Abstract
Bladder cancer is the second most prevalent urological malignancy. It's a big contributor to cancer-related deaths throughout the globe. Researchers discovered that the hedgehog signaling (HhS) pathway contributed to the onset and spread of many different kinds of cancer. Nevertheless, the present understanding of the function of HhS in the bladder cancer molecular landscape is incomplete. Raw data were gotten from the IMvigor210, the Gene Expression Omnibus, and The Cancer Genome Atlas databases. Bioinformatics was used to examine the HhS score of each sample, and the enrichment of differentially expressed genes (DEGs), differentiation characteristics, immunological infiltration, and metabolic activity. The HhS prognostic signature was developed with significant assistance from the least absolute shrinkage and selection operator regression and Cox regression. An HhS-related nomogram was developed to assist in the prediction of patients’ survival probability. We found that HhS was linked to poor prognosis in bladder cancer, and its activation was linked to the Basal subtype of bladder cancer. Bladder cancer with high HhS activity has higher glycolysis, nucleotide metabolism, amino acid metabolism, and other cancer-promoting metabolic activities. Furthermore, HhS mediates an immunosuppressive microenvironment in bladder cancer on the basis that HhS negatively correlates with the CD8 + T cells and correlates positively with immune checkpoints and T cell exhaustion scores. Finally, an HhS-related signature was developed for predicting the prognosis of patients with bladder cancer. Targeting HhS may be a potential therapy choice for bladder cancer.
Similar content being viewed by others
Introduction
Bladder cancer is the second most prevalent urological cancer. It is also one of the leading causes of cancer-related deaths worldwide today. According to statistics, there are more than 500,000 new cases of bladder cancer and approximately 213,000 bladder cancer-related deaths each year1. Although a variety of treatments have been developed, including neoadjuvant, adjuvant chemotherapy, immunotherapy, and targeted therapies, resulting in a high survival rate for bladder cancer. However, due to the propensity for recurrence and metastasis, most patients must undergo lifelong surveillance2,3. Despite having similar clinical and histopathological features, bladder cancer exhibits substantial variation in disease aggressiveness and response to therapy due to tumor heterogeneity2,3. Therefore, investigations of the relevant biomarkers with predictive value for clinical outcomes and treatment efficacy to develop reliable tools to guide individualized treatment need to be expedited.
Hedgehog (Hh) was originally discovered in the early 1980s4 in Drosophila by Nüsslein-Volhard and Wieschaus. Binding among Indian hedgehog (IHH), Sonic hedgehog (SHH), and Desert hedgehog (DHH) ligands attenuates the suppressive effect of their Patched (PTCH) transmembrane proteins/receptors on Smoothened (SMO), which is also located in the cell membrane5. The signaling cascade initiated by SMO culminates in the activation and nuclear deployment of GLI gene transcription elements, thus inducing the expression of Hh target genes; the majority of these genes are involved in proliferation, survival, and angiogenesis. The Hh signaling (HhS) pathway is aberrantly activated by overexpression of Hh ligands, loss of receptor function, or dysregulation of transcription factors5. Evidence suggests that The constitutive activation of the HhS pathway is definitely linked to the onset and progression of several types of cancer, such as lung, gastric, prostate, hepatocellular, pancreatic, colorectal, esophageal, and ovarian cancers6,7. Moreover, evidence shows that multiple uroepithelial cancer cell lines exhibit constitutive HhS8. Given the functional significance of HhS in cancer and its crucial role in the disease, novel strategies targeting this pathway seem to hold promising therapeutic options for bladder cancer treatment9. However, the current comprehension of the function of HhS in the bladder cancer molecular landscape is incomplete.
The aim of this study is to determine HhS's predictive value for clinical outcomes in bladder cancer, analyze its phenotypic characteristics, and its potential impact on the immune environment and molecular landscape of bladder cancer. Finally, to construct and validate an HhS prognostic signature, thereby offering novel perspectives for the development of personalized therapy for bladder cancer.
Materials and methods
Data processing
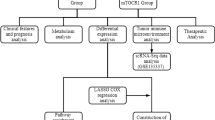
Figure 1 clearly shows the study’s framework design. For TCGA-BLCA, RNA-seq data from 408 BLCA patients in fragments per kilobase of exon per million mapped fragments (FPKM) values and matched medical data were retrieved from the UCSC Xena’s data portal. Subsequently, FPKM values were converted into transcripts per kilobase million (TPM) values. Somatic mutation data were obtained utilizing the R package TCGAbiolinks10 and then grouped in the Mutation Annotation Format (maf) form, followed by evaluation using the R package maftools.
From GEO, GSE31684 datasets 11 contained 93 bladder cancer samples that were used for external validation. In addition, a single-cell RNA sequencing (scRNA-seq) dataset of bladder cancer (GSE13533712) was used for scRNA-seq analysis.
