Abstract
TDP-43 is a major pathological protein in sporadic and familial amyotrophic lateral sclerosis (ALS) and mediates mRNA fate. TDP-43 dysfunction leads to causes progressive degeneration of motor neurons, the details of which remain elusive. Elucidation of the molecular mechanisms of RNA binding could enhance our understanding of this devastating disease. We observed the involvement of the glycine-rich (GR) region of TDP-43 in the initial recognition and binding of G-quadruplex (G4)-RNA in conjunction with its RNA recognition motifs (RRM). We performed a molecular dissection of these intramolecular RNA-binding modules in this study. We confirmed that the ALS-linked mutations in the GR region lead to alteration in the G4 structure. In contrast, amino acid substitutions in the GR region alter the protein structure but do not void the interaction with G4-RNA. Based on these observations, we concluded that the structural distortion of G4 caused by these mutations interferes with RRM recruitment and leads to TDP-43 dysfunction. This intramolecular organization between RRM and GR regions modulates the overall G4-binding properties.
Similar content being viewed by others
Amyotrophic lateral sclerosis (ALS) is a progressive neurodegenerative disorder characterized by the selective death of motor neurons in the motor cortex, brainstem, and spinal cord1,2. Approximately 10% of the cases involve genetic mutations, whereas the remaining cases are considered sporadic due to unknown causes3. Currently, more than 50 causative or potentially related genes have been identified, of which the majority encode RNAs and RNA-binding proteins (RBPs)4,5. Therefore, identifying and analyzing the state of RNA molecules common to all is essential. Indeed, without exception, all these RNAs and RBPs are associated with G-quadruplexes (G4) and are components of ribonucleoprotein (RNP) granules assembled through liquid–liquid phase separation (LLPS)6 (Table S1). This is surprising because there are not many RBPs that specifically recognize G47,8. Since these gene mutations are thought to be involved in the onset of ALS prominent, the research on the structure and function of these RNAs and proteins is essential for understanding ALS pathogenesis9.
TDP-43 (43 kDa TAR DNA-binding protein, encoded by TARDBP) is an ALS-linked RBP4. The well-studied TDP-43 is a major disease-related protein involved in the pathogenesis of ALS, leading to form abnormal intraneuronal inclusions detected in more than 90% of patients10. Mutations in the TDP-43 gene have also been identified in familial and sporadic ALS patients3,11. Previously, we identified that TDP-43 binds to G4-containing mRNAs and transports them to distal neurites for local translation12. G4 is a higher-order DNA/RNA structure consisting of two or more guanine tetrads originating from four guanine bases assembled in a square planar arrangement via a Hoogsteen hydrogen bond scheme13. G4-binding RBPs may play essential roles in the regulation of mRNA transport and function, and the dysregulation of RBPs is feared to cause neurological disorders14,15. However, no detectable destabilization was observed for mutant TDP-43 proteins16,17,18, thus dysfunction of TDP-43 might be the cause of the disease onset.
Herein, we performed the molecular dissection of TDP-43. It contains two RNA recognition motifs (RRM) and a C-terminal glycine-rich (GR) region and forms a homodimer under physiological cellular conditions19. Most of the disease-associated amino acid substitutions are located within the uncharacterized GR region3. With the use of TDP-43 with ALS-linked mutations in RRM or GR region, we found a reduction of G4-mRNA transport into distal neurites for local translation, suggesting the participation of both regions in binding and function12,20. We then speculated the involvement of multiple modules in the recognition of G4-RNA. We confirmed that the RRM and GR regions bind in concert to parallel-stranded G4 conformations21.
To clarify the roles of each RRM and GR region in the recognition of G4-RNA, we investigated the influence of ALS-linked TDP-43 mutations on the G4-RNA binding activity using these RNA-binding module segments. The mutations in GR segment alone induced the conformation alteration of G4-RNA while it showed the effects on G4-RNA binding only in the simultaneous presence of the RRM and GR regions. We proposed the participation of GR region in the recognition and binding of G4-RNA, thereby inducing the recruitment of RRM.
Results
ALS-linked mutations in the G4-RNA binding
The recognition properties of TDP-43 have been analyzed in vitro using segment containing two RRMs22,23,24,25. However, the RRM of TDP-43 is not the only module that participates in RNA recognition23,25. Therefore, we purified a full-length untagged TDP-43 natural dimer and performed a more accurate RNA binding analysis12,20,21. Using PSD-95 (postsynaptic density protein 95) and CaMKIIα (calcium/calmodulin-dependent protein kinase II alpha subunit) (Table S2), two dendritic G4-mRNAs that bind TDP-43 in vivo26,27, we determined the effects of 10 ALS-linked amino acid substitution mutations20. All the mutant TDP-43 proteins showed reduced interaction with G4-RNA using SPR (surface plasmon resonance)12,20. In this study, we used these G4-RNAs and TDP-43 proteins that had the G287S or M337V mutations and the lowest G4-RNA binding properties (Fig. 1A).
