Abstract
The ability to conditionally rewire pathways in human cells holds great therapeutic potential. Transcription activator-like effectors (TALEs) are a class of naturally occurring specific DNA binding proteins that can be used to introduce targeted genome modifications or control gene expression. Here we present TALE hybrids engineered to respond to endogenous signals and capable of controlling transgenes by applying a predetermined and tunable action at the single-cell level. Specifically, we first demonstrate that combinations of TALEs can be used to modulate the expression of stably integrated genes in kidney cells. We then introduce a general purpose two-hybrid approach that can be customized to regulate the function of any TALE either using effector molecules or a heterodimerization reaction. Finally, we demonstrate the successful interface of TALEs to specific endogenous signals, namely hypoxia signaling and microRNAs, essentially closing the loop between cellular information and chromosomal transgene expression.
Similar content being viewed by others
Introduction
Transcription activator-like effectors (TALEs) were first discovered in plant pathogenic bacteria Xanthomonas1,2. Most naturally occurring TALEs contain a central domain of tandem, 33–35 amino acid repeats, followed by a single truncated repeat of 20 amino acids ( Fig. 1a ). Each repeat is largely identical except for two variable amino acids at positions 12 and 13, the repeat variable di-residues (RVDs). Protein crystallography (PX) studies reveal that each TALE repeat contains two helices connected by a short RVD-containing loop. The protein forms a right-handed, superhelical structure with RVDs contacting the major groove of the DNA double helix. The 12th residue helps stabilize the RVD loop, while the 13th residue participates in the base-specific contact3,4,5. Further studies have shown that the four most common RVDs each preferentially bind to one of the four bases (HD to C, NI to A, NG to T, NN to G)6,7,8.
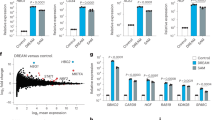
Design and conceptual elements of the TALE-hybrid approach.
(a) Schematic of general TALE design guidelines in this study. TALE hybrids were designed to control expression of target genes and their functions regulated by small molecules or endogenous signals. (b) Illustration of TALE-based rewiring of endogenous signals to chromosomal gene expression. (c) Schematic illustration of the stably integrated AmCyan transcript. The DNA binding sequences for three TALEs included in this study are shown.
The straightforward TALE-DNA binding specificity provides important new tools for genome engineering and targeting9,10. TALEs were fused with the catalytic domain of the FokI endonuclease to generate a new class of sequence-specific nucleases, the TAL effector nucleases (TALENs)11,12,13,14. TALENs, when used in pairs, can produce double-strand breaks between the target sequences and induce non-homologous end-joining and homologous recombination in endogenous target genes, such as a mutant form of the human β-globin (HBB) gene associated with sickle cell disease15. Secondly, TALE fusion proteins which contain transactivation domains were generated to induce the expression of specific genes and thus could potentially be used as therapeutic tools for hereditary diseases16. For example, TALEs which specifically target the human frataxin promoter were fused with VP64 transcription activator and the resulting fusion increased endogenous frataxin gene expression17. Finally, TALEs were fused with the KRAB transcriptional repression domain and these fusion TALE repressors were able to efficiently repress in transient transfections the synthetic fluorescent reporter gene which contains target sequences of TALEs18, as well as the transcription of endogenous human SOX2 gene19.
Venturing towards rewiring endogenous signals to chromosomal gene expression ( Fig. 1b ), we first performed a comprehensive characterization of TALE hybrids engineered for transgene activation and repression. Subsequently, based on a 2-hybrid approach, we engineered two different mechanisms to modulate any TALE function in cells. The first system is based on fusing a custom TALE protein to synthetic heterodimers that bind depending on the concentration of an externally delivered effector molecule and accordingly result to the recruitment of transcriptional components and the initiation of transcription of a target transgene. The second system is based on a fusion of a custom TALE protein to a sequence that forms a heterodimer with an endogenous transcription factor that translocates into the nucleus only under specific cellular conditions and again results to the initiation of the target transgene transcription. Finally, we successfully interfaced functional TALEs with endogenous microRNAs (miR-16 and miR-17) and a transcription factor (HIF-1α), essentially closing the loop between specific cellular signals and chromosomal gene expression.