For the BLCA immunotherapy-related cohort, the complete expression data as well as the detailed clinical information of 348 BLCA patients that received atezolizumab treatment were obtained by using the R package IMvigor210CoreBiologies13.
Computation of the enrichment scores of different gene signatures
Utilizing nonparametric and unsupervised methods, gene set variation analysis (GSVA) can predict the specific pathway’s activity based on transcriptomic data14. HhS gene signature were acquired from the Molecular Signatures Database (MsigDB, https://www.gsea-msigdb.org/gsea/downloads.jsp) hallmark gene set collection The Bladder Cancer Molecular Taxonomy Group-related study yielded 12 bladder cancer signatures that are specific to various molecular subtypes 15. Metabolism-related gene sets were obtained from KEGG gene set collections on MsigDB. Therapeutic-related signatures, including immune-inhibited oncogenic pathways, targeted therapy-linked gene signatures, and gene signatures anticipating radiotherapy responses were collected from the study of Hu et al.16. All the gene sets are shown in Table S1.
Screening and functional annotation of differentially expressed genes (DEGs)
DEGs were screened by the limma R package. P < 0.05 and fold change (FC) > 3/2 were set as the selection conditions for screening the downregulated and upregulated DEGs. The ClusterProfiler package performed Gene Ontology (GO), KEGG enrichment, and hallmark gene set enrichment. The R package ClusterProfiler, on the other hand, was used to perform gene set enrichment analysis (GSEA).
Prediction of the molecular subtypes in bladder cancer
ConsensusMIBC R packages was utilized to ascertain the molecular subtypes of bladder cancer in different molecular subtype systems, encompassing CIT, MDA, Lund, Baylor, TCGA, and UNC subtypes.
Evaluating immune cell infiltration
CIBERSORT, which can identify 22 tumor-infiltrating immune cells from each specimen, was used to measure the cellular density of the immune cells. The cancer immunity cycle gene sets can be grouped into seven stages: Stage 1 is the production of cancer cell antigens, Stage 2 is the presentation of cancer antigen, Stage 3 is the priming and activation, Stage 4 is the trafficking of immune cells to tumors, Stage 5 is the infiltration of immune cells to tumors, Stage 6 is the identification of cancer cells by T cells, and Stage 7 is the killing of cancer cells17. Then GSVA calculated the score of these gene sets.
Predicting response to chemotherapy
The IC50 of common chemotherapeutic drugs was estimated by using the “pRRophetic” R package. The Drugbank database (https://go.drugbank.com/) was employed in the screening of the drug-target genes.
Construction of the HhS-related signature (HhSRS)
The HhSRS was developed on the basis of the TCGA-BLCA cohort. The analysis of the univariate Cox regression was performed to find genes linked to overall survival (OS) from DEGs between the low-and-high- HhS group. Next, the LASSO regression played a crucial function in selecting candidate genes. Then, utilizing the multivariate Cox regression, the regression coefficient as well as the multiple regression model of genes linked to survival were determined. In these steps, the HhSRS was developed as follows: Risk score = Σn1 coefi*xi. The HhSRS was used to divide the groups into low- and high-risk categories, with the median score serving as the cutoff value.
Construction of the HhSRS-related nomogram
This study used a multivariate Cox analysis of HhSRS grouping and clinical variables to identify potential independent prognostic indicators. The HHSRS-related nomogram was then constructed using the regplot software, using age, stage, and HhSRS group as parameters. To further aid in the display of the closeness between the actual and expected OS, calibration curves were generated.
scRNA-Seq data analysis
The Seurat R package was used to perform unsupervised clustering of single cells using the read count matrix as input. The cell clustering, quality control, and annotation of scRNA-seq data were performed as per the previous description18. CellChat was employed in the intercellular communication network-related analysis from scRNA-seq data. The cytotoxic signature and exhaustion score of T cells were from the study of Zheng et al.(Table S2)19.
Statistical analysis
The Pearson correlation analysis played a crucial role in the examination of correlations between variables. A t-test was employed in the comparison of continuous variables among binary groups that conform to a normal distribution. For comparisons of more than two groups, Kruskal–Wallis tests were used to compare the differences. The log-rank test was used to uncover statistically significant differences, and the Kaplan–Meier approach generated survival curves for the subgroups in each data set. SPSS 22.0, SangerBox ( http://sangerbox.com )20 and R 4.0.0 implemented all statistical analyses. P values were two-sided. P values < 0.05 were statistically significant.
