Key Points
-
Predicting cis-regulatory modules (CRMs) can both expand our understanding of biology and have applications in medicine and other fields. For example, most of the genetic variants that are significantly associated with susceptibility to disease are not in protein-coding regions, and we surmise that many affect regulation of gene expression.
-
Clusters of transcription factor binding sites (TFBSs) do not provide sufficient specificity to be used for effective prediction of CRMs in large-scale investigations. However, when applied to identified or likely CRMs, they provide important insights into potentially cooperating transcription factors and help to build a regulatory code.
-
Predicting CRMs on the basis of strong constraint in non-coding sequences finds an important subset of the CRMs: that is, those that control developmental regulatory genes. However, they miss a large number of, possibly most, transcription-factor-occupied segments.
-
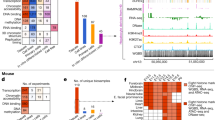
High-quality data sets of epigenetic features associated with gene regulation, such as DNase-hypersensitive sites, transcription factor occupancy and diagnostic histone modifications, provide a reasonably unbiased, sensitive view of the regulatory landscape. The integrative analysis of these features is currently the best approach for predicting CRMs.
-
Evolutionary and motif patterns in predicted CRMs should be used to partition the identified regions into categories that could have different functions in regulation.
-
Predicting and testing CRMs is essential for deciphering a regulatory code. Synthetic biology approaches, in which DNA sequences inferred for a particular function are synthesized and tested for that function, can assess the accuracy of models for regulatory codes and point to needed improvements.
Abstract
Differential gene expression is the fundamental mechanism underlying animal development and cell differentiation. However, it is a challenge to identify comprehensively and accurately the DNA sequences that are required to regulate gene expression: namely, cis-regulatory modules (CRMs). Three major features, either singly or in combination, are used to predict CRMs: clusters of transcription factor binding site motifs, non-coding DNA that is under evolutionary constraint and biochemical marks associated with CRMs, such as histone modifications and protein occupancy. The validation rates for predictions indicate that identifying diagnostic biochemical marks is the most reliable method, and understanding is enhanced by the analysis of motifs and conservation patterns within those predicted CRMs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Davidson, E. H. & Erwin, D. H. Gene regulatory networks and the evolution of animal body plans. Science 311, 796–800 (2006).
King, M. C. & Wilson, A. C. Evolution at two levels in humans and chimpanzees. Science 188, 107–116 (1975).
Hindorff, L. A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl Acad. Sci. USA 106, 9362–9367 (2009).
Tuupanen, S. et al. The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling. Nature Genet. 41, 885–890 (2009).
Jia, L. et al. Functional enhancers at the gene-poor 8q24 cancer-linked locus. PLoS Genet 5, e1000597 (2009).
Harismendy, O. et al. 9p21 DNA variants associated with coronary artery disease impair interferon-γ signalling response. Nature 470, 264–268 (2011).
Farrell, J. J. et al. A 3 bp deletion in the HBS1L-MYB intergenic region on chromosome 6q23 is associated with HbF expression. Blood 117, 4935–4945 (2011).
Maston, G. A., Evans, S. K. & Green, M. R. Transcriptional regulatory elements in the human genome. Annu. Rev. Genom. Hum. Genet. 7, 29–59 (2006).
Schones, D. E. & Zhao, K. Genome-wide approaches to studying chromatin modifications. Nature Rev. Genet. 9, 179–191 (2008).
Rando, O. J. & Chang, H. Y. Genome-wide views of chromatin structure. Annu. Rev. Biochem. 78, 245–271 (2009).
Noonan, J. P. & McCallion, A. S. Genomics of long-range regulatory elements. Annu. Rev. Genom. Hum. Genet. 11, 1–23 (2010).
Hawkins, R. D., Hon, G. C. & Ren, B. Next-generation genomics: an integrative approach. Nature Rev. Genet. 11, 476–486 (2010).
Lenhard, B., Sandelin, A. & Carninci, P. Metazoan promoters: emerging characteristics and insights into transcriptional regulation. Nature Rev. Genet. 13, 233–245 (2012).