Results
Characterization of TALE-based activation and inhibition
We first explored the transactivation activities of TALE fusion proteins. In order to use a well-controlled environment, we opted for the Flp-In system (Invitrogen) to generate a single-copy isogenic HEK293 stable cell line which contains an AmCyan fluorescent reporter gene under the control of a tetracycline responsive element (TRE) and minimum CMV promoter ( Fig. 1c ). We note that the particular stably integrated gene cassette contains the reverse tetracycline-controlled transactivator (rtTA) protein transcript under the control of a CMV promoter, as well as other regulatory elements not relevant to this work ( Supplementary Fig. 1 ). In the presence of doxycycline, rtTA binds the TRE element and drives the expression of AmCyan ( Fig. 2a ).
TAL effectors controlling the expression of the AmCyan transgene cassette.
(a) Induction of expression of AmCyan fluorescent proteins by TALETRE-VP16 activators. Different amounts of these two TALE fusion constructs were transiently transfected. 48 hours post transfection, cells were subjected to flow cytometry analysis. The fluorescence readings of wells which received control vector were used as the baseline and were subtracted from all other experimental samples. Bar graphs show expression levels of AmCyan as determined by flow cytometry and represent average and standard deviation from three replicates. TALETRE#3 and TALETRE#4 were designed to target the TRE sequence and fused with VP16 transactivation domain. Left, top: Both TALETRE-VP16 fusion proteins strongly induced the expression of AmCyan. Right, top: Fluorescence microscopy images of TALE-induced AmCyan expression in selected samples (I–IV). Bottom: 3-D illustration of overlaid flow cytometry histograms of AmCyan expression under the induction of TALETRE-VP16. I, control vector (250 ng); II, TALETRE#3-VP16 (250 ng); III, TALETRE#4-VP16 (250 ng); IV, TALETRE#3-VP16 (125 ng) + TALETRE#4-VP16 (125 ng). (b) Induction of expression of AmCyan fluorescent proteins by TALETRE-p65 activators. Left: Bar graphs representing expression levels of AmCyan as determined by flow cytometry. Left, inserts: Fluorescence microscopy images of selected samples (II, III). Right: Overlaid flow cytometry histograms of AmCyan expression between samples (II, III) and sample I (control). I, control vector (250 ng); II, TALETRE#3-p65 (250 ng); III, TALETRE#4-p65 (250 ng). (c) Suppression of expression of AmCyan fluorescent proteins by TALETRE-KRAB transcriptional repressors. The cells were induced by 10 ng TALETRE#4-VP16 and were co-transfected with different amounts of TALETRE#3-KRAB, TALETRE#4-KRAB, or TALECMV-KRAB plasmids. 72 hours post transfection the cells were subjected to the same analysis as above. The fluorescence readings of wells which received no TALETRE#4-VP16 induction were used as the baseline and were subtracted from all other experimental samples. Top: Fluorescence microscopy images of AmCyan expression in selected samples (I–VI). Middle: Bar graphs representing expression levels of AmCyan as determined by flow cytometry. Bottom: Overlaid flow cytometry histograms of AmCyan expression between samples (II–VI, color filled histograms) and sample I (positive control, black line histogram).
We then generated two TALEs, TALETRE#3 and TALETRE#4, which bind to 7 repetitive sequences within the TRE elements ( Fig. 1c and Supplementary Table 1 ). Both TALEs were fused with a VP16 transactivation domain and transiently transfected into the stable cells. After 48 hours both TALE fusion proteins strongly induced the expression of AmCyan, as quantified by microscopy and flow cytometry ( Fig. 2a ). The results hold for 72 hours measurements (data not shown). The transactivation activities of the TALE-VP16 proteins were significantly higher than doxycycline-induced rtTA. Notably, 10 ng of TALETRE-VP16 fusion protein ( Fig. 2a , TALETRE#3 and TALETRE#4 panels) resulted in stronger AmCyan expression than the saturation concentration of doxycycline (1 µg/ml) ( Fig. 2a ). To investigate possible synergistic effects between the two TALE fusion proteins, equal amounts of both constructs were co-transfected, but no obvious such effects were observed ( Fig. 2a ). We then sought to fuse our two TALEs with a weaker transactivation domain, the NF-kB p65 activation domain. The TALETRE#3-p65 and TALETRE#4-p65 fusions effectively induced the expression of AmCyan ( Fig. 2b ), as expected at lower levels (approximately 3-fold) compared to the VP16 fusions.
