Abstract
The trans-Himalayan region of India, although have xeric features, still supports a unique assemblage of biodiversity, including some of the charismatic and endemic species. In the present study, we studied blue sheep (Pseudois nayaur) across the distribution range in the Western trans Himalayas of India and found about 18,775 km2 area suitable for blue sheep. The explicit Bayesian based spatial and non-spatial population structure analysis assigned blue sheep into two genetic populations, i.e., Ladakh and Lahaul-Spiti. We found relatively high genetic divergence in blue sheep which is also supported by the low current flow in Circuitscape model. With the multiple evidences, we explain landscape resistance facilitated by the landscape heterogeneity, and large patches of unsuitable habitats forced population divergence and poor functional connectivity. We found that blue sheep population has been demographically stable in the past, but showed a slight decline within the last few decades. This study is the first range-wide attempt to exhibit landscape features in shaping the spatial distribution, genetic structure and demography patterns of blue sheep in Western Himalayas, and will be of use in the conservation and management planning of blue sheep.
Similar content being viewed by others
Introduction
Heterogeneity in large landscape often delineate species boundary and primarily forms the foundation of genetic differentiation in free ranging wildlife. In this context, human-induced habitat loss and climate change are known to be the key threats to biodiversity conservation worldwide1,2. Despite the profound impact of such changes on global biodiversity, there is a scarcity of direct evidence on how species respond to such changes, while landscape changes (also due to climate change) are more evident and visible than the species adaptive response. Therefore, to facilitate long term persistence of a species, functional connectivity need to ensured so as to enable gene flow and sharing of alleles throughout the landscape3,4. In free ranging terrestrial wildlife, gene flow is often limited by the degree of landscape permeability and connectivity between different habitats patches5. Assessing the factors impacting habitat connectivity and dispersal are essential measure to interpret evolutionary consequences and to forbid local and regional extirpations. Landscape genetics is a trans-disciplinary approach, particularly useful in elucidating the cumulative influence of ecological, landscape and other anthropogenic factors on gene flow and natural selection6,7,8. In this context, maternally inherit mtDNA and nuclear microsatellite markers are used in addressing phylogeography, evolutionary history and current population genetics parameters with respect to the landscape features and habitat connectivity6,9. With the ensemble approach of coupling environmental, ecological and other anthropogenic variables with the genetic information of the natural populations may depict the current spatial genetic structure and provide empirical information of management interventions e.g., assigning large geographical barriers10, finding unique taxonomic units11 and identifying habitats for population6,7,8.
Blue sheep (Pseudois nayaur) is one of the widely distributed medium-sized alpine ungulate distributed from the Qilian Mountains in the north to The Himalayas in the south. Globally, it is distributed in China, Bhutan, India, Myanmar, Nepal, Pakistan and Tajikistan12,13. In India, it is distributed in Ladakh, Himachal Pradesh, Uttarakhand, Sikkim and Arunachal Pradesh between the elevation ranges from 2500 to 5500 m asl14,15,16. They typically prefer plateau, alpine barren and inter-valley grassland16 and play a key ecological role in the maintaining the mountain ecosystems by imparting a crucial prey base to the large carnivores and determine their density and distribution in that region14,17,18. Blue sheep are listed as least concern (LC) under the IUCN red list and listed as Schedule I under the Wildlife (Protection) Act, 1972 of India16,19. However, no firm information is available of the population size of blue sheep19. Previous studies reported climate change and urbanization are the major threats to blue sheep across distribution range and interestingly, blue sheep show relatively less seasonal altitudinal migration with respect to changing seasons 20,21. Moreover, blue sheep usually found in large herd and avoid steep slope. While, the mountain ranges of the Western Trans Himalayas of India i.e. the Ladakh and Zanskar range appear to separate the mountainous landscape by challenging topography such as steep cliffs and canyons that could act as barriers to blue sheep movement. In addition, blue sheep also compete with other mountain ungulates and livestock for limited resources and often impacted with their high densities22,23,24. In the present study, we adopted a landscape genetics approach to testify the functionality of biological corridor to support the movement and gene flow of blue sheep in Western trans-Himalaya.
Results
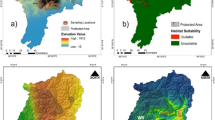
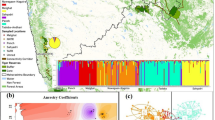
Of about the 106,014 km2 of the study area, only 18,775 km2 (<20%) was found suitable for the blue sheep that lies primarily in Leh region of Ladakh (LA) and Spiti region of Lahaul-spiti (LS) (Fig. 1b). The estimated AUC was 0.89 ± 0.081 indicating all the selected variables significantly contributed to building the model. Jackknife estimates that most contributing variables was Precipitation Seasonality (16%) followed by Mean Temperature of Driest Quarter (13.2%) and Euclidean distance from grasslands (11.9%) while Topography influence index and Euclidean distance from barren land showed least contribution 1.3% and 1.1% respectively (Table S4, Fig. S1).