Frazer, K. A., Elnitski, L., Church, D., Dubchak, I. & Hardison, R. C. Cross-species sequence comparisons: a review of methods and available resources. Genome Res. 13, 1–12 (2003).
Wasserman, W. W. & Sandelin, A. Applied bioinformatics for the identification of regulatory elements. Nature Rev. Genet. 5, 276–287 (2004).
Elnitski, L., Jin, V. X., Farnham, P. J. & Jones, S. J. Locating mammalian transcription factor binding sites: a survey of computational and experimental techniques. Genome Res. 16, 1455–1464 (2006).
Su, J., Teichmann, S. A. & Down, T. A. Assessing computational methods of cis-regulatory module prediction. PLoS Comput. Biol. 6, e1001020 (2010). This paper provides a comprehensive evaluation and comparison of sequence-based computational approaches for identifying cis -regulatory modules in human and D. melanogaster using large test sets.
Zhang, Y. et al. Primary sequence and epigenetic determinants of in vivo occupancy of genomic DNA by GATA1. Nucleic Acids Res. 37, 7024–7038 (2009).
Cao, Y. et al. Genome-wide MyoD binding in skeletal muscle cells: a potential for broad cellular reprogramming. Dev. Cell 18, 662–674 (2010).
Cheng, Y. et al. Erythroid GATA1 function revealed by genome-wide analysis of transcription factor occupancy, histone modifications, and mRNA expression. Genome Res. 19, 2172–2184 (2009).
Pribnow, D. Nucleotide sequence of an RNA polymerase binding site at an early T7 promoter. Proc. Natl Acad. Sci. USA 72, 784–788 (1975).
Carninci, P. et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nature Genet. 38, 626–635 (2006). This paper demonstrated that high-throughput sequencing of the 5′ ends of transcripts can be used to identify transcription start sites. Using this approach revealed different classes of mammalian promoter architecture.
Banerji, J., Rusconi, S. & Schaffner, W. Expression of a β-globin gene is enhanced by remote SV40 DNA sequences. Cell 27, 299–308 (1981).
Fromm, M. & Berg, P. Simian virus 40 early- and late-region promoter functions are enhanced by the 72 base-pair repeat inserted at distant locations and inverted orientations. Mol. Cell. Biol. 3, 991–999 (1983).
Gillies, S. D., Morrison, S. L., Oi, V. T. & Tonegawa, S. A tissue-specific transcription enhancer element is located in the major intron of a rearranged immunoglobulin heavy chain gene. Cell 33, 717–728 (1983).
Rusche, L. N., Kirchmaier, A. L. & Rine, J. The establishment, inheritance, and function of silenced chromatin in Saccharomyces cerevisiae. Annu. Rev. Biochem. 72, 481–516 (2003).
Martowicz, M. L., Grass, J. A., Boyer, M. E., Guend, H. & Bresnick, E. H. Dynamic GATA factor interplay at a multicomponent regulatory region of the GATA-2 locus. J. Biol. Chem. 280, 1724–1732 (2005).
Jing, H. et al. Exchange of GATA factors mediates transitions in looped chromatin organization at a developmentally regulated gene locus. Mol. Cell 29, 232–242 (2008).
Maniatis, T., Goodbourn, S. & Fischer, J. A. Regulation of inducible and tissue-specific gene expression. Science 236, 1237–1245 (1987).
Lettice, L. A. et al. A long-range Shh enhancer regulates expression in the developing limb and fin and is associated with preaxial polydactyly. Hum. Mol. Genet. 12, 1725–1735 (2003).
Schirm, S., Jiricny, J. & Schaffner, W. The SV40 enhancer can be dissected into multiple segments, each with a different cell type specificity. Genes Dev. 1, 65–74 (1987).
Ondek, B., Gross, L. & Herr, W. The SV40 enhancer contains two distinct levels of organization. Nature 333, 40–45 (1988).
Arnosti, D. N., Barolo, S., Levine, M. & Small, S. The eve stripe 2 enhancer employs multiple modes of transcriptional synergy. Development 122, 205–214 (1996).