Next, we replaced the VP16 transactivation domain with a KRAB transcriptional repressor domain in the above TALE fusion proteins to determine their suppression effects. The expression of AmCyan was induced by 10 ng of TALETRE#4-VP16 ( Fig. 2c ) corresponding to the saturating doxycycline conditions. Different amounts of TALETRE#3-KRAB or TALETRE#4-KRAB were co-transfected and 72 hrs after transfection, both fusion proteins show strong suppression of the expression of AmCyan ( Fig. 2c ). 150 ng of either construct were able to abolish the expression of AmCyan. In fact, the TALETRE#4-KRAB at 150 ng suppressed AmCyan below its basal (due to TRE leakage) expression level ( Fig. 2c , IV).
We then generated a TALE that binds to the TATA box of the CMV promoter and fused it with KRAB domain (TALECMV-KRAB) ( Supplementary Table 1 ). The particular TALECMV has two binding sites within our stably integrated gene circuit: one within the CMV promoter ( Supplementary Figure 1 ) and the other one within the CMVmin portion of the TRE/CMVmin promoter ( Fig. 1 ). Compared to the aforementioned TALETRE-KRAB fusions, the suppression capacity of TALECMV-KRAB was significantly weaker. For instance, 50 ng of TALETRE#3-KRAB suppressed 84% of TALETRE#4-VP16-induced amCyan signal and 50 ng of TALETRE#4-KRAB suppressed 80%. In comparison, the same amount of TALECMV-KRAB only suppressed 19% of the original fluorescent protein signal ( Fig. 2c ). To rule out of the possibility that our TALECMV-KRAB fusion does not efficiently bind to its target sequence, we co-transfected this construct with 100 ng of CMV-mKate-PEST into plain HEK293 cells. Strong suppression of CMV-mKate-PEST was observed, as 100 ng of TALECMV-KRAB reduced its expression by 93% ( Supplementary Fig. 2 ). We note that compared to amCyan, mKate fluorescent protein used in this experiment was fused with a PEST domain, which increases its turnover rate and may contribute to its higher susceptibility to TALE’s suppression. Given that the TALETRE-KRAB and TALECMV-KRAB bind to different target sequences, we explored their possible synergistic effects by co-transfecting equal amount of each construct. These combinations, tested at different total DNA levels, did not result in a greater suppression compared to the individual constructs ( Fig. 2c ).
We also tested whether our TALE-KRAB fusion proteins can suppress the induction effect of doxycycline. The stable cells were first transfected with different amounts of three TALE-KRAB constructs and then induced by 0.3 µg/ml doxycycline after 24 hrs. All three TALE-KRAB fusions still significantly suppressed the expression of AmCyan, though none were able to fully abolish the doxycycline-induced expression of the transgene ( Supplementary Fig. 3 ). The difference between the TALETRE#4-VP16 and doxycycline-induction experiments may be partially attributed to the transient transfection properties. When two plasmids are co-transfected (TALETRE#4-VP16 and TALE-KRAB) it is reasonable to assume higher probability for delivery into the same population of cells. In contrast, when inducing with doxycycline and transfecting TALE-KRAB there will be a subpopulation of induced cells that do not receive the TALE-KRAB plasmid and thus cannot be suppressed by TALE, increasing the population mean AmCyan level. To probe this effect we transfected the cells with a constitutive YFP plasmid and induced the AmCyan using doxycycline; indeed, a significant portion of the AmCyan positive population failed to overlap with the YFP positive population (data not shown).
To conclude, the combination of TALETRE-KRAB and TALETRE#4-VP16 competing for overlapping or adjacent binding sites in the TRE element resulted in the most efficient repression of the transgene expression in the population of cells. We note that adjusting the ratio between the two proteins can essentially reverse the inhibitory action. When we co-transfected 150 ng of either TALETRE-KRAB constructs with different amount of TALETRE#4-VP16 into our stable cells, TALETRE#4-VP16 eventually counteracted the inhibitory effects of TALETRE-KRABs and induces the expression of AmCyan in a dose-dependent manner ( Supplementary Fig. 4 ).