(a) Geographical map of the study area displaying sampling location of blue sheep (Pseudois nayaur) was created using ArcGIS 10.6 (http://www.esri.com) from Indian Trans-Himalayas. (b) Map of predicted habitat suitability for blue sheep in Ladakh and Lahaul-Spiti. The species distribution model of blue sheep was generated using Maximum Entropy species distribution model (MaxEnt v.3.3 http://biodiversityinformatics.amnh.org/open_source/maxent/) and ArcGIS 10.6 (http://www.esri.com) was used to visualize the map. (c) Genetic divergence plot of blue sheep in the studied landscape using the Genetic Landscape GIS Toolbox (https://www.sciencebase.gov/catalog/item/57e97d09e4b09082500c9210) in ArcGIS 10.6 (http://www.esri.com). High divergence at the peripheral population. (d) Map showing predicted corridors in the study landscape (High-brown and Low-blue). The connectivity map was generated with Circuitscape software v. 4.0 (https://circuitscape.org/downloads/) and plotted onto the map using ArcGIS 10.6 (http://www.esri.com).
On analysing 137 sequences of cytochrome b gene of blue sheep, we obtained seven haplotypes (Fig. 2a) with 0.331 ± 0.095 and π 0.012 ± 0.00393, respectively (Table 1). No statistical significance for Tajima’s D and Fu and Li’s F and D test were detected (> 0.01). We did not find blue sheep in Kargil and Lahaul region as none of the faecal samples was identified as blue sheep. The studied population showed a multimodal pattern of mismatch distribution (Fig. 2c). However, the BSP indicated a demographically stable population with slight decline during the last few decades (Fig. 2b).
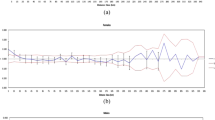
(a) Haplotype network generated in NETWORK software (v 10.2) using cytb gene of blue sheep in Ladakh and Lahaul-Spiti regions. The generated haplotype was then visualized on the map to understand the sharing of the haplotype in the studied area using ArcGIS 10.6 (ESRI, Redlands, CA). (b) Demographic history of (Pseudois nayaur) population estimated using Bayesian skyline plot in BEAST software v 2.6.3 https://www.beast2.org/ (Bayesian skyline plot showing a stable population in the recent past). The solid line is the median estimates of Ne τ (Ne = effective population size; τ = generation time), and the grey lines around median estimates is the 95% highest posterior density (HPD) estimate of the historic effective population size. The timing of events was estimated assuming a substitution rate of 2.0 × 10–6 substitutions/site/year (Joshi et al. 2020; Johns et al.1998). (c) The multimodal pattern of mismatch distribution was generated using DnaSP (v 6.12.03 http://www.ub.edu/dnasp/), indicating the blue sheep population under demographic equilibrium.
We generated 137 consensus genotypes with nine loci with considerably good amplification success (≥ 80%) (Table 2). The ADO and FA in all the loci were non-significant. Majority loci (8/9) were highly informative (PIC value > 0.5) and all the loci deviated from HWE except BM1824 (P < 0.05). None of the pairs of loci showed significant linkage disequilibrium (P < 0.05).
Unique individuals were identified using a panel of six microsatellite loci and the cumulative probability of identity assuming all individuals were siblings (PID sibs) was 2.1 × 10–3 (2.1 match in 1000 genotypes). In total, 118 unique individuals were identified out of 137 consensus genotypes generated. Genetic diversity indices of blue sheep population showed a mean Na and Ne of 9.89 ± 1.15 (LA) to 10 ± 0.71 (LS) and 4.37 ± 0.58 (LA) to 3.80 ± 0.50 (LS), respectively. The mean Ho was 0.44 ± 0.06 in LA and 0.35 ± 0.06 in Spiti, while the mean He was 0.72 ± 0.05 in LA and 0.70 ± 0.03 in Spiti (Table S6). A broad variation inbreeding coefficient value among loci ranging from 0.099 (BM1824 locus) to 0.722 (INRA35 locus) indicates slight inbreeding between the populations (Table 2).
STRUCTURE analysis identified two genetic clusters (ΔK = 2). The genetic partitioning showed the presence of geo-spatial structure in the population i.e. 56 individuals from LA assigned to Cluster I while 47 individuals from LS were assigned to Cluster II (Fig. 4a). Only 14 (#8 individuals from LA and #6 individuals from LS) showed admixed ancestry. GENELAND inferred the presence of four genetic clusters based on the highest average posterior probability where majority of individuals were assigned into two clusters supporting the geo-spatial clusters i.e., LA and LS. The LA population showed a further break-down into four smaller groups, indicating the presence of meta-populations of blue sheep in LA (Fig. 4c). The multi variation DAPC also show similar clustering pattern to STRUCTURE and identified two major clusters (Fig. 4b).