Pennacchio, L. A. et al. In vivo enhancer analysis of human conserved non-coding sequences. Nature 444, 499–502 (2006). The authors used extreme evolutionary conservation to identify predicted enhancers and showed that half of those tested drove reproducible tissue-specific expression patterns in mouse embryos.
Landry, J. R. et al. Expression of the leukemia oncogene Lmo2 is controlled by an array of tissue-specific elements dispersed over 100 kb and bound by Tal1/Lmo2, Ets, and Gata factors. Blood 113, 5783–5792 (2009).
Valenzuela, L. & Kamakaka, R. T. Chromatin insulators. Annu. Rev. Genet. 40, 107–138 (2006).
Wallace, J. A. & Felsenfeld, G. We gather together: insulators and genome organization. Curr. Opin. Genet. Dev. 17, 400–407 (2007).
Chung, J. H., Whiteley, M. & Felsenfeld, G. A. 5′ element of the chicken β-globin domain serves as an insulator in human erythroid cells and protects against position effect in Drosophila. Cell 74, 505–514 (1993).
Bell, A. C., West, A. G. & Felsenfeld, G. The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell 98, 387–396 (1999).
Schoborg, T. A. & Labrador, M. The phylogenetic distribution of non-CTCF insulator proteins is limited to insects and reveals that BEAF-32 is Drosophila lineage specific. J. Mol. Evol. 70, 74–84 (2010).
Recillas-Targa, F. et al. Position-effect protection and enhancer blocking by the chicken β-globin insulator are separable activities. Proc. Natl Acad. Sci. USA 99, 6883–6888 (2002).
Huang, S., Li, X., Yusufzai, T. M., Qiu, Y. & Felsenfeld, G. USF1 recruits histone modification complexes and is critical for maintenance of a chromatin barrier. Mol. Cell. Biol. 27, 7991–8002 (2007).
Phillips, J. E. & Corces, V. G. CTCF: master weaver of the genome. Cell 137, 1194–1211 (2009).
Kim, T. H. et al. Analysis of the vertebrate insulator protein CTCF-binding sites in the human genome. Cell 128, 1231–1245 (2007).
Wasserman, W. W. & Fickett, J. W. Identification of regulatory regions which confer muscle-specific gene expression. J. Mol. Biol. 278, 167–181 (1998). This was one of the first analyses to combine motif discovery and sequence conservation in a predictive model that identifies tissue-specific regulatory sequences.
Frith, M. C., Spouge, J. L., Hansen, U. & Weng, Z. Statistical significance of clusters of motifs represented by position specific scoring matrices in nucleotide sequences. Nucleic Acids Res. 30, 3214–3224 (2002).
Berman, B. P. et al. Exploiting transcription factor binding site clustering to identify cis-regulatory modules involved in pattern formation in the Drosophila genome. Proc. Natl Acad. Sci. USA 99, 757–762 (2002).
Markstein, M., Markstein, P., Markstein, V. & Levine, M. S. Genome-wide analysis of clustered Dorsal binding sites identifies putative target genes in the Drosophila embryo. Proc. Natl Acad. Sci. USA 99, 763–768 (2002).
Rebeiz, M., Reeves, N. L. & Posakony, J. W. SCORE: a computational approach to the identification of cis-regulatory modules and target genes in whole-genome sequence data. Site clustering over random expectation. Proc. Natl Acad. Sci. USA 99, 9888–9893 (2002).
Halfon, M. S., Grad, Y., Church, G. M. & Michelson, A. M. Computation-based discovery of related transcriptional regulatory modules and motifs using an experimentally validated combinatorial model. Genome Res. 12, 1019–1028 (2002).
Schroeder, M. D. et al. Transcriptional control in the segmentation gene network of Drosophila. PLoS Biol. 2, E271 (2004).
Zhou, Q. & Wong, W. H. CisModule: de novo discovery of cis-regulatory modules by hierarchical mixture modeling. Proc. Natl Acad. Sci. USA 101, 12114–12119 (2004).