Result
HhS activation is a marker of poor prognosis in BLCA patients
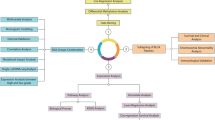
First, HhS scores were calculated and its expression characteristics were analyzed for patients in the TGCA-BLCA cohort. The results showed that HhS was overactivated in patients with higher stage and older age (Fig. 2A, B). And there is no significant difference in HhS scores among patients of different genders (Fig. 2C). Next, patients were classified into a high-HhS group and a low-HhS group as per the median as the cut-off value (Fig. 2D). Variations in the expression of 36 genes involved in the HhS pathway were analyzed between the two groups of patients, and the findings affirmed that 28 of the 36 genes involved in HhS were remarkably expressed in the high HhS group (Fig. 2D).
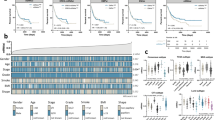
HhS scores, survival status, and expression of 36 HhS pathway genes in TCGA-BLCA patients were shown in Fig. 3A. KM analysis was performed and exhibited that the prognosis was worse in the high HhS group (Fig. 3B). Then, multivariate Cox regression analysis in the study, deduced that HhS acted as an independent prognostic f in bladder cancer patients (Fig. 3C).
These results indicated that HhS activation is a marker of poor prognosis in BLCA patients.
Discovery of DEGs and functional annotations
Regarding the DEGs, 551 downregulated DEGs and 2498 upregulated DEGs that belonged to the high HhS group were screened (Fig. S1A, B). Afterward, functional annotations of DEGs were performed. KEGG analysis showed that upregulated DEGs were mainly involved in “Pathways in cancer”, “Cytokine-cytokine receptor interaction” and “PI3K-Akt signaling pathway” (Fig. 4A). GO analysis demonstrated that upregulated DEGs were remarkably enriched in “Extracellular region” (cellular component, CC) “Immune system process” (biological process, BP), and “Antigen binding” (molecular function, MF) (Fig. 4B). The results of Hallmark gene set analysis showed that “Epithelial-mesenchymal transition”, “Inflammatory response”, and “TNFa signaling via NK-KB” (Fig. 4C).
Functional annotation by KEGG, GO and Hallmark gene sets enrichment. (A–C) (A) KEGG, (B) GO and (C) Hallmark pathways of upregulated DEGs. (D–F) (D) KEGG, (E) GO and (F) Hallmark pathways of downregulated DEGs. (G–H) Functional analysis of HhS by GSEA based on gene sets from (G) KEGG and (H) Hallmark gene sets.
For the downregulated DEGs, the KEGG pathways were “Chemical carcinogenesis”, “Metabolic pathways”, “Xenobiotics metabolism by cytochrome P450”, etc. (Fig. 4D). GO analysis manifested that the most substantially enriched pathways were “Epithelial cell differentiation” in BP, “Troponin complex” in CC, and “Oxidoreductase activity” in MF (Fig. 4E). And Hallmark gene sets analysis demonstrated that “Estrogen response early” “Estrogen response late”, and “Xenobiotic metabolism” were enriched (Fig. 4F).
Next, GSEA was performed and the results showed that the KEGG “Glioma”, “Long term potentiation”, and “JAK-STAT signaling pathway”, and the Hallmark “KRAS signaling up”, “UV response up” and “Myogenesis” were the most enriched in high HhS patients (Fig. 4G, H).
These results indicated that HhS was associated with differentiation characteristics, metabolism, and immunity in bladder cancer.
Association between HhS and molecular characteristics
The molecular subtypes of bladder cancer are closely linked to prognosis as well as response to chemotherapy, immunotherapy, and other therapies15,21. The association between HhS and molecular subtypes of bladder cancer was analyzed (Table. 1), and the results showed that six different molecular subtype prediction algorithms all suggest that bladder cancer with high expression of HhS is more likely to be basal type bladder cancer. And the analysis of molecular characteristics affirmed that the patients in the high HhS group had higher immune differentiation, basal differentiation, EMT differentiation, myofibroblasts, keratinization, smooth muscle, neuroendocrine differentiation, and interaction response scores, but lower urinary differentiation, Ta path and Luminal differentiation (Fig. 5A, B).
Correlation analysis between HhS and molecular subtype in bladder cancer. (A) Differential expression of specific bladder cancer-related signatures in high and low HhS score group patients. (B) Heatmap of specific bladder cancer-related signatures in high and low HhS score group patients. *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001.
Correlations between HhS and metabolism pathways
Next, the association of HhS with metabolic pathways was explored (Fig. 6). The patients in the high HhS group showed extensive upregulation of metabolic activity. Among all 70 KEGG metabolic pathways, 42 were upregulated and 12 were downregulated. Patients with high HhS have significantly activated glycolysis, pentose phosphate pathway, and nucleotide metabolism, which are considered to be the dependent pathways of malignant tumor progression22. And the pathways related to drug metabolism, including "Xerobiotics’ metabolism by cytochrome p450" and "Drugs’ metabolism cytochrome p450", were inhibited, which suggests that related drugs may be more effective for patients with high HhS.