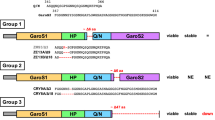
The ALS-linked mutations affect the interaction between human TDP-43 and G4-RNA. (A) The structural features of human TDP-43. The positions of two patient derived mutations are indicated. The dissociation rates of two types of G4 measured by SPR in previous studies are shown on the right20. (B) The purified untagged full-length dimeric proteins of wild-type and mutant proteins were used for this study. One microgram each of wild-type and mutant proteins were separated by 10% SDS-PAGE and detected by Coomassie Brilliant Blue staining (for original images, see Fig. S1A). (C) Far-UV CD spectrum of 0.2 μM of TDP-43 wild-type and two mutant proteins, TDP-43G287S and TDP-43M337V. The x-axis represents wavelength, and the y-axis represents the five scans’ average molar CD (Δε). The percentages of specific secondary structure motifs calculated using the BeStSel algorithm are shown below (D) The agarose gel mobility shift assay. 0.5 pmol of fluorescently labeled G4 probes, PSD-95 and CaMKIIα, were mixed with the indicated amounts of TDP-43 wild-type and mutant proteins and electrophoresed under non-denaturing conditions. Each over-shifted band was quantified, and the standard error (± SEM) obtained after three independent experiments (Exp1, Exp2, and Exp3) is shown in the graph below. The statistical significance was determined by a two-tailed Student’s t-test *P < 0.05, **P < 0.01, ***P < 0.001. When electrophoresed under inappropriate pH conditions, most bound signals are detected at below as smear signals, indicating that these dissociated during migration (see Fig. S1B).
First, to confirm the effects of ALS-linked mutations on the overall structure, far-UV circular dichroism (CD) spectroscopy was used (Fig. 1B,C). Secondary structure compositions of TDP-43 wild-type and mutant proteins were estimated using the BeStSel (Beta Structure Selection) algorithm (https://bestsel.elte.hu/index.php)28. TDP-43G287S has a more helical content, whereas TDP-43M337V showed a decrease and an increase in β-turn. We, therefore, hypothesized that the G287S and M337V mutations induced changes in the secondary and tertiary structures.
Next, to compare the interactions with G4-RNA under the unified experimental conditions of this study, an electrophoretic mobility shift assay was performed using a non-denaturing agarose gel (Fig. 1D). The binding of G4-RNA to specific interacting proteins with multiple binding modules often does not migrate normally when using a non-denaturing acrylamide gel6. As a result, this attempt was effective. By adding 2.5, 5, and 10 times the amount of protein to G4-RNA, we confirmed a dose-dependent over-shift in the G4-RNA mobility. The two mutant proteins showed a reduction in shifting bands, consistent with the SPR and secondary structure prediction data. The smeared bands of intermediate mobility might be dissociated RNAs during migration because the maximum signal is detected at this location when run at inappropriate pH condition for binding (Fig. S1B). This, combined with the recently reported results, we speculated that ALS-derived amino acid mutations alter the G4 associating structure of the GR region, resulting in the decreased direct binding affinity between G4-RNA and the GR region.
ALS-linked mutations in GR segment do not affect direct binding to G4-RNA
We confirmed the binding affinity to G4 by a segment of the GR region alone fused to GST (glutathione S-transferase)20. To confirm the effects of the mutations, we examined the possible influence of GST-fused GR mutant proteins on the G4-RNA immobilizing sensor chip using SPR (Fig. 2A). In the case of GST-GR fusion, however, no significant difference was observed in the interaction with G4-RNA between the wild-type GST-GR and mutant GST-GRs. To confirm this contradictory result between the full-length TDP-43 and the GR segment, these interactions were further analyzed by gel shift assay. No shifting bands were detectable on agarose gels, suggesting that additional binding modules were required for the GR region to form a stable complex (Fig. S1C). It was possible that the GR region alone cannot withstand even slight structural changes in gels forced to operate under low-salt conditions29,30,31. Therefore, we reinforced the association by UV cross-linking and confirmed the complex by SDS-PAGE21. The cross-linking and gel shift assay supported the SPR results, and two mutations did not affect the interaction of GST-GR with G4-RNAs (Fig. 2B). We repeated the experiment with different binding conditions such as buffer and temperature; however, the results were consistently reproducible.
The ALS-mutations in the GR segment do not affect the binding to G4-RNA. (A) The structural features of GST fusion GR wild-type and mutant proteins and SPR sensorgrams of the interaction between G4-RNA and GST fusion proteins. Terminal biotinylated poly dT16 was bound to the streptavidin-coated sensor chip through streptavidin–biotin complex, to which poly-dA16 tailed G4-RNA (PSD-95 dA16 and CaMKIIα dA16, 20 nM) was immobilized by hybridization. The test protein (50 nM) was added to measure interaction with G4-RNAs. (B) UV cross-linking for detection of G4-RNA and binding protein complexes. The reaction mixtures were irradiated at 254 nm at 200 mJ/cm2 and analyzed by denaturing gel mobility shift assay. Shifting bands were enhanced in both fluorescent probes (Table S2) with increased GST-GR proteins (1, 2 or, 4 pmol each). Several minor bands are seen that may have been cleaved during electrophoresis, but only the major band was used for quantification (for original images, see Fig. S2). The gel was stained with Coomassie Brilliant Blue and the blocking agent BSA signals were used to correct the data as a loading control. The experiments were performed thrice, and the y-axis values represent the mean ± SEM at 4 pmol. The statistical significance was not confirmed by a two-tailed Student’s t-test (n.s.).