Smith, A. D., Sumazin, P., Xuan, Z. & Zhang, M. Q. DNA motifs in human and mouse proximal promoters predict tissue-specific expression. Proc. Natl Acad. Sci. USA 103, 6275–6280 (2006).
Gertz, J., Siggia, E. D. & Cohen, B. A. Analysis of combinatorial cis-regulation in synthetic and genomic promoters. Nature 457, 215–218 (2009).
Chan, E. T. et al. Conservation of core gene expression in vertebrate tissues. J. Biol. 8, 33 (2009).
Ludwig, M. Z. et al. Functional evolution of a cis-regulatory module. PLoS Biol. 3, e93 (2005).
Hardison, R., Oeltjen, J. & Miller, W. Long human-mouse sequence alignments reveal novel regulatory elements: a reason to sequence the mouse genome. Genome Res. 7, 959–966 (1997).
Hardison, R. C. Conserved noncoding sequences are reliable guides to regulatory elements. Trends Genet. 16, 369–372 (2000).
Pennacchio, L. A. & Rubin, E. M. Genomic strategies to identify mammalian regulatory sequences. Nature Rev. Genet. 2, 100–109 (2001).
Dermitzakis, E. T., Reymond, A. & Antonarakis, S. E. Conserved non-genic sequences — an unexpected feature of mammalian genomes. Nature Rev. Genet. 6, 151–157 (2005).
Tagle, D. A. et al. Embryonic χ and γ globin genes of a prosimian primate (Galago crassicaudatus): nucleotide and amino acid sequences, developmental regulation and phylogenetic footprints. J. Mol. Biol. 203, 7469–7480 (1988).
Gumucio, D. L. et al. Phylogenetic footprinting reveals a nuclear protein which binds to silencer sequences in the human γ and χ globin genes. Mol. Cell. Biol. 12, 4919–4929 (1992).
Hardison, R. et al. Comparative analysis of the locus control region of the rabbit beta-like globin gene cluster: HS3 increases transient expression of an embryonic χ-globin gene. Nucl. Acids Res. 21, 1265–1272 (1993).
Elnitski, L., Miller, W. & Hardison, R. Conserved E boxes function as part of the enhancer in hypersensitive site 2 of the β-globin locus control region: role of basic helix-loop-helix proteins. J. Biol. Chem. 272, 369–378 (1997).
Xie, X. et al. Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434, 338–345 (2005).
Stark, A. et al. Discovery of functional elements in 12 Drosophila genomes using evolutionary signatures. Nature 450, 219–232 (2007).
Kheradpour, P., Stark, A., Roy, S. & Kellis, M. Reliable prediction of regulator targets using 12 Drosophila genomes. Genome Res. 17, 1919–1931 (2007).
Emorine, L., Kuehl, M., Weir, L., Leder, P. & Max, E. E. A conserved sequence in the immunoglobulin Jk-Ck intron: possible enhancer element. Nature 304, 447–449 (1983).
Loots, G. G. et al. Identification of a coordinate regulator of interleukins 4, 13, and 5 by cross-species sequence comparisons. Science 288, 136–140 (2000).
Frazer, K. A. et al. Noncoding sequences conserved in a limited number of mammals in the SIM2 interval are frequently functional. Genome Res. 14, 367–372 (2004).
Grice, E. A., Rochelle, E. S., Green, E. D., Chakravarti, A. & McCallion, A. S. Evaluation of the RET regulatory landscape reveals the biological relevance of a HSCR-implicated enhancer. Hum. Mol. Genet. 14, 3837–3845 (2005).
Johnson, D. S., Davidson, B., Brown, C. D., Smith, W. C. & Sidow, A. Noncoding regulatory sequences of Ciona exhibit strong correspondence between evolutionary constraint and functional importance. Genome Res. 14, 2448–2456 (2004).
Woolfe, A. et al. Highly conserved non-coding sequences are associated with vertebrate development. PLoS Biol. 3, e7 (2005). This paper demonstrated that many non-coding regions that are conserved between human and Fugu rubripes show tissue-specific enhancer functions in zebrafish embryos.