HhS relation to tumor immune microenvironment (TIME)
We evaluated the expression levels of immune checkpoint genes, the activity of the cancer immunity cycle, and the infiltration status of tumor-infiltrating immune cells to investigate the possible link between HhS and immunological features (Fig. 7). First, the CIBERSORT algorithm was used to deconvolute the infiltration of immune cells in TME. It was ascertained that patients with high expression of HhS had less infiltration of memory B cells, naive CD4 + T cells, CD8 + T cells, monocytes, resting NK cells, and the activated dendritic cells (DCs), and higher infiltration of M0 macrophages, naive B cells, M2 macrophages and M1 macrophages (Fig. 7A). And HhS scores were positively linked to a majority of immune checkpoints (Fig. 7B). Interestingly, the results of immunity cycle analysis deduced that HhS score was positively linked to positive and negative regulation scores of all immune cycle steps (Fig. 7C).
Correlation analysis between UCPs and tumor immune microenvironment. (A) Heatmap of immune cells in high and low HhS score group patients. (B–C) Correlation heatmap (up) of (B) immune checkpoints and (C) immune cycle score correlation with HhS and violin plot (down) in high and low HhS groups. *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001.
To further explore the function of HhS in TIME, scRNA-seq data of seven bladder cancer patients were obtained from GSE135337 and were annotated (Fig. 8A, B). Seven samples were classified into a low-HhS group and a high-HhS group as per the average HhS expression of samples (Fig. 8C). We used CellChat to infer the presumed intercellular interaction based on ligand-receptor signal transduction and observed that the T cell signal in the high HhS group was weakened (Fig. 8D, E). Consistently, T cells in the high HhS group showed higher exhaustion scores (Fig. 8F).
High HhS implies an Immunosuppressed tumor microenvironment in the single cell cohort. (A, B) t-SNE plot showing the composition of (A)7 bladder cancer samples and (B) 6 main subtypes. (C) Distribution in the HhS of seven bladder cancer samples (divided into 2 patterns). (D–E) Circos plots displaying putative ligand-receptor interactions between T cells and other cell clusters from (D) high-HhS and (E) low-HhS group. The brand links pairs of interacting cell types, and corresponding number of events were labeled in the graph. (F) Differences in cytotoxic score and exhausted scoreof T cells between high and low HhS groups. *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001.
These results suggest a significant antagonistic effect between immune invasion and immunosuppression in tumor tissues with high HhS and an overall predominance of immunosuppression.
HhS predicts therapeutic opportunities
Next, the association of HhS with chemotherapy, radiotherapy, and numerous targeted therapies' clinical responses was analyzed. The high HhS group showed higher enrichment scores for EGFR network (EGFR ligands) and radiotherapy-predicted pathways (cell cycle, hypoxia, and DNA replication), while the low HhS group expressed higher enrichment for immune-inhibited oncogenic pathways (PPARG network, WNTγ catenin network, IDH1, VEGFA) (Fig. 9A). The IC50 values calculated by pRRophetic revealed that the high HhS group had higher sensitivity to gemcitabine and cisplatin, chemotherapeutic agents commonly used in bladder cancer (Fig. 9B). In addition, results from the Drugbank database showed that the high HhS group expressed higher levels of EGFR inhibitors as well as chemotherapeutic agents-related targets (Fig. 9C). In addition, data from the IMvigor210 cohort showed that patients who were effective after receiving immunotherapy expressed lower HhS (Fig. 9D).
Correlation between HhS and the therapeutic response to several therapies in bladder cancer (A–C) Differential expression of (A) enrichment scores of therapeutic signatures such as targeted therapy and radiotherapy, (B) IC50 of gemcitabine and Cisplatin, and (C) drug-target genes in high and low HhS groups. (D) HhS score in patients with different clinical response of cancer immunotherapy in the IMvigor210 cohort.
The mutation profile was analyzed in the TCGA-BLCA cohort (Fig. S2). The low HhS group showed higher overall mutation frequency and had higher mutation frequencies of RYR2, FGFR3, and ELF3.
HhSRS development and validation
High- and low-HhS groups were compared using univariate Cox regression to identify 1290 DEGs from DEGs, and HhSRS was then developed. LASSO regression analysis (Fig. 10A, B) and multivariate Cox regression (Fig. 10C) analysis were used to reduce the overfitting of the HhSRS. Ultimately, 2 genes were recognized to derive the HhSRS: Risk score = 0.14135* cerebral endothelial cell adhesion molecule (CERCAM)-0.21481* granulysin (GNLY). ScRNA-seq analysis showed that GNLY was primarily expressed in T cells (Fig. 10D) and CERCAM was mainly shown in fibroblasts (Fig. 10E).