These unexpected results suggested that the mutations in the GR region do not affect GR-G4-RNA binding but influence other molecular characteristics of GR such as the modulation of G4-RNA structures and/or intramolecular signal transmission to the RRM for modulation of its RNA-binding activity. Indeed, mutations or deletions of G4 binding proteins have been reported to alter the G4 conformation32,33. Therefore, we analyzed the conformational changes in G4-RNA caused by disease-related amino acid substitutions in the GR region.
Alteration of G4-RNA structure after binding to GR mutant proteins
The possible impact of GR mutations on the conformations of G4-RNA, PSD-95 and CaMKIIα was analyzed by CD spectroscopy, which is frequently used to evaluate the conformational properties of nucleic acids. Recently, we identified that the CD spectral patterns of these two G4-RNAs were essentially the same even after binding of the full-length wild-type TDP-43, unlike FUS (fused in sarcoma)20. TDP-43 binds to cover the entire G4-RNA and maintains a stable conformation20. A positive peak near 265 nm and a negative peak near 240 nm characterized the parallel-stranded G4 conformation (Fig. 3A). Wild-type GST-fused wild-type GR segment, GST-GRWT, did not affect the positive peak of the CD spectral pattern, as expected. However, the presence of the mutant proteins, GST-GRG287S or GST-GRM337V, both increased the positive peak due to conformational alternation of G4-RNAs (Figs. 3A,B and S3), which is generally characterized as an altered stacking arrangement of guanine tetrads15,34. In addition, binding of the mutant proteins affected the negative peak of PSD-95 G4-RNA. This observation supported the prediction that the structure of the poly-A loop was affected by the binding35.
The GR mutant proteins alter the G4 structure. (A) CD spectra of G4-RNAs, PSD-95 and CaMKIIα (Table S2) in the absence and presence of wild-type and mutant GST-fused GR proteins. The blue arrows indicated that positive peaks were due to the conformational effects of G4-RNAs. The red arrows indicated that negative peaks were due to the structure of the poly-A loop and were affected by protein binding. CaMKIIα without poly-A did not show this decrease in negative peaks. The x-axis represents wavelength, and the y-axis represents the five scans’ average molar CD (Δε). The CD spectrum values of G4-RNA plus GST-fused GR protein were subtracted from each protein spectrum (For protein spectra, see Fig. S3). (B) The change of the positive peak at 265 nm by the addition of each GST-GR proteins from (A). The experiments were performed thrice, and the y-axis values represents the mean ± SEM. The statistical significance was determined by a two-tailed Student’s t-test *P < 0.05. (C) The measurement of G4 levels using the G4-binding fluorescent sensor TO1B. The fluorescence emission spectra of TO1B after binding with two G4-RNAs (PSD-95 and CaMKIIα) were measured in the absence or presence of GST-GR wild-type or mutant proteins. The buffer control and G4-independent fluorescence intensity are also shown. The graphs on the right showed the average fluorescence intensities at 535 nm. The experiments were performed thrice, and the y-axis values represents the mean ± SEM. A two-tailed Student’s t-test determinedthe statistical significance *P < 0.05. The effects of the three proteins on G4-independent fluorescence intensity are shown in Fig. S4.
Next, the effects on the G4 stabilization were analyzed by a turn-on assay using biotinylated thiazole orange, TO1B. Although, the biotin modification significantly reduces non-specific interactions with RNA, G4 structures can rigidify TO1B and enhance the fluorescence36,37,38. The wild-type GST-GR protein increased G4-dependent fluorescence enhancement, but the ALS-linked mutations G287S and M337V inhibited the increase (Fig. 3C). In the absence of G4, no enhancement of TO1B fluorescence was observed by adding any protein (Fig. S4).
From the CD spectroscopy and the TO1B turn-on assay together, we concluded that mutations altered the maintenance of the G4-RNA conformation. Does this GR mutation-induced structural change in G4 affect other RNA-binding modules such as RRM, altering its overall affinity for G4-RNA? The isolated RRMs have limited individual affinity for RNA, but the participation of partner regions with RNA recognition complements stable binding39. Full-length-TDP-43 specifically recognizes only parallel-stranded G4, whereas the RRM segment alone exhibits broad binding affinities with DNA/RNA including UG-rich sequences12,20,40,41.
