (2008).
Timmers, S. et al. Calorie restriction-like effects of 30 days of resveratrol supplementation on energy metabolism and metabolic profile in obese humans. Cell Metab. 14, 612–622 (2011). This paper is the first description of resveratrol treatment in humans, mimicking the effects of caloric restriction by showing improved (mitochondrial) metabolism in obese subjects.
Milne, J. C. et al. Small molecule activators of SIRT1 as therapeutics for the treatment of type 2 diabetes. Nature 450, 712–716 (2007).
Pacholec, M. et al. SRT1720, SRT2183, SRT1460, and resveratrol are not direct activators of SIRT1. J. Biol. Chem. 285, 8340–8351 (2010).
Beher, D. et al. Resveratrol is not a direct activator of SIRT1 enzyme activity. Chem. Biol. Drug Des. 74, 619–624 (2009).
Um, J. H. et al. AMP-activated protein kinase-deficient mice are resistant to the metabolic effects of resveratrol. Diabetes 59, 554–563 (2010).
Hawley, S. A. et al. Use of cells expressing γ-subunit variants to identify diverse mechanisms of AMPK activation. Cell Metab. 11, 554–565 (2010).
Zheng, J. & Ramirez, V. D. Inhibition of mitochondrial proton F0F1-ATPase/ATP synthase by polyphenolic phytochemicals. Br. J. Pharmacol. 130, 1115–1123 (2000).
Bouche, C., Serdy, S., Kahn, C. R. & Goldfine, A. B. The cellular fate of glucose and its relevance in type 2 diabetes. Endocr. Rev. 25, 807–830 (2004).
Liu, Y. et al. A fasting inducible switch modulates gluconeogenesis via activator/coactivator exchange. Nature 456, 269–273 (2008). This study describes how SIRT1, CRTC2 and FOXO1 are temporally regulated during fasting.
Frescas, D., Valenti, L. & Accili, D. Nuclear trapping of the forkhead transcription factor FoxO1 via Sirt-dependent deacetylation promotes expression of glucogenetic genes. J. Biol. Chem. 280, 20589–20595 (2005).
Herzog, B., Hall, R. K., Wang, X. L., Waltner-Law, M. & Granner, D. K. Peroxisome proliferator-activated receptor-γ coactivator-1α, as a transcription amplifier, is not essential for basal and hormone-induced phosphoenolpyruvate carboxykinase gene expression. Mol. Endocrinol. 18, 807–819 (2004).
Bordone, L. et al. SIRT1 transgenic mice show phenotypes resembling calorie restriction. Aging Cell 6, 759–767 (2007).
Rutanen, J. et al. SIRT1 mRNA expression may be associated with energy expenditure and insulin sensitivity. Diabetes 59, 829–835 (2010).
Wang, R. H. et al. Hepatic Sirt1 deficiency in mice impairs mTorc2/Akt signaling and results in hyperglycemia, oxidative damage, and insulin resistance. J. Clin. Invest. 121, 4477–4490 (2011). This article demonstrates that hepatic SIRT1 deficiency causes whole-body insulin resistance owing to hyperglycaemia-induced oxidative stress.
Purushotham, A. et al. Hepatocyte-specific deletion of SIRT1 alters fatty acid metabolism and results in hepatic steatosis and inflammation. Cell Metab. 9, 327–338 (2009). This study shows that hepatic SIRT1 deletion increases susceptibility to hepatic steatosis and body weight gain upon high-fat feeding.
Rodgers, J. T. & Puigserver, P. Fasting-dependent glucose and lipid metabolic response through hepatic sirtuin 1. Proc. Natl Acad. Sci. USA 104, 12861–12866 (2007).
Kim, J. W., Tchernyshyov, I., Semenza, G. L. & Dang, C. V. HIF-1-mediated expression of pyruvate dehydrogenase kinase: a metabolic switch required for cellular adaptation to hypoxia. Cell Metab. 3, 177–185 (2006).
Lim, J. H. et al. Sirtuin 1 modulates cellular responses to hypoxia by deacetylating hypoxia-inducible factor 1α. Mol. Cell 38, 864–878 (2010).