Patients were divided into two groups based on their HhSRS median (Fig. 11A). Then, the predictive efficacy of HhSRS was confirmed using KM survival curves (Fig. 11B). Patients in the TCGA-BLCA cohort with a high HhSRS risk experienced a significantly lower chance of survival. The HhSRS was next subjected to test in the GSE31684 cohort, where it also showed promising findings (Fig. 11C, D). In the IMvigor210 cohort, we then re-verified the prognostic impact of HhSRS on immunotherapy, and similarly, patients with low HhSRS had a considerably better prognosis than those with high HhSRS (Fig. 11E, F).
We resorted a nomogram to build a predictive model, taking clinicopathological covariates into account, thus establishing a clinically effective technique that could predict the likelihood that a patient would survive. Based on the cox analysis (Fig. 12A), a nomogram to predict OS rates was built (Fig. 12B). Further, the calibration curves showed that the HhSRS-related nomogram accurately predicted the likelihood of survival (Fig. 12C).
Establishment of the HhSRS related nomogram. (A) The forest plots for multivariate regression of clinical factors and HhSRS. (B) Nomogram for predicting the OS probability, by setting age, HhSRS group and tumour stage as parameters. (C) Calibration curves for verifying the prediction accuracy; red represents the 1-year prediction, blue represents the 3-year prediction, and green represents the 5-year prediction.
Discussion
Bladder cancer comprises various heterogeneous subtypes, and every subtype possesses its own clinical and biological characteristics15. The identification of reliable prognostic biomarkers to pinpoint bladder cancer patients at high risk who might benefit from aggressive treatment represents one of the latest hotspots in the disease23. Dysfunction or aberrant activation of the HhS pathway is linked to developmental deformities and cancers5. Shin et al. found that when invasive urothelial carcinoma develops, SHH expression is lost inevitably. Inhibiting stromal response to SHH genetically hastens disease progression and shortens the time of survival24. As per findings of a few studies, dysregulation of SHH ligand or one of its downstream mediators, for instance, PTCH1, SMO, or GLI1, has been linked to urothelial carcinoma initiation and progression8,25,26 and in modulating cancer stem cell activities27. Uncontrolled activation of the HhS pathway is linked to tissue healing and tissue homeostasis, whereas regulated activation of the pathway enhances cancer advancement28. Numerous small molecules have been identified as HhS pathway antagonists at various levels to date6. However, due to technical constraints, most previous studies have been limited to focusing on a few key proteins in HhS rather than the entire pathway.
The fast development of transcriptome, genome, and bioinformatics approaches over the last decade has greatly aided therapeutic options pertaining to cancer via their impact on the discovery of cancer biomarkers and the personalization of cancer therapy29,30,31. The present research confirmed that HhS may be a key factor for bladder cancer-related prognosis and is linked to poor outcomes based on a bioinformatics technique and RNA-seq data. Furthermore, a reliable HhSRS was created for bladder cancer-related prognosis prediction, thus aiding in improving the diagnosis and individualization of treatment for bladder cancer patients.
Molecular subtyping may provide light on the molecular heterogeneity of bladder cancer and can be used to make predictions about patients’ prognoses and responses to therapy15,21. The results of this research suggest that Basal bladder cancers have significantly elevated HhS activation. In terms of mechanism, basal bladder cancers have higher invasiveness, which relies on their higher levels of cancer stem cells and EMT characteristics32. And sustained activation of HhS contributes to tumor stem cell characteristics and EMT process activation33,34. Therefore, blocking the HhS pathway may be a potential approach to improve survival in patients with basal bladder cancer.
Cancer cells have remarkable metabolic plasticity to obtain nutrients for growth and survival under the harshest conditions, with aerobic glycolysis (Warburg effect) being one of the best-known features of tumor metabolic reprogramming22. Previous evidence suggests that activation of HhS results in a cellular transition to aerobic glycolysis35. Our study shows that patients with high HhS have elevated levels of amino acid metabolism, glycolysis, pentose phosphate pathway, and nucleotide metabolism, which are all fuels required for malignant cell growth and proliferation. Thus, Hh activity regulation is involved in cancer cell metabolic plasticity in migration and metastasis.
Anticancer immunotherapy is vital in the treatment strategy of bladder cancer, and immune checkpoint inhibitors (ICIs), anti-PD1 or anti-PD-L1 antibodies, have exhibited efficacy in clinical trials alone or in conjunction with chemotherapy36. Nevertheless, the mechanisms of resistance to these treatments are largely unknown. The association of Hh signaling with immunotherapy has been explored previously. Although ICIs maintained strong anti-tumor activity in the Hh-independent form of medulloblastoma, the work by Pham et al. confirmed that they had low efficacy in the Hh-dependent model37. Low immune infiltration and poor response to ICIs in cancer patients are linked to the Wnt pathway38, which is mechanistically related to HhS signaling39. Consistently, the findings of the presented study affirmed that bladder cancer patients with higher HhS had less CD8 + T cell infiltration and their T cells exhibited a higher depletion profile, in addition, patients who were not sensitive to immunotherapy had Higher HhS scores. Therefore, combination therapy of HhS signaling pathway targeting and immune checkpoint blockers may benefit bladder cancer patients.