BAYESASS showed low migrations i.e. ≤ 0.1% from either side from LA to LS and vice versa. The sPCA revealed a cline in allele frequencies from LA to LS (Fig. 4d). While the mantel test detected a gradual pattern of IBD across the study (r = 0.291; P > 0.01) (Fig. 3). The genetic divergence model also corroborated with the IBD and showed high genetic divergence between the LA and LS region and very high divergence in the Nubra and Changtang valley of LA which could be due to the peripheral population and edge effect (Fig. 1c).
Scatterplot showing the result of Mantel test for presence of IBD (Isolation by distance) between significance of geographical distance on the genetic distance (r = 0.291) in the blue sheep population. Colour represent the relative density of point higher densities are represented by warmer colour while show the correlation between the distance matrices. IBD was performed in R.4.0 software with Adegenet version 1.3.4 package.
The Circuitscape model showed cumulative currents confining within LA and LS and no high intensity current flows between regions indicating the lack of physical connectivity between the regions (Fig. 1d).
Discussion
Previous studies on blue sheep in India mainly focused on the ecological aspect and no study was conducted on the population genetics of blue sheep15,22,25,26. For accurate assessment of the spatial genetic patterns of blue sheep inhabiting Ladakh and Lahaul-spiti, we identified the landscape features that influenced genetic structure. Here, we included habitat and corridor modelling in addition to genetic data to understand the impact of landscape on the genetic diversity of blue sheep. In general, blue sheep avoids forested habitats, prefers alpine pasture and frequently found near the escape covers16,17. We also observed the similar patterns in the present study, as habitat suitability is positively influenced by the grassland area. Since, there is low human habitation in the studied landscape, the anthropogenic variables had insignificant effects on the blue sheep habitat, rather the habitat suitability was mainly influenced by the bioclimatic variable and land use (precipitation, temperature and grassland). Blue sheep prefers low precipitation and rely on the mean Temperature of Driest Quarter. Blue sheep are usually found in large herds and have relatively less dispersal which is also supported by our study showing low migration rate between the two regions i.e., Ladakh and Lahaul-Spiti (Home range: − 3.7 km2 male and females 2.9 km2 Cui27; Zhang et al.28). The present study suggested that low connectivity between LA and LS in addition to complex topography and high expanses among habitat patches.
We found comparable genetic diversity of blue sheep with previous studies29 and other caprinae species e.g., wild goats30; Hangul31; and Iberian ibex32. The neutrality tests were non-significant, indicating a stable population size. Mismatch distribution curve also revealed a stable population of blue sheep with no bottleneck in the past. The BSP analysis showed a stable population in the past with experiencing a slight demographic decline in the recent past. Thus, all the demographic data corroborate with each other. Mitochondrial data suggested gene flow in the past as one haplotype (hap_1) was distributed in all the region and Zanskar haplotype (hap_7) (Fig. 2a) is closer to the Spiti haplotype rather than the LA as geographically it is closer to Spiti.
Population genetic structure analysis based on microsatellite marker revealed all the individuals grouped into two distinct clusters with limited gene flow. One of the factors contributing to the limited gene flow between LA and LS area is likely to be limited physical corridor and patchiness in suitable area. The previous studies also suggested influence of geographic distance and landscape barrier on the genetic distance in blue sheep population33. Likewise, we also obtained a significant IBD and sPCA results suggested that individuals sampled on the periphery have higher genetic divergence than the central populations (Figs. 3, 4D). In addition to the landscape barrier, mating strategies and breeding system of blue sheep may also influence the genetic diversity between the landscapes34. The Zanskar range that divides the two regions could be resistance that impedes connectivity however, this needs a further detailed investigation. In other studies, much attention has been focused on the rare and specialist species while widely distributed species are often neglected and considered safe without recognizing their peripheral population31.
(a) Bayesian clustering patterns of blue sheep population at K2; Assignment at population level (cluster 1: Ladakh, cluster 2: Lahaul-spiti); A (i): Mean L (K), A (ii) The ad hoc quantity (delta K) over 20 runs for each K value. The structure analysis was performed using STRUCTURE v 2.3.4. (b) Non-Bayesian pattern of blue sheep with DAPC, (c) Estimated cluster membership shows spatial distribution of the four inferred genetic clusters across the study landscape using 118 blue sheep individuals (K4) based on clustering Bayesian model GENELAND (GENELAND v 4.0.3 4 package of R https://www.sciencebase.gov/catalog/item/57e97d09e4b09082500c9210). (d) Interpolation using a globally weighted regression of component 1 score for sPCA and contours are component score representing similarity across the landscape. DAPC and sPCA generated with Adegenet version 1.3.4 package of R (https://CRAN.R-project.org/package=adegenet).