TALE-based two-hybrid system
Regulating the function of TALE-fusions using small molecules was our next objective. We observed that both TALE-VP16 and TALE-KRAB fusion constructs consist of two functional domains: first, the DNA-binding domain and, second, the transactivation or the repressor domain, which could potentially be separated into two components in a two-hybrid system (for reviews of mammalian two-hybrids, see20,21). To test the feasibility of this TALE-based two-hybrid system, we fused the TALETRE#3 and TALETRE#4 DNA binding domain with the Rheo Receptor (New England Biolabs). The Rheo Activator contains a VP16 transactivation domain but by itself lacks the capability of inducing expression of target genes. Upon induction with GenoStat ligand (Millipore), the Rheo Receptor and Rheo Activator form a heterodimer, which brings the VP16 domain to the proximity of TRE element and thus induces the expression of AmCyan. We co-transfected the two plasmids into the stable cells and tested a range of different concentrations of GenoStat. 72 hours post transfection, the induction levels of AmCyan ( Fig. 3 ) show a clear correlation with the GenoStat concentration, indicating that GenoStat specifically induced the association of TALETRE-Rheo Receptor and Rheo Activator, which resulted in a functional transactivation complex.
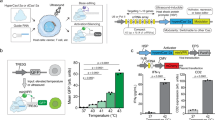
Induction of expression of AmCyan fluorescent proteins by TALE-based two-hybrid system.
(a) Schematic illustration of the TALE-based two-hybrid method. The TALETRE#3 and TALETRE#4 were fused with Rheo Receptor. 500 ng of these two TALE fusion plasmids were co-transfected into the cells with 500 ng of EF1-Rheo Activator. The cells were treated with different concentration of GenoStat ligand. 72 hours post transfection the cells were subjected to fluorescence microscopy and flow cytometry analysis. All experiments were performed in triplicates. (b) 3-D illustration of overlaid flow cytometry histograms of AmCyan in cell samples treated with different concentration of GenoStat. (front to back: 0, 4, 20, 100 and 500 nM of GenoStat). (c) Fluorescence microscopy images and bar graph representations of AmCyan expression in same samples. The AmCyan signals correlate with GenoStat concentrations.
Interface of TALEs with endogenous signals
After the successful TALE-based control of chromosomal transgene expression and modulation of the activity of the TALEs using small molecules, our final objective was to interface a functional TALE to endogenous signals and consequently close the loop between cellular information and the transgene expression. We first attempted to connect to an endogenous signaling pathway using a modification of the proposed TALE-based two-hybrid approach. We selected to interface a TALE with the hypoxia pathway, given its general importance to cell health, but the approach should generally apply to any cellular heterodimerization reaction.
The central transcription factor of the hypoxia signaling, HIF-1 (hypoxia-inducible factor-1) is composed of two subunits: HIF-1α and ARNT (aryl hydrocarbon receptor nuclear translocator). ARNT is constitutively expressed, whereas HIF-1α is targeted to proteasome degradation under normoxia. Under hypoxia or CoCl2 treatment, HIF-1α is stabilized and translocates to the nucleus, where it forms an active heterodimer with ARNT22.
We fused the amino acids 1–474 of human ARNT protein, which contain the HIF-1α-interacting domain bHLH-PAS but lack the transactivation domain, to our TALETRE#3 and TALETRE#4 DNA binding domains ( Fig. 4a ). The treatment of cells with CoCl2 (100 μM) significantly increased the expression of AmCyan fluorescent protein ( Fig. 4b ), indicating that the stabilized HIF-1α protein formed a functional heterodimer with the TALETRE-ARNT1-474 fusions. In comparison, only minimal level of AmCyan expression was observed when the negative control, TALETRE#4-Rheo Receptor, was transfected. We note that even without CoCl2 treatment ( Fig. 4b ), TALETRE-ARNT1-474 fusions induced an intermediate expression level of AmCyan. This result mostly likely arises from bHLH-PAS-dependent protein-protein cross-talk, as ARNT protein has been demonstrated to interact with other transactivators such as MOP1 and MOP223.
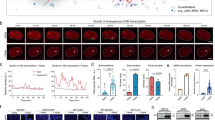
TALE interface with endogenous transcription factor and microRNA signals.