Visel, A. et al. Ultraconservation identifies a small subset of extremely constrained developmental enhancers. Nature Genet. 40, 158–160 (2008).
Attanasio, C. et al. Assaying the regulatory potential of mammalian conserved non-coding sequences in human cells. Genome Biol. 9, R168 (2008).
Clark, A. G. et al. Evolution of genes and genomes on the Drosophila phylogeny. Nature 450, 203–218 (2007).
Loots, G. G., Ovcharenko, I., Pachter, L., Dubchak, I. & Rubin, E. M. rVista for comparative sequence-based discovery of functional transcription factor binding sites. Genome Res. 12, 832–839 (2002).
Gibbs, R. A. et al. Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428, 493–521 (2004).
Sinha, S., Schroeder, M. D., Unnerstall, U., Gaul, U. & Siggia, E. D. Cross-species comparison significantly improves genome-wide prediction of cis-regulatory modules in Drosophila. BMC Bioinformat. 5, 129 (2004).
Blanchette, M. et al. Genome-wide computational prediction of transcriptional regulatory modules reveals new insights into human gene expression. Genome Res. 16, 656–668 (2006). This study combines the identification of motif clusters with sequence constraint to produce a set of cis -regulatory module predictions that capture many known modules.
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nature Genet. 39, 311–318 (2007). By using a map of histone modifications and binding locations of key transcription factors, the authors have generated a model that predicts novel promoters and enhancers.
Gotea, V. et al. Homotypic clusters of transcription factor binding sites are a key component of human promoters and enhancers. Genome Res. 20, 565–577 (2010).
Donaldson, I. J. et al. Genome-wide identification of cis-regulatory sequences controlling blood and endothelial development. Hum. Mol. Genet. 14, 595–601 (2005).
Narlikar, L. et al. Genome-wide discovery of human heart enhancers. Genome Res. 20, 381–392 (2010).
Sinha, S. & He, X. MORPH: probabilistic alignment combined with hidden Markov models of cis-regulatory modules. PLoS Comput. Biol. 3, e216 (2007).
Majoros, W. H. & Ohler, U. Modeling the evolution of regulatory elements by simultaneous detection and alignment with phylogenetic pair HMMs. PLoS Comput. Biol. 6, e1001037 (2010).
Taylor, J. et al. ESPERR: learning strong and weak signals in genomic sequence alignments to identify functional elements. Genome Res. 16, 1596–1604 (2006).
Wang, H. et al. Experimental validation of predicted mammalian erythroid cis-regulatory modules. Genome Res. 16, 1480–1492 (2006). In reference 87, the authors describe a 'motif-blind' model for predicting CRMs based on patterns in multi-sequence alignments of known regulatory regions. Reference 88 shows that these the predictions were validated at a good rate.
Miller, W. et al. 28-way vertebrate alignment and conservation track in the UCSC Genome Browser. Genome Res. 17, 1797–1808 (2007).
Kantorovitz, M. R. et al. Motif-blind, genome-wide discovery of cis-regulatory modules in Drosophila and mouse. Dev. Cell 17, 568–579 (2009).
King, D. C. et al. Finding cis-regulatory elements using comparative genomics: some lessons from ENCODE data. Genome Res. 17, 775–786 (2007).
Schmidt, D. et al. Five-vertebrate ChIP–seq reveals the evolutionary dynamics of transcription factor binding. Science 328, 1036–1040 (2010).
Boffelli, D., Nobrega, M. A. & Rubin, E. M. Comparative genomics at the vertebrate extremes. Nature Rev. Genet. 5, 456–465 (2004).
Petrykowska, H., Vockley, C. & Elnitski, L. Detection and characterization of silencers and enhancer-blockers in the greater CFTR locus. Genome Res. 18, 1238–1246 (2008).
Goldberg, A. D., Allis, C. D. & Bernstein, E. Epigenetics: a landscape takes shape. Cell 128, 635–638 (2007).