Despite being important biodiversity hotspot of unique and endemic fauna, not much attention has been given in the context of genetic monitoring of large mammals in these landscapes. Further, there are six protected areas in these landscapes where local live in harmony with the indigenous wildlife. Both, Changthang (CWS) and Karakoram (KWS) sanctuaries of LA where blue sheep majorly reported have no definite boundary till now. Moreover, these regions had encountered drastic changes in the last few years including increased presence of defense forces and large influx of tourists leading to unplanned construction that have disturbed the harmonious association and affected the native wildlife. A few earlier studies have identified that anthropogenic changes often restrict gene flow among populations and exhibited the utility of landscape genetics in conservation and management planning in ungulates35. Likewise, with the blend of genetic data and landscape modeling, we provide pragmatic potential connecting corridors supporting the movement and gene flow in blue sheep in the studied landscape. Although, we tested the present landscape genetic model on blue sheep, we believe the validated connecting corridors could be useful in multi species conservation and management.
Materials and methods
Study area
We prioritized sampling of blue sheep in the reported distribution of the Lahul and Spiti (LS) districts of Himachal Pradesh and Leh and Kargil districts of Ladakh union territory (LA-UT), falling under the Western Trans Himalayan region covering a total area of 1,06,014 km2 (LA 86,983 km2 and LS 19,032 km2) (Fig. 1). The Lahaul valley possesses mix condition of Himalaya and trans-Himalaya while Spiti and Ladakh regions harbour low faunal diversity and the trans-Himalayan features. The study landscape is consisting of six protected areas i.e. Pin Valley National Park (PNP), Kibber Wildlife Sanctuary KBS, and Chandra Tal Wildlife Sanctuary (CWS) located in Spiti valley of Himachal Pradesh and Changthang Cold Desert Sanctuary (CWS), Karakoram Wildlife Sanctuary (KWS), Hemis National Park (HNP) located in LA UT.
Species occurrence data
Field surveys were performed from 2018 to 2021 in the studied landscape. We opted well established contour transect in the studied landscape and in total, 350 occurrence points of blue sheep were recorded (275 pellet samples, 75 sign survey). In addition to the primary occurrence records, secondary data was extracted from the Global Biodiversity Information Facility (GBIF) database (www.gbif.org). For distribution modelling, combination of primary as well as secondary data were used. To reduce over fitting of the model spatially independent locations were used after rectifying the location points based on the home range of blue sheep27,28.
Ethics declarations
This study was based on samples collected non-invasively where no animal was captured or harmed. Thus, ethical clearance was not needed.
Sampling and DNA extraction
We collected 275 non-invasive (pellet) samples, i.e., 118 pellet samples from LS region and 157 samples from LA region possibly belonging to blue sheep (Fig. 1a). The collected samples were sun dried and stored in sterile containers with ¼ filled with silica crystals. Genomic DNA from pellets was extracted using QIAamp Fast DNA Stool Mini Kit (QIAGEN, Germany) following manufacturer's instructions.
Species identification
For species verification, we used partial fragments of the mitochondrial cytochrome b36. We set-up the PCR in a 10μL containing 10X PCR buffer, 2 mM MgCl2, 2.5 mM dNTPs mix, 0.1 μM primers each, and 20-30 ng template. The amplification conditions were as follows: 94 °C for 5 min followed by 35 cycles at 94 °C for 30 s, 53 °C for 45 s, and 72 °C for 45 s, and a final extension at 72 °C for 10 min on a Veriti Thermocycler (Applied Biosystems, USA). The sanger sequencing was performed following Big-Dye Terminator Cycle Sequencing Kit version 3.1 (Thermo Scientific, USA) after exo-sap clean up on an ABI 3730 Genetic analyzer (Applied Biosystems, USA).
Microsatellite genotyping
All the genetically identified samples of blue sheep were processed for genotyping. We selected 22 microsatellites (15 loci of Bovidae;30 and 7 loci of cervidae;37 (Table S1). Total, 16 microsatellites were successfully amplified (Table S2). PCRs were set up following Gao et al.33 (Table 2). The fluorescence-based genotyping was performed on Genetic Analyzer (Applied Biosystem, Foster City, USA) and allele scoring was performed using Gene Mapper (version 4.1, Applied Biosystems, USA).