RRM recognizes G4-RNA
RRM is one of the most abundant RNA-binding modules42,43. The isolated RRM binds the UG-rich sequence in vitro22,24,44,45, whereas in vivo screening of the full-length TDP-43 interacting RNAs revealed that UG-rich sequences were neither necessary nor sufficient to specify a TDP-43 binding site27. Our previous study obtained by SPR analysis supported these findings and showed that, unlike the readily binding RRM segment, full-length native homodimer has no affinity for UG-rich sequence and binds only parallel-stranded G4 structures12,20. However, it remains unclear whether isolated RRM interacts directly with the G4 motif. Therefore, we performed a gel mobility shift assay to confirm the reproducibility of protein-UG-rich interaction under the same experimental conditions employed in this study (Figs. 1D and 4A). As reported in a previous study12, no binding was detected even when UG10 probe was reacted with 10 times the amount of full-length TDP-43 dimer (Table S2). On the other hand, the isolated RRM segment over-shifted most of the signals even at the lowest amount. The double-mutant RRM segment, which harbors two amino acid substitutions deleterious to RNA binding, GST-RRMwm (R151A and D247A within RRM1 and RRM2)24, showed conditional reduction of binding at low concentrations, but when 10 times the amount was added, a binding signal almost equivalent to that of the wild-type RRM segment was detected. The results obtained corroborated our previous report12; therefore, we analyzed the interaction between RRM and G4-RNA.
The binding specificity of RRM. (A) The structural feature of GST-fused RRM and double-mutant protein, RRMwm are indicated. To confirm the reproducibility under the experimental conditions different from the previous study, the binding to the fluorescently labeled UG10 probe (0.5 pmol) was confirmed by agarose gel mobility shift assay using an increased amount of TDP-43, GST-fused RRM, and RRMwm. (B) Agarose gel mobility shift assay using two G4 probes (0.5 pmol) and increased amount of GST-fused RRM. No band shift was observed in the GST-only control, but a concentration-dependent shift was confirmed with GST-RRM. Lanes 1 and 7 were run simultaneously with unfolded RNA heat denatured with 50% formamide. The experiments were performed thrice, and the y-axis values represent the mean ± SEM.
RRM could confirm the band shift of both PSD-95 G4 and CaMKIIα G4, but there was a difference in the binding affinity, i.e., higher for PSD-95 G4 harboring A-loop (Fig. 4B). While this trend was similar for the full-length dimeric protein (Fig. 1D), there was no significant difference between PSD-95 G4 and CaMKIIα G4 for the GR protein cross-linked with G4-RNA (Fig. 2B). This indicated that the strength of TDP-43 binding to G4-RNA is derived from its interacting affinity to RRM rather than the GR region. At the same time, it was suggested that G4 recognition by RRM is hypersensitive to conformational changes of G4.
Mutations in the GR region interfere with the recruitment of RRM to G4-RNA
Since both GR region and RRM bind to G4-RNA, they may cooperate in a specific recognition and stable binding. To examine this hypothesis, we prepared a His-tagged RRM segment and examined its binding activity to GST-GR in the presence and absence of G4-RNA (Figs. 5A and S5). First, G4-RNA was bound to GST-GRs immobilizing beads, and after washing, His-tagged RRM detected the ternary complex formation. The bound RRM was then detected by western blotting (Fig. 5B). Co-precipitation of the wild-type GST-GR with His-tagged RRM was observed in the presence of G4-RNAs (PSD-95 and CaMKIIα). In contrast, two GR mutant proteins, GST-GRG287S and GST-GRM337V, significantly reduced the levels of His-RRM precipitation. Under this condition, we could not detect any G4-RNA-independent interaction between the RRM and GR regions. This finding indicated the G4 structure dependent intramolecular organization of RRM and GR regions, leading to modulation of its G4-RNA binding property.
GR mutations interfere with the recruitment of RRM. (A) A cartoon of experimental flow. (B) The His-RRM protein (Fig. S5) was co-pulled with GST-GR wild-type and mutant proteins by glutathione sepharose beads (GSB) in the absence and presence of G4-RNAs, PSD-95 and CaMKIIα (Table S2) and detected by western blotting analysis (for original images, see Fig. S6). The graphs show the average of G4 dependent binding ratio compared to the GST-GR wild-type protein and the SEM of three experiments is shown. Two-tailed Student’s t-test was used to determine the statistical significance **P < 0.01.
Discussion
Although TDP-43, the major protein responsible for ALS, is an RNA-binding protein, its binding target of complete dimer form has remained unknown for a long time. Based on a SELEX (Systematic Evolution of Ligands by Exponential Enrichment) screening, we first identified a parallel-stranded G4 structure as a specific binding target, and proposed G4-containing RNAs as the initial selection signal of mRNAs to be transported into neurites for local translation by TDP-4312,15,20. In this mRNA selection process, the GR region is essential. Moreover, the RRM of TDP-43 is a rather promiscuous binding module with a high affinity for RNA. When exposed alone, RRM prefers a UG-repeat sequence for which full-length native TDP-43 has no affinity22,24,35.
We hypothesized that G4-RNA binding affinity exists independently on the RRM and GR regions, and ALS-linked amino acid substitutions in the GR region of TDP-43 simply resulted in reduced binding of the GR region. Unexpectedly, the GR region with the two mutations herein analyzed showed the same G4-RNA binding ability as the wild-type. Hence, it is interesting that the CD spectra and TO1B turn-on assay showed the two amino acid mutations affected the G4 conformation. A subsequent ternary complex formation experiment suggested that the conformational change of G4-RNA interferes with the recruitment of RRM and reduces the binding affinity of TDP-43.