Boyd, K. E. & Farnham, P. J. Myc versus USF: discrimination at the cad gene is determined by core promoter elements. Mol. Cell. Biol. 17, 2529–2537 (1997).
Ren, B. et al. Genome-wide location and function of DNA binding proteins. Science 290, 2306–2309 (2000).
Wold, B. & Myers, R. M. Sequence census methods for functional genomics. Nature Methods 5, 19–21 (2008).
Boyle, A. P. et al. High-resolution mapping and characterization of open chromatin across the genome. Cell 132, 311–322 (2008).
Hesselberth, J. R. et al. Global mapping of protein-DNA interactions in vivo by digital genomic footprinting. Nature Methods 6, 283–289 (2009).
ENCODE Project Consortium. A user's guide to the encyclopedia of DNA elements (ENCODE). PLoS Biol. 9, e1001046 (2011).
Gerstein, M. B. et al. Integrative analysis of the Caenorhabditis elegans genome by the modENCODE project. Science 330, 1775–1787 (2010).
Roy, S. et al. Identification of functional elements and regulatory circuits by Drosophila modENCODE. Science 330, 1787–1797 (2010).
Bernstein, B. E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nature Biotech. 28, 1045–1048 (2010).
Trinklein, N. D., Aldred, S. J., Saldanha, A. J. & Myers, R. M. Identification and functional analysis of human transcriptional promoters. Genome Res. 13, 308–312 (2003).
Landolin, J. M. et al. Sequence features that drive human promoter function and tissue specificity. Genome Res. 20, 890–898 (2010).
Roh, T. Y., Cuddapah, S. & Zhao, K. Active chromatin domains are defined by acetylation islands revealed by genome-wide mapping. Genes Dev. 19, 542–552 (2005).
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
Visel, A. et al. ChIP–seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009).
Blow, M. J. et al. ChIP–seq identification of weakly conserved heart enhancers. Nature Genet. 42, 806–810 (2010).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011). This study uses a probabilistic model to combine multiple chromatin marks maps into an integrated model to aid the interpretation of chromatin signatures.
Cheng, Y. et al. Transcriptional enhancement by GATA1-occupied DNA segments is strongly associated with evolutionary constraint on the binding site motif. Genome Res. 18, 1896–1905 (2008).
Wilson, N. K. et al. Combinatorial transcriptional control in blood stem/progenitor cells: genome-wide analysis of ten major transcriptional regulators. Cell Stem Cell 7, 532–544 (2010).
Tuan, D. Y., Solomon, W. B., London, I. M. & Lee, D. P. An erythroid-specific, developmental-stage-independent enhancer far upstream of the human “β-like globin” genes. Proc. Natl Acad. Sci. USA 86, 2554–2558 (1989).
West, A. G., Gaszner, M. & Felsenfeld, G. Insulators: many functions, many mechanisms. Genes Dev. 16, 271–288 (2002).
Boyle, A. P. et al. High-resolution genome-wide in vivo footprinting of diverse transcription factors in human cells. Genome Res. 21, 456–464 (2011). This paper demonstrates the use of DNase–seq for mapping open chromatin to predict regulatory regions and for footprinting individual transcription factor binding sites.
Rhee, H. S. & Pugh, B. F. Comprehensive genome-wide protein–DNA interactions detected at single-nucleotide resolution. Cell 147, 1408–1419 (2011).
Dostie, J. & Dekker, J. Mapping networks of physical interactions between genomic elements using 5C technology. Nature Protoc. 2, 988–1002 (2007).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Machanick, P. & Bailey, T. L. MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics 27, 1696–1697 (2011).
Kasowski, M. et al. Variation in transcription factor binding among humans. Science 328, 232–235 (2010).
Stranger, B. E. et al. Population genomics of human gene expression. Nature Genet. 39, 1217–1224 (2007).
Cheung, V. G. & Spielman, R. S. Genetics of human gene expression: mapping DNA variants that influence gene expression. Nature Rev. Genet. 10, 595–604 (2009).