Data analysis
Habitat suitability model
To identify suitable habitat for blue sheep, we used a Maximum Entropy (MaxEnt) species distribution model (version 3.338). We started with a set of 30 habitat variables including present bioclimatic variable, topographic variables, physiological variables and anthropogenic variables. The 19 bioclimatic variables were obtained from the WorldClim data base at 30 arc second scale (https://www.worldclim.org/)39 (Table S3). Topographic variables such as elevation, slope, aspect, ruggedness and topography wetness index were generated from digital elevation model using ArcGIS 10.6 software (ESRI, REDLANDS, CA). MODIS data were downloaded for land use/land cover (LULC) from USGS Earth Explorer (https://earthexplorer.usgs.gov) and ArcGIS 10.6 (ESRI, REDLANDS, CA) was used for calculating the Euclidian distance. Human influence index was downloaded from Socioeconomic Data and Application (SEDAC). The variables were resampled at 1 km resolution and converted to ASCII format in ArcGIS 10.6 (ESRI, REDLANDS, CA). Multi- collinearity among the variables were tested in the R platform using ENM tool40 and the variable with Person Correlation Coefficient (r) more than 0.8 were dropped from the analysis. Finally, eighteen spatially independent predictors (Table S4) were used for mapping the suitable habitat of blue sheep and best fit model was determined in the R package ENMeval41 on the basis of lowest Akaike information criterion (AICc) value42. Further, for the model evaluation, we used 70% of species presence points as training data and 30% as testing data. The modelling was assessed based on Receiver Operating Characteristics (ROC) Area under the curve (AUC) with the threshold value higher than 0.8 < AUC < 1 which provide good prediction model.
Sequencing data
Generated sequences were cleaned with Sequencher 5.4.6 (Gene Code Corp) and validated through the BLAST tool of NCBI/GenBank. The identified sequences were aligned by multiple sequence alignment using CLUSTAL W in MEGA version X43 and trimmed to generate similar length of sequences for further genetic analysis.
Using the program DnaSP version 6.12.0344, Neutrality tests (Tajima’s D, Fu and Li’s F and D statistics) and genetic diversity indices i.e. no. of haplotypes (H), nucleotide diversity (π), haplotype diversity (Hd), average number of nucleotide differences (K) and mismatch distribution test for demographic expansion, equilibrium or bottleneck were computed. The NETWORK45 software was used to build a median-joining network. Subsequently, the obtained haplotype was then plotted on the map to visualize the haplotype sharing across the studied areas, using ArcMap 10.6 (ESRI, REDLANDS, CA). Bayesian Skyline Plot (BSP) was constructed to determine the history of population over times using BEAST software version 2 6.346 by Bayesian Skyline method47. The HKY model and the Markov Chains Monte Carlo (MCMC) were run for 108 steps that yielded effective sample sizes (ESS) under a strict molecular clock and a stepwise skyline model. The molecular evolution was calibrated assuming a substitution rate of 2.0E−7 substitutions/ site/year following11,48 and the results were visualized using Tracer version 1.749.
Microsatellite data
For individual identification, we selected a panel of six loci on the basis of their least or no genotyping error, relatively short amplicon size, high amplification success and a high discriminating power to avoid the overestimation of individuals. The locus wise and cumulative probability of identity for unrelated individuals (PID) and siblings (PID sibs) were calculated in GenAlEx version 6.5150 and the number of unique individuals were estimated in GeneCap version 1.2.251 and the PIC value for each locus was estimated using CERVUS version 3.052. While the nine markers were used to calculate the genetic diversity indices that included numbers of observed (Na) and effective alleles (Ne), observed heterozygosity (HO), expected heterozygosity (HE), and Wright's inbreeding coefficient at each locus utilizing GENAIEX version 6.550. We tested the deviation of any loci from Hardy–Weinberg equilibrium (HWE) using the program FSTAT version 2.9.453 and null allele frequencies were calculated using CERVUS version 3.052. Linkage disequilibrium (LD) was tested using GENEPOP version 4.7 following 10,000 dememorizations, 100 batches and 10,000 iterations per batch54.
To determine whether discrete population structure was present in the blue sheep population, we used two Bayesian clustering (STRUCTURE and GENELAND) and one Non-Bayesian clustering (DAPC) method. First Bayesian clustering approach was performed in STRUCTURE version 2.3.455 with a burn-in period of 50,000 and 500,000 (MCMC) repetitions and 20 independent replicates at each ‘K’. The appropriate K value was determined by calculating ad hoc quantity (ΔK)56. Individuals were assigned to the inferred clusters using a threshold proportion of membership (q) i.e., q ≥ 0.80, otherwise an individual was determined as admixed if the q value was less than 0.8057,58,59. Secondly to spatially explicit the genetic discontinuities between the populations, we performed Bayesian clustering analysis in GENELAND version 4.0.360 through an extension of R package. In the analysis, along with multi-locus genotype, spatial coordinates were also taken into account. We conducted 20 independent runs for each K ranging from 1 to 10, with 1,000,000 iterations and 1000 thinning with other parameters were kept as default values (maximum rate of the Poisson process, 100; uncertainty on spatial coordinates, 0; maximum number of nuclei, 300; null allele model, FALSE). The runs showing the highest average logarithm of the posterior probability were selected and post-processing was conducted using 100 × 100 pixels in the spatial domain with a burn in period of 200. Finally non-Bayesian clustering methods i.e., DAPC were run to assign the possible clusters in Adegenet version 1.3.4 package of R61. To evaluate the migration rate between the LA and LS population, we run BAYESASS version 1.362 at 9 × 106 MCMC iterations, 106 were discarded as burn-in and sampling frequency was done at every 2000 intervals. Runs were carried out at migration rate prior of 0.05, while other parameters were kept at default.