Regarding binding specificity, the essential fact is that the RNA-binding module, the RRM segment, bind the UG-rich sequences, whereas the full-length TDP-43 completely loses this binding capacity. We interpreted that the loss of strong RNA binding affinity may be the result of a perturbation by another intramolecular module such as the GR region, thereby inhibiting the binding to targets other than parallel-stranded G4-RNA. Another possibility is the involvement of the N-terminal domain (NTD). It has been reported that NTD behaves as a scaffold for nucleic acid binding and contributes to binding specificity23,25. Furthermore, a compound that targeting NTD inhibits RRM-RNA binding via allosteric modulation42. However, we have so far failed to detect inhibition of the binding between RRM and UG-rich RNA by an isolated GR region or NTD.
Over 50 missense mutations have been identified in the TDP-43 cording gene in familial and sporadic patients46. Most of these amino acid substitutions were in the GR region. It remains unclear whether all of them interfere with RRM recruitment via the conformational changes in the G4-RNA structure without affecting G4 binding, as determined in this study. Further detailed analysis will be carried out to determine whether there are amino acid substitutions that affect direct interaction with G4. In addition, we recently found that G4-RNA promotes LLPS during FUS RNP granule formation47. The GR region of TDP-43 was involved in LLPS48,49, and interaction with G4-RNA may have complex effects. The insights provide essential information for the studying G4-binding proteins involved in ALS pathogenesis.
Materials and methods
Plasmids
E. coli expression plasmid pGEX-GRWT was described previously20. pGEX-GRG287S and pGEX-GRM337V were constructed by PCR-based mutagenesis method using several primer pairs20. E. coli expression plasmid pET-His-RRM was constructed by replacing the cDNA fragment between the EcoRI and BamHI sites of pGEX-RRM (101-261aa) between the same sites of pET28a ( +) (Merck, NJ USA)12.
Proteins
Full-length, untagged natural dimer proteins TDP-43WT, TDP-43G287S and TDP-43M337V were described previously12,20. Glutathione S-transferase (GST) and GST fusion proteins, GST-GRWT, GST-GRG287S and GST-GRM337V were purified as described20. His-RRM was overproduced in E. coli BL21(DE3) carrying a pET-His-RRM plasmid as noted above. The transformants were grown at 30 °C up to OD600 = 0.4, and then Isopropyl β-D-1-thiogalactopyranoside (IPTG) was added at a final concentration of 0.5 mM. After 4 h of culture at 30 °C, the cells were harvested by centrifugation and suspended in lysis buffer (10% glycerol, 50 mM Tris–HCl pH 8.0, 250 mM NaCl, 10 mM 2-mercaptoethanol, 0.1% Nonidet-P40, 10 mM imidazole and 0.1% benzamidine hydrochloride) containing 1.5 mg/ml lysozyme, stored for 30 min on ice, and then sonicated. After centrifugation, the supernatant was loaded onto the Ni–NTA agarose (Qiagen, Hulsterweg Nederland) open column. The column was washed with lysis buffer and the bound protein was eluted with a buffer containing 250 mM imidazole.
All proteins were further purified by Superdex 75 10/300 GL (Sytiva, MA USA) (10% glycerol, 20 mM HEPES [4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid]-NaOH, pH 6.8, 300 mM NaCl and 0.1 mM EDTA) and the peak fractions were collected with use of ÄKTA explorer 10 S/100 system (Sytiva, MA USA). The purified proteins were concentrated by a centrifugal filter (Vivaspin 6-10 kDa; Sytiva, MA USA) down to 0.5 mg/ml and stored frozen at −80 °C.
Agarose gel mobility shift assay
The fluorescence-labeled oligonucleotides, Cy3-PSD-Ax647, Cy3-CaMKII-Ax647 and UG10- Ax647 (Table S2) (0.5 pmol) were incubated with the indicated amount of diluted protein in a 10 μl buffer (10% glycerol, 20 mM PIPES–NaOH pH 6.8, 0.8 mM MgCl2 and 150 mM NaCl) containing 0.1 mg/ml bovine serum albumin (BSA) and 0.1 mg/ml poly-dI-dC at 25 °C for 30 min. The samples were loaded onto a 1% agarose gel run at 50 V for 60 min at 25 °C. Although the buffer was 0.5 × TBE (pH 6.8), only the dissociation assay was performed under the condition of pH 8.3 (Fig. S1B). The gels were visualized and quantified by Typhoon 9410 imaging system (Sytiva, MA USA). The unfolded RNA samples were prepared by heat denaturation with 50% formamide at 98 °C for 3 min prior to agarose gel electrophoresis, indicated as “UF” in the diagram.
Surface plasmon resonance (SPR) analysis
SPR analysis was carried out as described previously20 in the SPR buffers (20 mM PIPES [1,4-Piperazinediethanesulfonic acid sesquisodium salt]-NaOH, pH 6.8, at 25 °C, 12 mM KCl, 0.8 mM MgCl2, 0.05% Tween-20, and 150 mM NaCl) using a BIAcoreJ instrument (Sytiva, MA USA) at 25 °C. In this method, simultaneous analysis was performed using two flow cells. The dT16 oligomer was immobilized onto flow cell 2 and flow cell 1 was left blank to serve as an in-line reference surface. The analyte protein was injected into the flow cells-1 and -2 of the sensor chip. The plasmon resonance values (resonance unit; RU) were obtained from the flow cell-2 data after subtracting the flow cell-1 data.