(a) The 1–454 amino acids of human ARNT protein fused to the TALETRE#3 and TALETRE#4 DNA binding domains reacting with HIF-1α under hypoxic conditions and inducing the transgene amCyan. (b) Induction of expression of AmCyan fluorescent proteins by TALETRE-ARNT1-454 fusions under treatment of CoCl2 (100 μM). The TALETRE#3 and TALETRE#4 were fused with amino acids 1–454 of human ARNT protein. 800 ng of these two TALE fusion plasmids were transfected into the cells with or without treatment of CoCl2 (100 μM). 72 hours post transfection the cells were subjected to flow cytometry analysis. All experiments were performed in triplicates. Overlaid flow cytometry histograms of AmCyan in cell samples with or without CoCl2 treatment. Incubation with 100 μM of CoCl2 significantly increased the expression of AmCyan fluorescent protein. Bar graphs represent AmCyan expression in same samples. TALETRE#4-Rheo Receptor was included as the negative control. (c) MiR-16 and miR-17 target sequences incorporated into 3′-UTR regions of TALETRE#3-VP16 and TALETRE#4-VP16 constructs. FF4 targets are used as the negative control. 10 ng of each construct were transiently transfected into cells. 72 hours post transfection cells were subjected to fluorescence microscopy and flow cytometry analysis. The fluorescence readings of wells which received control vector were used as the baseline and were subtracted from all other experimental samples. All experiments were performed in triplicates. (d) Overlaid flow cytometry histograms of AmCyan between samples with (color filled histograms) or without (black line histograms) miR-16 or -17 target sequences. (e) Fluorescence microscopy images and bar graphs showing relative mRNA level of TALE-VP16 and expression level of AmCyan signals. The induction capacity of TALETRE-VP16 was significantly lower when miR-16 or -17 targets were inserted. ** denotes p<0.01.
In addition to the hypoxia signaling, we also selected microRNAs given their critical role in cells24 and on cell fate25. To interface with endogenous microRNAs, we invoke a method of microRNA-mediated repression that involves miRISC (microRNA and RISC complex) and direct endonucleolytic mRNA cleavage in a mechanism that highly resembles RNAi. Although rare in mammalian cells, it is known to occur when perfect complementarity between the microRNA target site and the miRISC group exists26,27.
We first focused on two of the most abundantly expressed miRNAs in HEK293 cells, miR-16 and miR-17 and incorporated 4 copies of their reverse complementary sequences into the 3’-UTR regions of TALETRE#3-VP16 and TALETRE#4-VP16 ( Fig. 4c ). A negative control construct was also generated by inserting 4 copies of the reverse complementary sequences of the artificial microRNA miR-FF4 (described in28,29). Both miR-16 and miR-17 reduced the mRNA levels of TALETRE-VP16 by more than 95% and effectively suppressed TALETRE-VP16’s induction of AmCyan signals ( Fig. 4d ), while no down-regulation could be observed in the TALETRE-VP16 constructs which contain miR-FF4 targets ( Fig. 4e ). Note that the induction capacity of these constructs can be partially reduced by co-transfection of miR-FF4 ( Supplementary Fig. 5 ). In addition to the most abundant miRNAs (miR-16 and miR-17), we further tested the suppression effects of miR-10b, which expression in HEK293 cells is at intermediate level and miR-146a, which is absent in HEK293 cells. Four copies of the reverse complementary sequences of these two miRNAs were inserted into our TALETRE-VP16 constructs. Compared to miR-16 and miR-17, as expected, the suppression effects of miR-10b on TALETRE-VP16 were mild, resulting in as expected intermediate AmCyan activation ( Supplementary Fig. 6 ).
Discussion
The ability of signaling networks to detect, process and react specifically to various signals is a key property of living cells and the implementation of systems30,31,32 that reliably rewire such endogenous pathways can be a future therapeutic TALE-based application. The results presented here point to new generations of TALE hybrids and synthetic circuits engineered to detect and monitor endogenous signals with the capability of interfacing with biological pathways to apply predetermined and controllable action at the single-cell level.
We show that competitive action of TALEs can be used to effectively control chromosomal gene expression. Furthermore, we introduced a novel 2-hybrid system that can be used to regulate the activity of any TALE. Finally, as a proof of principle, we demonstrated the successful interface of TALEs with hypoxia signaling and endogenous microRNA, essentially closing the loop by activating the stably integrated transgene cassette. In the future, TALE-based synthetic networks will be able to interface with the cellular environment to filter, amplify and reliably transduce signals applying custom and fine-tuned control.
Methods
Recombinant DNA constructs
All TALE constructs were prepared using the Golden Gate TALEN and TAL effector kit (Addgene, catalog number: 1000000016) developed by Cermak et al16. The TAL effector target sequences and their according RVD sequences were designed using the online tool TAL Effector Targeter (http://boglabx.plp.iastate.edu/TALENT/). The target sequences are between 12–18 bp and preceded by a T (Supplementary Table S1). For the detailed cloning plan, see Supplementary Experimental Procedures, DNA constructs section and Supplementary Table S2.