Our study identified two genes, namely, GNLY and CERCAM, which were linked to high-risk and low-risk, correspondingly. GNLY, belonging to the saposin-like family of proteins, is a cytolytic and proinflammatory molecule constitutively. GNLY was previously revealed to be also linked to the cytotoxicity and characteristic of effector memory T cells in metastatic melanoma patients40. Consistently, the current study also showed that GNLY is specifically expressed in bladder cancer T cells. CERCAM is an adhesion molecule whose expression is at high levels in brain microvasculature, which leukocytes can use to stretch across the blood–brain barrier, regulate leukocyte migration capacity and adhesion, and has been found to be a negative prognostic marker in several tumors41,42,43. Its overexpression promotes cell viability, proliferation, and invasion and is associated with the PI3K/AKT pathway44. The presented study found that the expression of CERCAM was higher in fibroblasts than in cancer cells, so further studies are needed to investigate its role in TME.
Our study still had some limitations. Because raw data were gathered from publicly available datasets, the study was retrospective. Further in vitro or in vivo testing is required to confirm the findings. The two independent prognostic genes' fundamental processes were not further investigated. In summary. to confirm the findings and define the putative molecular mechanism at play, functional studies of the two independent prognostic genes were necessary.
Conclusion
This study demonstrates that HhS is a poor prognostic factor in bladder cancer and mediates the association of bladder cancer with molecular subtypes, metabolic promotion, and immunosuppression. In addition, HhS can predict bladder cancer response to several therapies. Therefore, it seems that targeting HhS might be a useful strategy for treating bladder cancer.
Data availability
The raw data of our study were downloaded from TCGA dataset (http://cancergenome.nih.gov/), GEO dataset (GSE31684, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE31684; GSE135337, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE135337) and IMvigor210CoreBiologies dataset (http://research-pub.gene.com/IMvigor210CoreBiologies/).
Abbreviations
- Hh:
-
Hedgehog
- SHh:
-
Sonic hedgehog
- IHh:
-
Indian hedgehog
- DHh:
-
Desert hedgehog
- Smo:
-
Smoothened
- Ptch:
-
Patched
- HhS:
-
Hh signaling
- TPM:
-
Transcripts per kilobase million
- FPKM:
-
Fragments per kilobase of exon per million mapped fragments
- MAF:
-
Mutation annotation format
- GEO:
-
Gene expression omnibus
- scRNA-seq:
-
Single-cell RNA sequencing
- MsigDB:
-
Molecular signatures database
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- DEGs:
-
Differentially expressed genes
- GO:
-
Gene ontology
- GSEA:
-
Gene set enrichment analysis
- FC:
-
Fold change
- OS:
-
Overall survival
- LASSO:
-
Least absolute shrinkage and selection operator
- BP:
-
Biological process
- CC:
-
Cellular component
- MF:
-
Molecular function
- TIME:
-
Tumor immune microenvironment
- CERCAM:
-
Cerebral endothelial cell adhesion molecule
- GNLY:
-
Granulysin
- ICIs:
-
Immune checkpoint inhibitors
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 70, 7–30. https://doi.org/10.3322/caac.21590 (2020).
Sanli, O. et al. Bladder cancer. Nat. Rev. Dis. Primers 3, 17022. https://doi.org/10.1038/nrdp.2017.22 (2017).
Witjes, J. A. et al. European association of urology guidelines on muscle-invasive and metastatic bladder cancer: Summary of the 2020 guidelines. Eur. Urol. 79, 82–104. https://doi.org/10.1016/j.eururo.2020.03.055 (2021).
Nüsslein-Volhard, C. & Wieschaus, E. Mutations affecting segment number and polarity in Drosophila. Nature 287, 795–801. https://doi.org/10.1038/287795a0 (1980).
Xin, M., Ji, X., De La Cruz, L. K., Thareja, S. & Wang, B. Strategies to target the hedgehog signaling pathway for cancer therapy. Med. Res. Rev. 38, 870–913. https://doi.org/10.1002/med.21482 (2018).
Hanna, A. & Shevde, L. A. Hedgehog signaling: Modulation of cancer properies and tumor mircroenvironment. Mol. Cancer 15, 24. https://doi.org/10.1186/s12943-016-0509-3 (2016).
Ruch, J. M. & Kim, E. J. Hedgehog signaling pathway and cancer therapeutics: Progress to date. Drugs 73, 613–623. https://doi.org/10.1007/s40265-013-0045-z (2013).