Circular dichroism (CD) spectra analysis
For conformational analysis of the wild-type and mutant TDP-43 proteins (0.5 mg/ml), the purified protein samples were diluted in PBS (phosphate-buffered saline) to make a 0.2 μM solution, and then subjected to CD analysis with by using a spectropolarimeter (J-820; Jasco, Tokyo Japan)47. The estimated G4 structure of the synthesized RNAs, PSD-95 and CaMKIIα (Table S2) was confirmed by CD spectra analysis using 1 μM oligonucleotide in each of the SPR buffer conditions at 25 °C by using a spectropolarimeter21. To analyze the influence of protein binding on G4 conformation, parallel-stranded G4-forming RNAs, PSD-95 and CaMKIIα, were confirmed by CD spectra analysis using 1 μM RNA in the presence or absence of 1 μM protein in the CD buffer (20 mM PIPES–NaOH pH 6.8, 0.8 mM MgCl2 and 150 mM NaCl) for 30 min at 25 °C. The scan was repeated five times, and Jasco Spectro Manager compiled the mean values. The x-axis represents the wavelength, whereas the y-axis represents the molar CD (Δε). The content of secondary structure was estimated by using the BeStSel online deconvolution webserver28.
UV cross-linking and gel shift assay
The fluorescence-labeled oligonucleotides, Cy3-PSD-Ax647 and Cy3-CaMKII-Ax647 (Table S2) (0.5 pmol) were incubated with the diluted protein (1, 2, or 4 pmol each) in a 10 μl buffer (10% glycerol, 20 mM PIPES–NaOH pH 6.8, 0.8 mM MgCl2 and 150 mM NaCl) containing 0.1 mg/ml BSA. The mixtures of G4-RNAs and proteins were subjected to cross-linking upon exposure to 254 nm UV at 200 mJ/cm2 using a UV cross-linker, FS-1500 (Funakoshi, Tokyo Japan). The cross-linked samples were separated by 10% SDS-PAGE and the fluorescent signal (Ax647, Alexa Fluor 647) was scanned by Typhoon 9410 imaging system (Sytiva, MA USA)21.
TO1B turn-on assay
The oligonucleotide (20 pmol) was incubated with the protein (40 pmol) in 180 μl buffer (20 mM PIPES-NaOH, pH 6.8, at 25 °C, 0.8 mM MgCl2, 0.05% Tween-20, and 150 mM NaCl) for 10 min at 25 °C. Twenty microliters of 1 μM thiazole orange-3PEG-Biotin, TO1B (ABM, BC Canada) was mixed in the same buffer and stored for 5 min at 25 °C. After 5 min, the ligand-enhanced fluorescence spectra were obtained using a fluorescence spectrometer (Shimadzu RF-5300PC) with an excitation wavelength of 505 nm and a scanning emission wavelength of 520–600 nm. The fluorescence intensities at 535 nm were used to plot the comparison graphs. The oligonucleotides used in this study were obtained from GeneDesign, Inc. (Osaka Japan) (Table S2).
GST pull-down assay
Glutathione sepharose beads (5 μl) (Sytiva, MA USA) immobilized with 4 μg of GST or GST fusion protein were incubated with G4-RNA, PSD-95 or CaMKIIα (Table S2) (800 pmol) in 250 μl of binding buffer at 4 °C for 2 h in binding buffer a 250 μl (10% glycerol, 20 mM PIPES-NaOH pH 6.8, 0.8 mM MgCl2 1.8 mM CaCl2 and 145 mM NaCl, 5.4 mM KCl and 0.05% Tween-20) containing 0.1 mg/ml BSA. These beads were washed two times and incubated with two μg of His-RRM in a 300 μl binding buffer containing 0.1 mg/ml BSA at 4° C for 2 h. These beads were washed thrice, and interacting proteins were eluted with 12.5 μl of 1 × SDS sample buffer. The samples were separated by 12.5% SDS-PAGE, transferred onto a porvinilidendifluoride membrane (Immobilon-P, Merck, NJ USA), and anti-TDP-43 antibodies (Cell Signaling Technology, MA USA) were used for detection as described previously20.
Data availability
The data sets generated during and/or analyzed during the current study are available from the corresponding authors on reasonable request. (akira.ishiguro.iu@hosei.ac.jp).
References
Taylor, J. P., Brown, R. H. Jr. & Cleveland, D. W. Decoding ALS: From genes to mechanism. Nature 539(7628), 197–206 (2016).
Deng, Z., Sheehan, P., Chen, S. & Yue, Z. Is amyotrophic lateral sclerosis/frontotemporal dementia an autophagy disease?. Mol. Neurodegener. 12(1), 90. https://doi.org/10.1186/s13024-017-0232-6 (2017).
Pesiridis, G. S., Lee, V. M. & Trojanowski, J. Q. Mutations in TDP-43 link glycine-rich domain functions to amyotrophic lateral sclerosis. Hum. Mol. Genet. 18(R2), R156–R162 (2009).