Gaulton, K. J. et al. A map of open chromatin in human pancreatic islets. Nature Genet. 42, 255–259 (2010).
Kharchenko, P. V. et al. Comprehensive analysis of the chromatin landscape in Drosophila melanogaster. Nature 471, 480–485 (2011).
Hoffman, M. M. et al. Unsupervised pattern discovery in human chromatin structure through genomic segmentation. Nature Methods 9, 473–476 (2012).
Badis, G. et al. Diversity and complexity in DNA recognition by transcription factors. Science 324, 1720–1723 (2009).
Gilmour, D. S. & Lis, J. T. Detecting protein–DNA interactions in vivo: distribution of RNA polymerase on specific bacterial genes. Proc. Natl Acad. Sci. USA 81, 4275–4279 (1984).
Johnson, D. S., Mortazavi, A., Myers, R. M. & Wold, B. Genome-wide mapping of in vivo protein–DNA interactions. Science 316, 1497–1502 (2007).
Robertson, G. et al. Genome-wide profiles of STAT1 DNA association using chromatin immunoprecipitation and massively parallel sequencing. Nature Methods 4, 651–657 (2007). These two papers introduce ChIP–seq, which enables the binding locations of transcription factors to be mapped to DNA.
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
He, H. H. et al. Nucleosome dynamics define transcriptional enhancers. Nature Genet. 42, 343–347 (2010).
Staden, R. Methods for calculating the probabilities of finding patterns in sequences. Comput. Appl. Biosci. 5, 89–96 (1989).
Claverie, J. M. & Audic, S. The statistical significance of nucleotide position-weight matrix matches. Comput. Appl. Biosci. 12, 431–439 (1996).
Schones, D. E., Smith, A. D. & Zhang, M. Q. Statistical significance of cis-regulatory modules. BMC Bioinformat. 8, 19 (2007).
Odom, D. T. et al. Tissue-specific transcriptional regulation has diverged significantly between human and mouse. Nature Genet. 39, 730–732 (2007).
Cuellar-Partida, G. et al. Epigenetic priors for identifying active transcription factor binding sites. Bioinformatics 28, 56–62 (2012).
Fujiwara, T. et al. Discovering hematopoietic mechanisms through genome-wide analysis of GATA factor chromatin occupancy. Mol. Cell 36, 667–681 (2009).
Wu, W. et al. Dynamics of the epigenetic landscape during erythroid differentiation after GATA1 restoration. Genome Res. 21, 1659–1671 (2011).
Pollard, K. S., Hubisz, M. J., Rosenbloom, K. R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 20, 110–121 (2010).
Acknowledgements
Support is from US National Institutes of Health grants R01 DK065806, RC2 HG005573 and U54 HG004695 and funds from Emory University to J.T.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Related links
Related links
FURTHER INFORMATION
CCGB: Miller lab (for BLASTZ and LASTZ)
LAGAN Toolkit Website (for MLAGAN and CHAOS)
Nature Reviews Genetics Series on Regulatory elements
Regulatory sequence analysis tools
Siggia lab Rockefeller Univesrity software list (for Ahab)
Software for computational molecular biology (from Miller lab; for yama)
Glossary
- Purifying selection
-
The evolutionary process by which deleterious mutations are removed from a population or genome. Also referred to as negative selection or evolutionary constraint.
- Enhancers
-
DNA sequences that cause increased expression of their target gene (or genes).
- Epigenetic features
-
Molecules and chemical modifications that are associated with genomic DNA, including covalent modifications of DNA and histones, RNA transcribed from the DNA, occupancy of DNA by transcription factors and accessibility of DNA in chromatin to DNases.
- Transcription factor binding site
-
(TFBS). A short segment of DNA that is bound by a particular transcription factor in vivo.
- TFBS motif
-
A short string of DNA base pairs (often 6–10 bp long) constituting the sequence recognized by the DNA-binding domain of the transcription factor.
- Promoter
-
The DNA sequence that directs RNA polymerase to initiate transcription at the correct place.