Cell culture and transient transfection
A HEK293 stable cell line that harbors the Tetracycline Responsive Element (TRE) AmCyan transcript was generated using Flp-In System (Invitrogen, catalog number: K6010-01) according to the manufacturer's instructions. The cells were maintained at 37°C, 100% humidity and 5% CO2. The cells were grown in Dulbecco's modified Eagle's medium (DMEM, Invitrogen, catalog number: 11965-1181) supplemented with 10% Fetal Bovine Serum (FBS, Invitrogen, catalog number: 26140), 0.1 mM MEM non-essential amino acids (Invitrogen, catalog number: 11140-050), 0.045 units/mL of Penicillin and 0.045 units/mL of Streptomycin (Penicillin-Streptomycin liquid, Invitrogen, catalog number: 15140) and 50 µg Hygromycin B (Invitrogen, catalog number: 10687-010). To pass the cells, the adherent culture was first washed with PBS (Dulbecco's Phosphate Buffered Saline, Mediatech, catalog number: 21-030-CM), then trypsinized with Trypsin-EDTA (0.25% Trypsin with EDTAX4Na, Invitrogen, catalog number: 25200) and finally diluted in a fresh medium upon reaching 50–90% confluence. To maintain plain HEK293 cells, the procedures were essentially the same, except that no Hygromycin B was included in the growth medium.
For transient transfections, ~300 thousand cells in 1 mL of complete medium were plated into each well of 12-well culture treated plastic plates (Griener Bio-One, catalog number: 665180) and grown for 16-20 hours. For Lipofectamine LTX transfection, up to 1 µg of the plasmid was added to 200 µL of DMEM and 2 µL Lipofectamine LTX (Invitrogen, catalog number: 94756). Transfection solutions were mixed and incubated at room temperature for 30 minutes. The transfection mixture was then applied to the cells and mixed with the medium by gentle shaking. When applicable, doxycycline (Clontech, catalog number: 631311) was added three hours after transfection.
Fluorescence microscopy
All microscopy was performed 48–72 hours post transfection. The live cells were grown on 12-well plates (Greiner Bio-One) in the complete medium. Cells were imaged using the Olympus IX81 microscope and a Precision Control environmental chamber. The images were captured using a Hamamatsu ORCA-03 Cooled monochrome digital camera. The filter sets (Chroma) are as follows: ET436/20x (excitation) and ET480/40 m (emission) for AmCyan, ET560/40x (excitation) and ET630/75 m (emission) for mKate, ET500/20x (excitation) and ET535/30 m (emission) for YFP (Yellow Fluorescent Protein). Data collection and processing was performed in software package Slidebook 5.0. All images within a given experimental set were collected with the same exposure times and underwent identical processing.
Flow cytometry
48–72 hours post transfection cells from each well of the 12-well plates were trypsinized with 0.1 mL 0.25% Trypsin-EDTA at 37°C for 3 mins. Trypsin-EDTA was then neutralized by adding 0.9 mL of complete medium. The cell suspension was centrifuged at 1000 rpm for 5 mins and after removal of supernatants, the cell pellets were resuspended in 0.5 mL PBS buffer. The cells were analyzed on a BD LSRFortessa flow analyzer. AmCyan was measured with a 445-nm laser and a 515/20 band-pass filter, mKate with a 561-nm laser, 610 emission filter and 610/20 band-pass filter and YFP with a 488-nm laser, a 535 emission filter and 545/35 band-pass filter.
For experiments performed in TRE_AmCyan HEK293 cells, 100,000 events were collected. A FSC (forward scatter)/SSC (side scatter) gate was generated using a un-transfected negative sample and applied to all cell samples. The mean values of AmCyan reporter fluorescence were then collected and processed by FlowJo. The average of the means of AmCyan from three control samples which were transfected with empty plasmids (EF1-FF3X3-FF4X3, see Supplementary Experimental Procedures, DNA constructs section) were set as baseline values and were subtracted from all other experimental samples. All experiments were performed in triplicates.
For experiments performed in plain HEK293 cells, 50,000 events were collected. A FSC (forward scatter)/SSC (side scatter) gate was first generated using a un-transfected negative sample and applied to all cell samples. The cells were further gated to select the YFP+ populations. The mean values of mKate and YFP were collected and processed by FlowJo. The ratios of mKate/YFP were then calculated.