Zhao, M., Kim, J. R., van Bruggen, R. & Park, J. RNA-binding proteins in amyotrophic lateral sclerosis. Mol. Cells. 41(9), 818–829 (2018).
Mejzini, R. et al. ALS genetics, mechanisms, and therapeutics: Where are we now?. Front. Neurosci. 13, 1310. https://doi.org/10.3389/fnins.2019.01310 (2019).
Ishiguro, A. & Ishihama, A. Essential roles and risks of G-quadruplex regulation: Recognition targets of ALS-linked TDP-43 and FUS. Front. Mol. Biosci. https://doi.org/10.3389/fmolb.2022.957502 (2022).
Rhine, K., Vidaurre, V. & Myong, S. RNA droplets. Annu. Rev. Biophys. 49, 247–265 (2020).
Jolma, A. et al. Binding specificities of human RNA-binding proteins toward structured and linear RNA sequences. Gen. Res. 30(7), 962–973 (2020).
Kapeli, K., Martinez, F. J. & Yeo, G. W. Genetic mutations in RNA-binding proteins and their roles in ALS. Hum. Genet. 136(9), 1193–1214 (2017).
Mackenzie, I. R. et al. Pathological TDP-43 distinguishes sporadic amyotrophic lateral sclerosis from amyotrophic lateral sclerosis with SOD1 mutations. Ann. Neurol. 61(5), 427–434 (2007).
Sreedharan, J. et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319(5870), 1668–1672 (2008).
Ishiguro, A., Kimura, N., Watanabe, Y., Watanabe, S. & Ishihama, A. TDP-43 binds and transports G-quadruplex-containing mRNAs into neurites for local translation. Genes Cells 21(5), 466–481 (2016).
Rhodes, D. & Lipps, H. J. G-quadruplexes and their regulatory roles in biology. Nucleic Acids Res. 43(18), 8627–8637 (2015).
Varshney, D., Spiegel, J., Zyner, K., Tannahill, D. & Balasubramanian, S. The regulation and functions of DNA and RNA G-quadruplexes. Nat. Rev. Mol. Cell. Biol. 21(8), 459–474 (2020).
Wang, E., Thombre, R., Shah, Y., Latanich, R. & Wang, J. G-Quadruplexes as pathogenic drivers in neurodegenerative disorders. Nucleic Acids Res. 49(9), 4816–4830 (2021).
Ling, S. C. et al. ALS-associated mutations in TDP-43 increase its stability and promote TDP-43 complexes with FUS/TLS. Proc. Natl. Acad. Sci. USA 107(30), 13318–13323 (2010).
Barmada, S. J. et al. Cytoplasmic mislocalization of TDP-43 is toxic to neurons and enhanced by a mutation associated with familial amyotrophic lateral sclerosis. J. Neurosci. 30(2), 639–649 (2010).
Watanabe, S., Kaneko, K. & Yamanaka, K. Accelerated disease onset with stabilized familial amyotrophic lateral sclerosis (ALS)-linked mutant TDP-43 proteins. J. Biol. Chem. 288(5), 3641–3654 (2013).
Shiina, Y., Arima, K., Tabunoki, H. & Satoh, J. TDP-43 dimerizes in human cells in culture. Cell. Mol. Neurobiol. 30(4), 641–652 (2010).
Ishiguro, A. et al. Molecular dissection of ALS-linked TDP-43—Involvement of the Gly-rich domain in interaction with G-quadruplex mRNA. FEBS Lett. 594(14), 2254–2265 (2020).
Ishiguro, A., Katayama, A. & Ishihama, A. Different recognition modes of G-quadruplex RNA between two ALS/FTLD-linked proteins TDP-43 and FUS. FEBS lett. 595(3), 310–323 (2021).
Ayala, Y. M. et al. Human, Drosophila, & C. elegans TDP43: Nucleic acid binding properties and splicing regulatory function. J. Mol. Biol. 348(3), 575–588 (2005).
Chang, C. K. et al. The N-terminus of TDP-43 promotes its oligomerization and enhances DNA binding affinity. Biochem. Bophys. Res. Commun. 425(2), 219–224 (2012).
Lukavsky, P. J. et al. Molecular basis of UG-rich RNA recognition by the human splicing factor TDP-43. Nat. Struct. Mol. Biol. 20(12), 1443–1449 (2013).
Mompeán, M. et al. The TDP-43 N-terminal domain structure at high resolution. FEBS J. 283(7), 1242–1260 (2016).
Subramanian, M. et al. G-quadruplex RNA structure as a signal for neurite mRNA targeting. EMBO Rep. 12(7), 697–704 (2011).
Polymenidou, M. et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat. Neurosci. 14(4), 459–468 (2011).
Micsonai, A. et al. BeStSel: webserver for secondary structure and fold prediction for protein CD spectroscopy. Nucleic Acids Res. 50(W1), W90–W98. https://doi.org/10.1093/nar/gkac345 (2022).
Sen, D. & Gilbert, W. Formation of parallel four-stranded complexes by guanine-rich motifs in DNA and its implications for meiosis. Nature 334(6180), 364–366 (1988).