Fei, D. L. et al. Hedgehog signaling regulates bladder cancer growth and tumorigenicity. Cancer Res. 72, 4449–4458. https://doi.org/10.1158/0008-5472.CAN-11-4123 (2012).
Salaritabar, A. et al. Targeting hedgehog signaling pathway: Paving the road for cancer therapy. Pharmacol. Res. 141, 466–480. https://doi.org/10.1016/j.phrs.2019.01.014 (2019).
Colaprico, A. et al. TCGAbiolinks: An R/Bioconductor package for integrative analysis of TCGA data. Nucleic acids Res. https://doi.org/10.1093/nar/gkv1507 (2016).
Riester, M. et al. Combination of a novel gene expression signature with a clinical nomogram improves the prediction of survival in high-risk bladder cancer. Clin. Cancer Res. 18, 1323–1333. https://doi.org/10.1158/1078-0432.CCR-11-2271 (2012).
Lai, H. et al. Single-cell RNA sequencing reveals the epithelial cell heterogeneity and invasive subpopulation in human bladder cancer. Int. J. Cancer 149, 2099–2115. https://doi.org/10.1002/ijc.33794 (2021).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548. https://doi.org/10.1038/nature25501 (2018).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7. https://doi.org/10.1186/1471-2105-14-7 (2013).
Kamoun, A. et al. A consensus molecular classification of muscle-invasive bladder cancer. Eur. Urol. 77, 420–433. https://doi.org/10.1016/j.eururo.2019.09.006 (2020).
Hu, J. et al. Siglec15 shapes a non-inflamed tumor microenvironment and predicts the molecular subtype in bladder cancer. Theranostics 11, 3089–3108. https://doi.org/10.7150/thno.53649 (2021).
Xu, L. et al. TIP: A web server for resolving tumor immunophenotype profiling. Cancer Res. 78, 6575–6580. https://doi.org/10.1158/0008-5472.CAN-18-0689 (2018).
Liu, Z. et al. Lipid-associated macrophages in the tumor-adipose microenvironment facilitate breast cancer progression. OncoImmunology 11, 2085432. https://doi.org/10.1080/2162402X.2022.2085432 (2022).
Zheng, Y. et al. Immune suppressive landscape in the human esophageal squamous cell carcinoma microenvironment. Nat. Commun. 11, 6268. https://doi.org/10.1038/s41467-020-20019-0 (2020).
Shen, W. et al. Sangerbox A comprehensive, interaction-friendly clinical bioinformatics analysis platform. iMeta 1, e36. https://doi.org/10.1002/imt2.36 (2022).
Necchi, A. et al. Impact of molecular subtyping and immune infiltration on pathological response and outcome following neoadjuvant pembrolizumab in muscle-invasive bladder cancer. Eur. Urol. 77, 701–710. https://doi.org/10.1016/j.eururo.2020.02.028 (2020).
Wu, Q., Yu, X., Li, J., Sun, S. & Tu, Y. Metabolic regulation in the immune response to cancer. Cancer Commun. 41, 661–694. https://doi.org/10.1002/cac2.12182 (2021).
Tran, L., Xiao, J.-F., Agarwal, N., Duex, J. E. & Theodorescu, D. Advances in bladder cancer biology and therapy. Nat. Rev. Cancer 21, 104–121. https://doi.org/10.1038/s41568-020-00313-1 (2021).
Shin, K. et al. Hedgehog signaling restrains bladder cancer progression by eliciting stromal production of urothelial differentiation factors. Cancer Cell 26, 521–533. https://doi.org/10.1016/j.ccell.2014.09.001 (2014).
Thievessen, I., Wolter, M., Prior, A., Seifert, H.-H. & Schulz, W. A. Hedgehog signaling in normal urothelial cells and in urothelial carcinoma cell lines. J. Cell Physiol. 203, 372–377. https://doi.org/10.1002/jcp.20248 (2005).
Sverrisson, E. F. et al. Clinicopathological correlates of Gli1 expression in a population-based cohort of patients with newly diagnosed bladder cancer. Urol. Oncol. 32, 539–545. https://doi.org/10.1016/j.urolonc.2014.03.006 (2014).
Hahn, H. et al. Mutations of the human homolog of Drosophila patched in the nevoid basal cell carcinoma syndrome. Cell 85, 841–851. https://doi.org/10.1016/s0092-8674(00)81268-4 (1996).
Islam, S. S. et al. Sonic hedgehog (Shh) signaling promotes tumorigenicity and stemness via activation of epithelial-to-mesenchymal transition (EMT) in bladder cancer. Mol. Carcinog. 55, 537–551. https://doi.org/10.1002/mc.22300 (2016).
Yu, X. et al. Identification of UCP1 and UCP2 as potential prognostic markers in breast cancer: A study based on immunohistochemical analysis and bioinformatics. Front. Cell Dev. Biol. 10, 12 (2022).