Finley, L. W. et al. SIRT3 opposes reprogramming of cancer cell metabolism through HIF1α destabilization. Cancer Cell 19, 416–428 (2011).
Muoio, D. M. & Newgard, C. B. Mechanisms of disease: molecular and metabolic mechanisms of insulin resistance and β-cell failure in type 2 diabetes. Nature Rev. Mol. Cell Biol. 9, 193–205 (2008).
Moynihan, K. A. et al. Increased dosage of mammalian Sir2 in pancreatic β-cells enhances glucose-stimulated insulin secretion in mice. Cell Metab. 2, 105–117 (2005).
Bordone, L. et al. Sirt1 regulates insulin secretion by repressing UCP2 in pancreatic β-cells. PLoS Biol. 4, e31 (2006).
Ahuja, N. et al. Regulation of insulin secretion by SIRT4, a mitochondrial ADP-ribosyltransferase. J. Biol. Chem. 282, 33583–33592 (2007).
Schug, T. T. & Li, X. Sirtuin 1 in lipid metabolism and obesity. Ann. Med. 43, 198–211 (2011).
Strable, M. S. & Ntambi, J. M. Genetic control of de novo lipogenesis: role in diet-induced obesity. Crit. Rev. Biochem. Mol. Biol. 45, 199–214 (2010).
Horton, J. D., Goldstein, J. L. & Brown, M. S. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J. Clin. Invest. 109, 1125–1131 (2002).
Li, X. et al. SIRT1 deacetylates and positively regulates the nuclear receptor LXR. Mol. Cell 28, 91–106 (2007).
Ponugoti, B. et al. SIRT1 deacetylates and inhibits SREBP-1C activity in regulation of hepatic lipid metabolism. J. Biol. Chem. 285, 33959–33970 (2010).
Walker, A. K. et al. Conserved role of SIRT1 orthologs in fasting-dependent inhibition of the lipid/cholesterol regulator SREBP. Genes Dev. 24, 1403–1417 (2010).
Pfluger, P. T., Herranz, D., Velasco-Miguel, S., Serrano, M. & Tschop, M. H. Sirt1 protects against high-fat diet-induced metabolic damage. Proc. Natl Acad. Sci. USA 105, 9793–9798 (2008).
Wang, R. H., Li, C. & Deng, C. X. Liver steatosis and increased ChREBP expression in mice carrying a liver specific SIRT1 null mutation under a normal feeding condition. Int. J. Biol. Sci. 6, 682–690 (2010).
Kim, H. S. et al. Hepatic-specific disruption of SIRT6 in mice results in fatty liver formation due to enhanced glycolysis and triglyceride synthesis. Cell Metab. 12, 224–236 (2010).
Wang, P., Mariman, E., Renes, J. & Keijer, J. The secretory function of adipocytes in the physiology of white adipose tissue. J. Cell Physiol. 216, 3–13 (2008).
Heikkinen, S., Auwerx, J. & Argmann, C. A. PPARγ in human and mouse physiology. Biochim. Biophys. Acta 1771, 999–1013 (2007).
Wang, F. & Tong, Q. SIRT2 suppresses adipocyte differentiation by deacetylating FOXO1 and enhancing FOXO1's repressive interaction with PPARγ Mol. Biol. Cell 20, 801–808 (2009).
Schreurs, M., Kuipers, F. & van der Leij, F. R. Regulatory enzymes of mitochondrial β-oxidation as targets for treatment of the metabolic syndrome. Obes. Rev. 11, 380–388 (2010).
Xu, F. et al. Lack of SIRT1 (mammalian sirtuin 1) activity leads to liver steatosis in the SIRT1± mice: a role of lipid mobilization and inflammation. Endocrinology 151, 2504–2514 (2010).
Li, Y. et al. Hepatic overexpression of SIRT1 in mice attenuates endoplasmic reticulum stress and insulin resistance in the liver. FASEB J. 25, 1664–1679 (2011).
Ajmo, J. M., Liang, X., Rogers, C. Q., Pennock, B. & You, M. Resveratrol alleviates alcoholic fatty liver in mice. Am. J. Physiol. Gastrointest. Liver Physiol. 295, 833–842 (2008).