Blice-Baum, A. C. & Mihailescu, M. R. Biophysical characterization of G-quadruplex forming FMR1 mRNA and of its interactions with different fragile X mental retardation protein isoforms. RNA 20(1), 103–114 (2014).
Haeusler, A. R. et al. C9orf72 nucleotide repeat structures initiate molecular cascades of disease. Nature 507(7491), 195–200 (2014).
Kosiol, N., Juranek, S., Brossart, P., Heine, A. & Paeschke, K. G-quadruplexes: A promising target for cancer therapy. Mol. Cancer. 20(1), 40. https://doi.org/10.1186/s12943-021-01328-4 (2021).
Lyu, K. et al. An RNA G-quadruplex structure within the ADAR 5’UTR interacts with DHX36 helicase to regulate translation. Angew. Chem. Int. Ed. Engl. 61(52), e202203553. https://doi.org/10.1002/anie.202203553 (2022).
Do, N. Q., Lim, K. W., Teo, M. H., Heddi, B. & Phan, A. T. Stacking of G-quadruplexes: NMR structure of a G-rich oligonucleotide with potential anti-HIV and anticancer activity. Nucleic Acids Res. 39(21), 9448–9457 (2011).
Zeraati, M. et al. Cancer-associated noncoding mutations affect RNA G-quadruplex-mediated regulation of gene expression. Sci. Rep. 7(1), 708. https://doi.org/10.1038/s41598-017-00739-y (2017).
Jeng, S. C., Chan, H. H., Booy, E. P., McKenna, S. A. & Unrau, P. J. Fluorophore ligand binding and complex stabilization of the RNA Mango and RNA Spinach aptamers. RNA 22(12), 1884–1892 (2016).
Trachman, R. J. 3rd. et al. Structural basis for high-affinity fluorophore binding and activation by RNA Mango. Nat. Chem. Biol. 13(7), 807–813 (2017).
Umar, M. I., Ji, D., Chan, C. Y. & Kwok, C. K. G-quadruplex-based fluorescent turn-on ligands and aptamers: From development to applications. Molecules https://doi.org/10.3390/molecules24132416 (2019).
Ozdilek, B. A. et al. Intrinsically disordered RGG/RG domains mediate degenerate specificity in RNA binding. Nucleic Acids Res. 45(13), 7984–7996 (2017).
Furukawa, Y. et al. A molecular mechanism realizing sequence-specific recognition of nucleic acids by TDP-43. Sci. Rep. 6, 20576. https://doi.org/10.1038/srep20576 (2016).
Zacco, E., Martin, S. R., Thorogate, R. & Pastore, A. The RNA-recognition motifs of TAR DNA-binding protein 43 may play a role in the aberrant self-assembly of the protein. Front. Mol. Neurosci. 11, 372. https://doi.org/10.3389/fnmol.2018.00372 (2018).
Auweter, S. D., Oberstrass, F. C. & Allain, F. H. Sequence-specific binding of single-stranded RNA: Is there a code for recognition?. Nucleic Acids Res. 34(17), 4943–4959 (2006).
Lunde, B. M., Moore, C. & Varani, G. RNA-binding proteins: modular design for efficient function. Nat. Rev. Mol. Cell. Biol. 8(6), 479–490 (2007).
Buratti, E. et al. Nuclear factor TDP-43 and SR proteins promote in vitro and in vivo CFTR exon 9 skipping. EMBO J. 20(7), 1774–1784 (2001).
Rengifo-Gonzalez, J. C. et al. The cooperative binding of TDP-43 to GU-rich RNA repeats antagonizes TDP-43 aggregation. eLife https://doi.org/10.7554/eLife.67605 (2021).
Buratti, E. Functional significance of TDP-43 mutations in disease. Adv. Genet. 91, 1–53 (2015).
Ishiguro, A., Lu, J., Ozawa, D., Nagai, Y. & Ishihama, A. ALS-linked FUS mutations dysregulate G-quadruplex-dependent liquid-liquid phase separation and liquid-to-solid transition. J. Biol. Chem. 297(5), 101284 (2021).
Li, H. R., Chiang, W. C., Chou, P. C., Wang, W. J. & Huang, J. R. TAR DNA-binding protein 43 (TDP-43) liquid-liquid phase separation is mediated by just a few aromatic residues. J. Biol. Chem. 293(16), 6090–6098 (2018).
Conicella, A. E. et al. TDP-43 α-helical structure tunes liquid-liquid phase separation and function. Proc. Natl. Acad. Sci. USA 117(11), 5883–5894 (2020).
Acknowledgements
We thank Dr. Daisaku Ozawa and Dr. Masako Ishiguro for their support and helpful advice.
Funding
This work was supported by the Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Scientific Research (17K07291, 22K07032 and 23H04265 to A. Ishiguro).
Author information
Authors and Affiliations
Contributions
A. Ishiguro and A. Ishihama designed the study; A. Ishiguro performed most of the experiment, and A. Ishiguro and A. Ishihama wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ishiguro, A., Ishihama, A. ALS-linked TDP-43 mutations interfere with the recruitment of RNA recognition motifs to G-quadruplex RNA. Sci Rep 13, 5982 (2023). https://doi.org/10.1038/s41598-023-33172-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-33172-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.