Quantitative reverse transcription-PCR
48 hours post transfection, total RNA was extracted from TRE_AmCyan HEK293 cells using an RNeasy Mini kit (Qiagen, catalog number: 74104) following the manufacturer's protocol. First-strand synthesis was performed using QuantiTect Reverse Transcription kit (Qiagen, catalog number: 205311). Quantitative PCR was performed using KAPA SYBR FAST Universal qPCR kit (KAPA Biosystems, catalog number: KK4601). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) sequences were used for normalization. The forward primer for GAPDH was 5′-AATCCCATCACCATCTTCCA-3′ and the reverse primer for GAPDH was 5′- TGGACTCCACGACGTACTCA-3′. The forward primer for TALE was 5′- CTCCACTTAGACGGCGAGGA-3′ and the reverse primer for TALE was 5′- GAAGTCGGCCGTATCCAGAG-3′. The thermal cycling conditions were 3 min at 95°C followed by 40 cycles of denaturation for 15 s at 95°C and annealing for 30 s at 60°C. Normalized data were used to compare relative levels of TALE-VP16 transcripts which contained different miRNA targets using ΔΔCt analysis.
Statistical analysis
The values of AmCyan reporter fluorescence are reported as mean with standard deviation. The significance values between sample groups were calculated by the Student's ‘t’-test and P-value less than 0.05 were taken as significance in all the experiments.
References
Sugio, A., Yang, B., Zhu, T. & White, F. F. Two type III effector genes of Xanthomonas oryzae pv. oryzae control the induction of the host genes OsTFIIAγ1 and OsTFX1 during bacterial blight of rice. Proc Natl Acad Sci U S A 104, 10720 (2007).
Boch, J. & Bonas, U. Xanthomonas AvrBs3 family-type III effectors: Discovery and function. Annu Rev Phytopathol 48, 419–436 (2010).
Mak, A. N. S., Bradley, P., Cernadas, R. A., Bogdanove, A. J. & Stoddard, B. L. The crystal structure of TAL effector PthXo1 bound to its DNA target. Science 335, 716–719 (2012).
Deng, D., Yan, C., Pan, X. et al. Structural basis for sequence-specific recognition of DNA by TAL effectors. Science 335, 720–723 (2012).
Bradley, P. Structural modeling of TAL effector–DNA interactions. Protein Science 21(4), 471–4 (2012).
Moscou, M. J. & Bogdanove, A. J. A simple cipher governs DNA recognition by TAL effectors. Science 326, 1501–1501 (2009).
Boch, J., Scholze, H., Schornack, S. et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326, 1509 (2009).
Streubel, J., Blücher, C., Landgraf, A. & Boch, J. TAL effector RVD specificities and efficiencies. Nat Biotechnol 30, 593–595 (2012).
Briggs, A. W., Rios, X., Chari, R. et al. Iterative capped assembly: rapid and scalable synthesis of repeat-module DNA such as TAL effectors from individual monomers. Nucleic Acids Res 40(15), e117 (2012).
Doyle, E. L., Booher, N. J., Standage, D. S. et al. TAL Effector-Nucleotide Targeter (TALE-NT) 2.0: tools for TAL effector design and target prediction. Nucleic Acids Res 40, W117–W122 (2012).
Li, T., Huang, S., Jiang, W. Z. et al. TAL nucleases (TALNs): hybrid proteins composed of TAL effectors and FokI DNA-cleavage domain. Nucleic Acids Res 39, 359–372 (2011).
Kim, H., Um, E., Cho, S. R., Jung, C., Kim, H. & Kim, J. S. Surrogate reporters for enrichment of cells with nuclease-induced mutations. Nat Methods 8, 941–943 (2011).
Christian, M., Cermak, T., Doyle, E. L. et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186, 757–761 (2010).
Kleinstiver, B. P., Wolfs, J. M., Kolaczyk, T., Roberts, A. K., Hu, S. X. & Edgell, D. R. Monomeric site-specific nucleases for genome editing. Proc Natl Acad Sci U S A 109, 8061 (2012).
Sun, N., Liang, J., Abil, Z. & Zhao, H. Optimized TAL effector nucleases (TALENs) for use in treatment of sickle cell disease. Mol.BioSyst. 8(4), 1255–63 (2012).
Cermak, T., Doyle, E. L., Christian, M. et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res 39, E82 (2011).