Yamazaki, Y. et al. Treatment with SRT1720, a SIRT1 activator, ameliorates fatty liver with reduced expression of lipogenic enzymes in MSG mice. Am. J. Physiol. Endocrinol. Metab. 297, 1179–1186 (2009).
Yamamoto, H. et al. NCoR1 is a conserved physiological modulator of muscle mass and oxidative function. Cell 147, 827–839 (2011). This paper shows that reduced co-repressor activity can improve metabolism similarly to enhanced co-activator activity.
Handschin, C. & Spiegelman, B. M. Peroxisome proliferator-activated receptor-γ coactivator 1 coactivators, energy homeostasis, and metabolism. Endocr. Rev. 27, 728–735 (2006).
Fernandez-Marcos, P. J. & Auwerx, J. Regulation of PGC-1α, a nodal regulator of mitochondrial biogenesis. Am. J. Clin. Nutr. 93, 884S–890S (2011).
Jager, S., Handschin, C., St- Pierre, J. & Spiegelman, B. M. AMP-activated protein kinase (AMPK) action in skeletal muscle via direct phosphorylation of PGC-1α. Proc. Natl Acad. Sci. USA 104, 12017–12022 (2007).
Gonzalez, A. A., Kumar, R., Mulligan, J. D., Davis, A. J. & Saupe, K. W. Effects of aging on cardiac and skeletal muscle AMPK activity: basal activity, allosteric activation, and response to in vivo hypoxemia in mice. Am. J. Physiol. Regul. Integr. Comp. Physiol. 287, R1270–R1275 (2004).
Schenk, S. et al. Sirt1 enhances skeletal muscle insulin sensitivity in mice during caloric restriction. J. Clin. Invest. 121, 4281–4288 (2011). This paper demonstrates that caloric restriction-induced insulin sensitivity is mediated by SIRT1 in skeletal muscle in mice.
Palacios, O. M. et al. Diet and exercise signals regulate SIRT3 and activate AMPK and PGC-1α in skeletal muscle. Aging 1, 771–783 (2009).
Philp, A. et al. Sirtuin 1 (SIRT1) deacetylase activity is not required for mitochondrial biogenesis or peroxisome proliferator-activated receptor-γ coactivator-1α (PGC-1α) deacetylation following endurance exercise. J. Biol. Chem. 286, 30561–30570 (2011).
Cantó, C. & Auwerx, J. PGC-1α, SIRT1 and AMPK, an energy sensing network that controls energy expenditure. Curr. Opin. Lipidol. 20, 98–105 (2009).
Tissenbaum, H. A. & Guarente, L. Increased dosage of a sir-2 gene extends lifespan in Caenorhabditis elegans. Nature 410, 227–230 (2001).
Rogina, B. & Helfand, S. L. Sir2 mediates longevity in the fly through a pathway related to calorie restriction. Proc. Natl Acad. Sci. USA 101, 15998–16003 (2004).
Bass, T. M., Weinkove, D., Houthoofd, K., Gems, D. & Partridge, L. Effects of resveratrol on lifespan in Drosophila melanogaster and Caenorhabditis elegans. Mech. Ageing Dev. 128, 546–552 (2007).
Kaeberlein, M. & Powers, R. W. Sir2 and calorie restriction in yeast: a skeptical perspective. Ageing Res. Rev. 6, 128–140 (2007).
Burnett, C. et al. Absence of effects of Sir2 overexpression on lifespan in C. elegans and Drosophila. Nature 477, 482–485 (2011). This study shows that Sir2 and sir-2.1 overexpression is not sufficient to extend lifespan in flies and worms, respectively.
Viswanathan, M. & Guarente, L. Regulation of Caenorhabditis elegans lifespan by sir-2.1 transgenes. Nature 477, 1–2 (2011).
Li, Y., Xu, W., McBurney, M. W. & Longo, V. D. SirT1 inhibition reduces IGF-I/IRS-2/Ras/ERK1/2 signaling and protects neurons. Cell Metab. 8, 38–48 (2008).
Herranz, D. et al. Sirt1 improves healthy ageing and protects from metabolic syndrome-associated cancer. Nature Commun. 1, 3 (2010). This report demonstates that SIRT1 overexpression in mice improves healthy ageing but does not extend lifespan.