Corridor connectivity using landscape genetics
To explore the potential landscape barriers and corridors effecting the gene flow and population divergence of blue sheep, we first estimated genetic divergence in GenAlEx version 6.5150. Then, we mapped the pattern of genetic distances (independent of any a priori model of landscape resistance) using the Genetic Landscape GIS Toolbox63 in ArcGIS 10.6 (ESRI, REDLANDS, CA). We used the Single Species module, to calculate surface of the pair-wise population genetic differentiation by generating a network. Midpoint of each edge of the network becomes associated with genetic diversity value. In the package, both spatial location points as well as relative genetic divergence values were used. In addition, presence of Isolation by distance (IBD) across the study landscape was also generated with Mantel test. The Mental test and sPCA was performed in R.4.0 software with Adegenet version 1.3.4 package61. Lastly, we built connectivity model based on circuit theory in the Circuitscape software version 4.064 to test the connectivity between the habitat patches to understand the dispersal and gene flow following Dalui et al.6. For conductance modelling resistance raster was constructed with ensemble of genetic divergence and habitat suitable model using the ArcGIS 10.6 software (ESRI, REDLANDS, CA) extension Gnarly landscape utilities package Resistance and Habitat Calculator65.
Data availability
The sampling location of blue sheep used in this study is available on DRYAR (https://doi.org/10.5061/dryad.t1g1jwt7z). The novel seven haplotypes generated are available on NCBI and their unique accession numbers are mentioned under supplementary table (Table S5).
References
Aryal, A., Brunton, D., Ji, W. & Raubenheimer, D. Blue sheep in the Annapurna conservation area, Nepal: Habitat use, population biomass and their contribution to the carrying capacity of snow leopards. Integr. Zool. 9, 34–45 (2014).
Liu, Z., Wang, X., Teng, L., Cui, D. & Li, X. Estimating seasonal density of blue sheep (Pseudois nayaur) in the Helan Mountain region using distance sampling methods. Ecol. Res. 23, 393–400 (2008).
Keyghobadi, N. The genetic implications of habitat fragmentation for animals. Can. J. Zool. 85, 1049–1064 (2007).
Radespiel, U. & Bruford, M. W. Fragmentation genetics of rainforest animals: Insights from recent studies. Conserv. Genet. 15, 245–260 (2014).
Benton, T. G. & Bowler, D. E. Linking dispersal to spatial dynamics. Dispersal Ecol. Evol. 251, 265 (2012).
Dalui, S. et al. Fine-scale landscape genetics unveiling contemporary asymmetric movement of red panda (Ailurus fulgens) in Kangchenjunga landscape India. Sci. Rep. 10, 15446 (2020).
Holderegger, R. & Wagner, H. H. Landscape genetics. Bioscience 58, 199–207 (2008).
Manel, S., Schwartz, M. K., Luikart, G. & Taberlet, P. Landscape genetics: Combining landscape ecology and population genetics. Trends Ecol. Evol. 18, 189–197 (2003).
Arif, I. A. et al. DNA marker technology for wildlife conservation. Saudi J. Biol. Sci. 18, 219–225 (2011).
Ghosh, A. et al. The Sela macaque (Macaca selai) is a distinct phylogenetic species that evolved from the Arunachal macaque following allopatric speciation. Mol. Phylogenet. Evol. 174, (2022).
Joshi, D. B. et al. Genetic evidence for allopatric speciation of the Siberian ibex Capra sibirica in India. Endanger. Species Res. 42, 1–5 (2020).
Liu, Z., Wang, X., Teng, L. & Cao, L. Food habits of blue sheep, Pseudois nayaur in the Helan Mountains China. Folia Zool. 56, 13–22 (2007).
Tan, S. et al. Molecular evidence for the subspecific differentiation of blue sheep (Pseudois nayaur) and polyphyletic origin of dwarf blue sheep (Pseudois schaeferi). Genetica 140, 159–167 (2012).
Harris, J. Wildlife Conservation in China: Preserving the Habitat of China’s Wild West. (Routledge, 2014). https://doi.org/10.4324/9781315698168.
Namgail, T., Fox, J. L. & Bhatnagar, Y. V. Habitat segregation between sympatric Tibetan argali Ovis ammon hodgsoni and blue sheep Pseudois nayaur in the Indian Trans-Himalaya. J. Zool. 262, 57–63 (2004).
Schaller G.B. Wildlife of the Tibetan Steppe,. Univ. Chicago Press. 373, (1998).