Li, W., Guo, X., Chen, C. & Li, J. The prognostic value of arachidonic acid metabolism in breast cancer by integrated bioinformatics. Lipids Health Dis. 21, 103. https://doi.org/10.1186/s12944-022-01713-y (2022).
Liu, S.-Q. et al. Single-cell and spatially resolved analysis uncovers cell heterogeneity of breast cancer. J. Hematol. Oncol. 15, 19. https://doi.org/10.1186/s13045-022-01236-0 (2022).
McConkey, D. J., Choi, W., Ochoa, A. & Dinney, C. P. N. Intrinsic subtypes and bladder cancer metastasis. Asian J. Urol. 3, 260. https://doi.org/10.1016/j.ajur.2016.09.009 (2016).
Takebe, N. et al. Targeting Notch, Hedgehog, and Wnt pathways in cancer stem cells: Clinical update. Nat. Rev. Clin. Oncol. 12, 445–464. https://doi.org/10.1038/nrclinonc.2015.61 (2015).
Xiao, K. et al. Mesenchymal stem cells reverse EMT process through blocking the activation of NF-κB and hedgehog pathways in LPS-induced acute lung injury. Cell Death Dis. 11, 863. https://doi.org/10.1038/s41419-020-03034-3 (2020).
Chen, Y. et al. Hedgehog controls hepatic stellate cell fate by regulating metabolism. Gastroenterology 143, 1319-1329.e11. https://doi.org/10.1053/j.gastro.2012.07.115 (2012).
Lenis, A. T., Lec, P. M., Chamie, K. & Mshs, M. D. Bladder cancer: A review. JAMA 324, 1980–1991. https://doi.org/10.1001/jama.2020.17598 (2020).
Pham, C. D. et al. Differential immune microenvironments and response to immune checkpoint blockade amongst molecular subtypes of murine medulloblastoma. Clin. Cancer Res. 22, 582–595. https://doi.org/10.1158/1078-0432.CCR-15-0713 (2016).
Katoh, M. Network of WNT and other regulatory signaling cascades in pluripotent stem cells and cancer stem cells. Curr. Pharm. Biotechnol. 12, 160–170. https://doi.org/10.2174/138920111794295710 (2011).
Spranger, S. & Gajewski, T. F. A new paradigm for tumor immune escape: β-catenin-driven immune exclusion. J. Immunother. Cancer 3, 43. https://doi.org/10.1186/s40425-015-0089-6 (2015).
Fairfax, B. P. et al. Peripheral CD8+ T cell characteristics associated with durable responses to immune checkpoint blockade in patients with metastatic melanoma. Nat. Med. 26, 193–199. https://doi.org/10.1038/s41591-019-0734-6 (2020).
Uhlen, M. et al. A pathology atlas of the human cancer transcriptome. Science 357, eaan2507. https://doi.org/10.1126/science.aan2507 (2017).
Jiang, M., Wang, H., Chen, H. & Han, Y. SMARCD3 is a potential prognostic marker and therapeutic target in CAFs. Aging 12, 20835–20861. https://doi.org/10.18632/aging.104102 (2020).
Roldán, F. L. et al. Prognostic gene expression-based signature in clear-cell renal cell carcinoma. Cancers 14, 3754. https://doi.org/10.3390/cancers14153754 (2022).
Zuo, Y., Xu, X., Chen, M. & Qi, L. The oncogenic role of the cerebral endothelial cell adhesion molecule (CERCAM) in bladder cancer cells in vitro and in vivo. Cancer Med. 10, 4437–4450. https://doi.org/10.1002/cam4.3955 (2021).
Acknowledgements
The authors thank professional English editor (Bullet Edits) for assistance in improving the quality of language.
Funding
This work was supported by Shanghai Pujiang Program (22PJD054) and a National Natural Science Foundation of China (NSFC) grant (Grant NO: 82203629) to Dr. Qi Wu. This work was also suported by a Research Foudation of Shanghai Artemed Hospital (Grant NO: ATM2022YJ01) to Dr. Wenge Li.
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.X., Y.F. and Q.W. Funding acquisition: Y.X., Y.F. and Q.W. Investigation: X.Y., W.L. and Z.G. Project administration: Y.X. and Y.F. Supervision: Y.X. and Y.F. Validation: X.Y. and W.L. Original draft writing: X.Y. and W.L. Writing-review and editing: X.Y. and W.L. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yu, X., Li, W., Feng, Y. et al. The prognostic value of hedgehog signaling in bladder cancer by integrated bioinformatics. Sci Rep 13, 6241 (2023). https://doi.org/10.1038/s41598-023-33140-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-33140-z
This article is cited by
-
Investigating the prognostic value of mTORC1 signaling in bladder cancer via bioinformatics evaluation
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.