- TFBS consensus
-
A string of DNA nucleotides describing the most frequently occurring short sequences in a collection of TFBSs, usually including ambiguous positions (for example, R refers to G or A nucleotides).
- Position-specific weight matrix
-
(PWM). A matrix providing the frequency at which each nucleotide is found at the positions of the TFBS consensus.
- Silencers
-
DNA sequences that cause reduced expression of their target gene (or genes).
- Insulators
-
DNA sequences that control the ability of an enhancer to regulate a promoter by an enhancer blocking activity or a domain barrier function or both.
- Position effects
-
The observation that the level of expression of some genes is affected by their position on chromosomes, with normal level of expression in one location but altered expression when translocated. For example, proximity to centromeres is associated with lowered expression for many genes.
- False positive
-
In a prediction experiment, a case in which the prediction is positive, but the true class is negative.
- Sensitivity
-
In a prediction experiment, the proportion of the true class that is predicted by the method: that is, (number of true positives)/(number of true positives + number of false negatives).
- Positive predictive value
-
In a prediction experiment, the proportion of positive predictions that are true positives.
- Position-specific scoring matrix
-
(PSSM). A matrix providing the log ratio of frequency at which each nucleotide is found at the positions of the TFBS consensus relative to a background model.
- TFBS motif instances
-
A match to a TFBS consensus or motif matrix within a longer DNA sequence (for example, a genome or chromosome).
- Specificity
-
In a prediction experiment, the proportion of the false class that is not predicted by the method: that is, (number of true negatives)/(number of true negatives + number of false positives).
- Logistic regression
-
A form of regression used when the output is binary. The predictor is a linear combination of the input variables transformed with the logistic function to form a probability. For classification, the coefficients are learned to maximize the (log) conditional likelihood of the training data.
- Homotypic clusters
-
Clusters of similar transcription factor binding sites that often bind the same transcription factor.
- Chromatin immunoprecipitation
-
(ChIP). A method for purifying the DNA segments that are in close contact with a transcription factor in living cells. After crosslinking DNA to native proteins in cells and preparing sheared chromatin, antibodies that specifically react with one transcription factor are used to isolate the DNA bound to that transcription factor.
- ChIP followed by high-throughput sequencing
-
(ChIP–seq). A technique for mapping the particular segments of DNA purified by chromatin immunoprecipitation (ChIP): it involves massively parallel short-read (second generation) sequencing and then aligning the reads to a reference genome. ChIP–seq is often highly accurate and has close to whole-genome coverage.
- Hidden Markov model
-
A statistical model in which internal states are not visible but the outputs of these states are, and the outputs can therefore be used to infer the internal states. This model can be used to determine biologically relevant states from chromatin immunoprecipitation followed by high-throughput sequencing (ChIP–seq) data sets.
- Morpholino oligonucleotides
-
Synthetic oligonucleotides in which the ribose portion of the nucleotide is replaced a morpholino compound; these are more stable than RNA and can be used to interfere with gene activity in transgenic zebrafish.
- ChIP–exo
-
An extension of ChIP–seq that includes exonuclease trimming after immunoprecipitation to increase the resolution of the mapped transcription-factor-bound sites.
Rights and permissions
About this article
Cite this article
Hardison, R., Taylor, J. Genomic approaches towards finding cis-regulatory modules in animals. Nat Rev Genet 13, 469–483 (2012). https://doi.org/10.1038/nrg3242
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3242
This article is cited by
-
Cell-type-specific prediction of 3D chromatin organization enables high-throughput in silico genetic screening
Nature Biotechnology (2023)
-
Structural and developmental dynamics of Matrix associated regions in Drosophila melanogaster genome
BMC Genomics (2022)
-
Detecting clusters of transcription factors based on a nonhomogeneous poisson process model
BMC Bioinformatics (2022)
-
MOCCA: a flexible suite for modelling DNA sequence motif occurrence combinatorics
BMC Bioinformatics (2021)
-
Comparative transcriptomic analysis highlights contrasting levels of resistance of Vitis vinifera and Vitis amurensis to Botrytis cinerea
Horticulture Research (2021)