Oli, M. K. Seasonal patterns in habitat use of blue sheep Pseudois nayaur (Artiodactyla, Bovidae) in Nepal. Mammalia 60, 187–193 (1996).
Chundawat, R. S. & Rawat, G. Food habits of snow leopard in Ladakh, India, Proceedings of the 7th International Snow Leopard Symposium, International Snow Leopard Trust, Seattle, Washington, USA. 127–132 (1994).
Zhang, M. et al. Population dynamics of blue sheep Pseudois nayaur in Ningxia Helan Mountain National Nature Reserve China. Folia Zool. 61, 121–128 (2012).
Aryal, A. et al. Predicting the distributions of predator (snow leopard) and prey (blue sheep) under climate change in the Himalaya. Ecol. Evol. 6, 4065–4075 (2016).
Cui, X. & Graf, H. F. Recent land cover changes on the Tibetan Plateau: A review. Clim. Change 94, 47–61 (2009).
Bhattacharya, A., Chatterjee, N., Rawat, G. S. & Habib, B. Blue sheep resource selection in alpine grasslands of a western himalayan landscape—A point process approach. Zool. Stud. 59, 1–15 (2020).
Mishra, C., Van Wieren, S. E., Ketner, P., Heitkönig, I. M. A. & Prins, H. H. T. International Snow Leopard Trust (India Program), Nature Conservation Foundation. J. Appl. Ecol. 41 (2004).
Suryawanshi, K. R. Overwintering strategies and demographic response of bharal (Pseudois nayaur) to livestock grazing and removal, in Kibber Wildlife Sanctuary (2008).
Bhardwaj, M., Uniyal, V. P. & Sanyal, A. K. Estimating relative abundance and habitat use of Himalayan blue sheep Pseudois nayaur in Gangotri national park, western Himalaya India. Galemys 22, 545–560 (2010).
Namgail, T. Winter habitat partitioning between asiatic ibex and blue sheep in ladakh, northern india. J. Mt. Ecol. 8, 7–13 (2006).
Cui, D. Home range, activity rhythm and foraging strategy of bharal (Pseudois nayaur) in Helan Mountain, China. A Diss. Philos. Dr. Ecol. East China Norm. Univ. Shanghai, China.(in Chinese (2007).
Zhang, M., Wang, X. & Ding, Y. Flight responses of blue sheep in Ningxia Helan mountain national nature reserve. Folia Zool. 62, 185–192 (2013).
Yang, M. et al. Development and characterization of nine polymorphic microsatellite markers for the blue sheep (Pseudois nayaur). Conserv. Genet. Resour. 7, (2015).
Maudet, C. et al. Microsatellite DNA and recent statistical methods in wildlife conservation management: Applications in Alpine ibex [Capra ibex (ibex)]. Mol. Ecol. 11, 421–436 (2002).
Thakur, M., Schättin, E. W. & McShea, W. J. Globally common, locally rare: Revisiting disregarded genetic diversity for conservation planning of widespread species. Biodivers. Conserv. 27, 3031–3035 (2018).
Angelone-Alasaad, S., Biebach, I., Pérez, J. M., Soriguer, R. C. & Granados, J. E. Molecular analyses reveal unexpected genetic structure in Iberian ibex populations. PLoS One 12, e0170827 (2017).
Gao, H. et al. Taxonomic status of Chinese blue sheep (Pseudois nayaur): New evidence of a distinct subspecies. Integr. Zool. 15, 202–212 (2020).
Wang, X. M., Cao, L. R., Liu, Z. S. & Fang, S. G. Mitochondrial DNA variation and matrilineal structure in blue sheep populations of Helan Mountain China. Can. J. Zool. 84, 1431–1439 (2006).
Epps, C. W., Wehausen, J. D., Bleich, V. C., Torres, S. G. & Brashares, J. S. Optimizing dispersal and corridor models using landscape genetics. J. Appl. Ecol. 44, 714–724 (2007).
Gupta, S. K., Kumar, A. & Hussain, S. A. Novel primers for sequencing of the complete mitochondrial cytochrome b gene of ungulates using non-invasive and degraded biological samples. Conserv. Genet. Resour. 6, 499–501 (2014).
Moore, S. S., Barendse, W., Berger, K. T., Armitage, S. M. & Hetzel, D. J. S. Bovine and ovine DNA microsatellites from the EMBL and GENBANK databases. Anim. Genet. 23, 463–467 (2009).
Phillips, S. J. & Dudík, M. Modeling of species distributions with Maxent: New extensions and a comprehensive evaluation. Ecography (Cop.) 31, 161–175 (2008).
Fick, S. E. & Hijmans, R. J. WorldClim 2: New 1-km spatial resolution climate surfaces for global land areas. Int. J. Climatol. 37, 4302–4315 (2017).