Flachsbart, F. et al. Sirtuin 1 (SIRT1) sequence variation is not associated with exceptional human longevity. Exp. Gerontol. 41, 98–102 (2006).
Bastin, J., Lopes-Costa, A. & Djouadi, F. Exposure to resveratrol triggers pharmacological correction of fatty acid utilization in human fatty acid oxidation-deficient fibroblasts. Hum. Mol. Genet. 20, 2048–2057 (2011).
Acknowledgements
The authors apologize to colleagues whose original work could not be cited owing to space limitations. The authors thank team members of the Auwerx laboratory for discussions. R.H.H. is supported by a Rubicon fellowship of the Netherlands Organization for Scientific Research (NWO), and E.P. is funded by the Academy of Finland and the Finnish Cultural Foundation. The work in the Auwerx laboratory is supported by grants of the École Polytechnique Fédérale de Lausanne, Faculty of Life Science, the European Union Ideas programme (ERC-2008-AdG-23118), the Velux Stiftung and the Swiss National Science Foundation (31003A-124713 and 31003A-125487).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Caloric restriction
-
A reduction of caloric intake (typically 20–50% less than average) that has been shown to increase lifespan in a variety of organisms.
- Nutriceutical
-
A food product that provides health benefits.
- ADP-ribosyltransferases
-
Enzymes that transfer an ADP-ribose group on a protein target.
- Demalonylate
-
The act of removing a malonyl group from a specific Lys residue of a target protein.
- Desuccinylate
-
The act of removing a succinyl group from a specific Lys residue of a target protein.
- Ketone bodies
-
Metabolic energy units derived from fat breakdown. Ketone bodies serve to fuel the brain in times of low blood glucose concentrations.
- Tricarboxylic acid cycle
-
(TCA cycle). Also known as Krebs cycle. Acetyl-CoA derived from glucose or fat breakdown is further metabolized to offer reduced energy equivalents that are used by mitochondrial oxidative phosphorylation to generate ATP.
- Oxidative phosphorylation
-
(OXPHOS). Mitochondrial enzymatic chain of events by which ATP is generated.
- Reactive oxygen species
-
(ROS). Oxygen radicals that are produced as by-products of oxidative phosphorylation in mitochondria. In excess, they can cause intracellular and mitochondrial damage, which promotes cell death.
- Steatosis
-
A condition characterized by abnormal accumulation of lipids within the cell.
- K m
-
(Michaelis constant). Enzymatic property describing the substrate concentration at which the enzyme works at half-maximal capacity.
- Hyperinsulinaemia
-
A condition that occurs when there is excess insulin in the circulation.
- Fasting hypoglycaemia
-
A condition during fasting that occurs when circulating glucose levels are abnormally low.
- Insulinoma cells
-
Tumour cells derived from pancreatic β-cells that secrete insulin.
- Single-nucleotide polymorphisms
-
Naturally occurring single nucleotide variations in a DNA sequence. The most common type of genetic polymorphism.
Rights and permissions
About this article
Cite this article
Houtkooper, R., Pirinen, E. & Auwerx, J. Sirtuins as regulators of metabolism and healthspan. Nat Rev Mol Cell Biol 13, 225–238 (2012). https://doi.org/10.1038/nrm3293
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3293
This article is cited by
-
Amifostine ameliorates bleomycin-induced murine pulmonary fibrosis via NAD+/SIRT1/AMPK pathway-mediated effects on mitochondrial function and cellular metabolism
European Journal of Medical Research (2024)
-
Dysregulation of histone deacetylases in ocular diseases
Archives of Pharmacal Research (2024)
-
SIRT6 in Regulation of Mitochondrial Damage and Associated Cardiac Dysfunctions: A Possible Therapeutic Target for CVDs
Cardiovascular Toxicology (2024)
-
Induction of Oxidative Stress in Sirtuin Gene-Disrupted Ashbya gossypii Mutants Overproducing Riboflavin
Molecular Biotechnology (2024)
-
CYB5R3 functions as a tumor suppressor by inducing ER stress-mediated apoptosis in lung cancer cells via the PERK-ATF4 and IRE1α-JNK pathways
Experimental & Molecular Medicine (2024)