Tremblay, J. P., Chapdelaine, P., Coulombe, Z. & Rousseau, J. TALE proteins induced the expression of the frataxin gene. Hum Gene Ther 23(8), 883–90 (2012).
Garg, A., Lohmueller, J. J., Silver, P. A. & Armel, T. Z. Engineering synthetic TAL effectors with orthogonal target sites. Nucleic Acids Res 10.1093/nar/gks404 (2012).
Cong, L., Zhou, R., Kuo, Y., Cunniff, M. & Zhang, F. Comprehensive interrogation of natural TALE DNA-binding modules and transcriptional repressor domains. Nat Commun 3, 968 (2012).
Lievens, S., Lemmens, I. & Tavernier, J. Mammalian two-hybrids come of age. Trends Biochem Sci 34, 579–588 (2009).
Lee, J. W. & Lee, S. K. Mammalian two-hybrid assay for detecting protein-protein interactions in vivo. Methods Mol Biol 439, 327 (2008).
Yuan, Y., Hilliard, G., Ferguson, T. & Millhorn, D. E. Cobalt inhibits the interaction between hypoxia-inducible factor-α and von Hippel-Lindau protein by direct binding to hypoxia-inducible factor-α. J Biol Chem 278, 15911 (2003).
Long, W. P., Chen, X. & Perdew, G. H. Protein kinase C modulates aryl hydrocarbon receptor nuclear translocator protein-mediated transactivation potential in a dimer context. J Biol Chem 274, 12391 (1999).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Wijnhoven, B., Michael, M. & Watson, D. MicroRNAs and cancer. Br J Surg 94, 23–30 (2007).
Stegmeier, F., Hu, G., Rickles, R. J., Hannon, G. J. & Elledge, S. J. A lentiviral microRNA-based system for single-copy polymerase II-regulated RNA interference in mammalian cells. Proc Natl Acad Sci U S A 102, 13212 (2005).
Xie, Z., Wroblewska, L., Prochazka, L., Weiss, R. & Benenson, Y. Multi-input RNAi-based logic circuit for identification of specific cancer cells. Science 333, 1307 (2011).
Bleris, L., Xie, Z., Glass, D., Adadey, A., Sontag, E. & Benenson, Y. Synthetic incoherent feedforward circuits show adaptation to the amount of their genetic template. Mol Syst Biol 7 (2011).
Rinaudo, K., Bleris, L., Maddamsetti, R., Subramanian, S., Weiss, R. & Benenson, Y. A universal RNAi-based logic evaluator that operates in mammalian cells. Nat Biotechnol 25, 795–801 (2007).
Holtz, W. J. & Keasling, J. D. Engineering static and dynamic control of synthetic pathways. Cell 140, 19–23 (2010).
Ruder, W. C., Lu, T. & Collins, J. J. Synthetic biology moving into the clinic. Science 333, 1248–1252 (2011).
Benenson, Y. Biomolecular computing systems: principles, progress and potential. Nat Rev Genet 13, 455–468 (2012).
Acknowledgements
This work was funded by the US National Institutes of Health (NIH) grant 1R21GM098984, the US National Science Foundation (NSF) grant CBNET-1105524 and the University of Texas at Dallas. We would like to thank A. Chandrahas and K. Ehrhardt for comments and technical support and M. Zhang and X. Xie for discussions.
Author information
Authors and Affiliations
Contributions
L.B., Y.L., R.M., M.G. conceived the project and designed the experiments. Y.L., R.M., M.G. performed the experiments. Y.L. analyzed the data. Y.L., L.B, M.G., R.M. prepared the manuscript. L.B. supervised the project.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplement Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Li, Y., Moore, R., Guinn, M. et al. Transcription activator-like effector hybrids for conditional control and rewiring of chromosomal transgene expression. Sci Rep 2, 897 (2012). https://doi.org/10.1038/srep00897
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00897
This article is cited by
-
Design principles for synthetic control systems to engineer plants
Plant Cell Reports (2023)
-
Orthogonal control of gene expression in plants using synthetic promoters and CRISPR-based transcription factors
Plant Methods (2022)
-
An engineered tale-transcription factor rescues transcription of factor VII impaired by promoter mutations and enhances its endogenous expression in hepatocytes
Scientific Reports (2016)
-
A transcription activator–like effector (TALE) induction system mediated by proteolysis
Nature Chemical Biology (2016)
-
Modular construction of mammalian gene circuits using TALE transcriptional repressors
Nature Chemical Biology (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.