Warren, D. L. et al. ENMTools 1.0: An R package for comparative ecological biogeography. Ecography (Cop.). 44, 504–511 (2021).
Muscarella, R. et al. ENM eval: An R package for conducting spatially independent evaluations and estimating optimal model complexity for Maxent ecological niche models. Methods Ecol. Evol. 5, 1198–1205 (2014).
Kass, J. M. et al. ENMeval 2.0: Redesigned for customizable and reproducible modeling of species’ niches and distributions. Methods Ecol. Evol. 12, 1602–1608 (2021).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Librado, P. & Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Bandelt, H.-J., Forster, P. & Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48 (1999).
Bouckaert, R. et al. BEAST 2: A software platform for bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537 (2014).
Drummond, A. J. Bayesian coalescent inference of past population dynamics from molecular sequences. Mol. Biol. Evol. 22, 1185–1192 (2005).
Johns, G. C. & Avise, J. C. A Comparative summary of genetic distances in the vertebrates from the mitochondrial cytochome b gene. Mol. Biol. Evol. 15, (1998).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in bayesian phylogenetics using tracer 1.7. Syst. Biol. 67, 901–904 (2018).
Peakall, R. & Smouse, P. E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research: An update. Bioinformatics 28, 2537–2539 (2012).
Wilberg, M. J. & Dreher, B. P. genecap: A program for analysis of multilocus genotype data for non-invasive sampling and capture-recapture population estimation. Mol. Ecol. Notes 4, 783–785 (2004).
Kalinowski, S. T., Taper, M. L. & Marshall, T. C. Revising how the computer program cervus accommodates genotyping error increases success in paternity assignment. Mol. Ecol. 16, 1099–1106 (2007).
Goudet, J. FSTAT (version 2.9.4), a program (for Windows 95 and above) to estimate and test population genetics parameters. Dep. Ecol. Evol. Lausanne Univ. Switz. 53, (2003).
Raymond, M. & Rousset, F. An exact test for population differentiation. Evolution (N. Y). 49, 1280 (1995).
Pritchard, J. K., Wen, W. & Falush, D. Documentation for the structure software, version 2 (University of Chicago, Chicago, 2003).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Lecis, R. et al. Bayesian analyses of admixture in wild and domestic cats (Felis silvestris) using linked microsatellite loci. Mol. Ecol. 15, 119–131 (2005).
Takur, M. et al. Loss of genetic diversity and inbreeding in Kashmir red deer (Cervus elaphus hanglu) of Dachigam National Park, Jammu & Kashmir India. BMC Res. Notes 6, 326 (2013).
Pierpaoli, M. et al. Genetic distinction of wildcat ( Felis silvestris ) populations in Europe, and hybridization with domestic cats in Hungary. Mol. Ecol. 12, 2585–2598 (2003).
Guillot, G., Santos, F. & Estoup, A. Analysing georeferenced population genetics data with Geneland: A new algorithm to deal with null alleles and a friendly graphical user interface. Bioinformatics 24, 1406–1407 (2008).
Jombart, T. adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405 (2008).
Wilson, G. A. & Rannala, B. Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163, 1177–1191 (2003).
Vandergast, A. G., Perry, W. M., Lugo, R. V. & Hathaway, S. A. Genetic landscapes GIS Toolbox: Tools to map patterns of genetic divergence and diversity. Mol. Ecol. Resour. 11, 158–161 (2011).
Mcrae, B. H. & Shah, V. B. Circuitscape user guide (Univ, 2011).
McRae, B., Shirk, A. & Platt, J. Gnarly Landscape Utilities: Resistance and Habitat Calculator User Guide. The Nature Conservancy, Fort Collins, CO. Available at: https://circuitscape.org/gnarly-landscape-utilities/ (2013).
Funding
This study was supported by the National Mission for Himalayan Studies, Ministry of Environment, Forest and Climate Change (MoEF&CC) New Delhi (Grant No. NMHS/2017–18/LG09/02/476).
Author information
Authors and Affiliations
Contributions
S.Do. and G.J. undertook field surveys and collected samples. S.Do., S.K.S., B.D.J. and S.Da. participated in analyzing the field collected data. S.Do., S.K.S., generated STR data and undertook genetic analysis in the mentorship of M.T. M.T. and L.K.S. conceptualized the idea and S.Do. wrote the primary draft of the manuscript. M.T., L.K.S. and V.K.S. edited and revised the MS. S.Do., M.T., and L.K.S. finalized the MS. K.C. supervised the overall activities and provided all the logistic support and administrative approval.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dolker, S., Jabin, G., Singh, S.K. et al. Landscape genetics identified conservation priority areas for blue sheep (Pseudois nayaur) in the Indian Trans-Himalayan Region. Sci Rep 13, 18152 (2023). https://doi.org/10.1038/s41598-023-44823-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-44823-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.