Abstract
The serine/threonine kinase Akt plays a central role in cell proliferation, survival and metabolism, and its hyperactivation is linked to cancer progression. Here we report that Akt undergoes K64 methylation by SETDB1, which is crucial for cell membrane recruitment, phosphorylation and activation of Akt following growth factor stimulation. Furthermore, we reveal an adaptor function of histone demethylase JMJD2A, which is important for recognizing Akt K64 methylation and recruits E3 ligase TRAF6 and Skp2-SCF to the Akt complex, independently of its demethylase activity, thereby initiating K63-linked ubiquitination, cell membrane recruitment and activation of Akt. Notably, the cancer-associated Akt mutant E17K displays enhanced K64 methylation, leading to its hyper-phosphorylation and activation. SETDB1-mediated Akt K64 methylation is upregulated and correlated with Akt hyperactivation in non-small-cell lung carcinoma (NSCLC), promotes tumour development and predicts poor outcome. Collectively, these findings reveal complicated layers of Akt activation regulation coordinated by SETDB1-mediated Akt K64 methylation to drive tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Mass spectrometry data have been deposited in ProteomeXchange with the primary accession code code PXD011966. The human lung adenocarcinoma data were derived from the TCGA Research Network: http://cancergenome.nih.gov/. The dataset derived from this resource that supports the findings of this study is available in http://www.cbioportal.org. The human cancer SETDB1 protein expression data were derived from the Human Protein Atlas Network and the supports the findings of this study is available in https://www.proteinatlas.org/ENSG00000143379-SETDB1/pathology. Source data for Figs. 4 and 7 and Supplementary Figs. 3 and 6 have been provided in Supplementary Table 4. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
Change history
29 April 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41556-021-00686-x
References
Cheung, M. & Testa, J. R. Diverse mechanisms of AKT pathway activation in human malignancy. Curr. Cancer Drug Targets 13, 234–244 (2013).
Manning, B. D. & Cantley, L. C. AKT/PKB signaling: navigating downstream. Cell 129, 1261–1274 (2007).
Yang, W. L. et al. The E3 ligase TRAF6 regulates Akt ubiquitination and activation. Science 325, 1134–1138 (2009).
Chan, C. H. et al. The Skp2-SCF E3 ligase regulates Akt ubiquitination, glycolysis, herceptin sensitivity, and tumorigenesis. Cell 149, 1098–1111 (2012).
Yang, W. L., Wu, C. Y., Wu, J. & Lin, H. K. Regulation of Akt signaling activation by ubiquitination. Cell Cycle 9, 487–497 (2010).
Hamamoto, R., Saloura, V. & Nakamura, Y. Critical roles of non-histone protein lysine methylation in human tumorigenesis. Nat. Rev. Cancer 15, 110–124 (2015).
Campaner, S. et al. The methyltransferase Set7/9 (Setd7) is dispensable for the p53-mediated DNA damage response in vivo. Mol. Cell 43, 681–688 (2011).
Kunizaki, M. et al. The lysine 831 of vascular endothelial growth factor receptor 1 is a novel target of methylation by SMYD3. Cancer Res. 67, 10759–10765 (2007).
Mazur, P. K. et al. SMYD3 links lysine methylation of MAP3K2 to Ras-driven cancer. Nature 510, 283–287 (2014).
Dasgupta, M., Dermawan, J. K., Willard, B. & Stark, G. R. STAT3-driven transcription depends upon the dimethylation of K49 by EZH2. Proc. Natl Acad. Sci. USA 112, 3985–3990 (2015).
Kim, E. et al. Phosphorylation of EZH2 activates STAT3 signaling via STAT3 methylation and promotes tumorigenicity of glioblastoma stem-like cells. Cancer Cell 23, 839–852 (2013).
Zhang, X. & Bruice, T. C. Enzymatic mechanism and product specificity of SET-domain protein lysine methyltransferases. Proc. Natl Acad. Sci. USA 105, 5728–5732 (2008).
Jacob, Y. et al. Regulation of heterochromatic DNA replication by histone H3 lysine 27 methyltransferases. Nature 466, 987–991 (2010).
Schultz, D. C. et al. SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev. 16, 919–932 (2002).
Fei, Q. et al. Histone methyltransferase SETDB1 regulates liver cancer cell growth through methylation of p53. Nat. Commun. 6, 8651 (2015).
Kim, M. S., Jeong, E. G., Yoo, N. J. & Lee, S. H. Mutational analysis of oncogenic AKT E17K mutation in common solid cancers and acute leukaemias. Br. J. Cancer 98, 1533–1535 (2008).
Bleeker, F. E. et al. AKT1(E17K) in human solid tumours. Oncogene 27, 5648–5650 (2008).
Carpten, J. D. et al. A transforming mutation in the pleckstrin homology domain of AKT1 in cancer. Nature 448, 439–444 (2007).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gyorffy, B., Surowiak, P., Budczies, J. & Lanczky, A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS ONE 8, e82241 (2013).
Uhlen, M. et al. A pathology atlas of the human cancer transcriptome. Science 357, eaan2507 (2017).
Berry, W. L. & Janknecht, R. KDM4/JMJD2 histone demethylases: epigenetic regulators in cancer cells. Cancer Res. 73, 2936–2942 (2013).
Cantley, L. C. The phosphoinositide 3-kinase pathway. Science 296, 1655–1657 (2002).
Datta, S. R., Brunet, A. & Greenberg, M. E. Cellular survival: a play in three Akts. Genes Dev. 13, 2905–2927 (1999).
Biggar, K. K. & Li, S. S. Non-histone protein methylation as a regulator of cellular signalling and function. Nat. Rev. Mol. Cell Biol. 16, 5–17 (2015).
Ea, C. K. & Baltimore, D. Regulation of NF-κB activity through lysine monomethylation of p65. Proc. Natl Acad. Sci. USA 106, 18972–18977 (2009).
Ceol, C. J. et al. The histone methyltransferase SETDB1 is recurrently amplified in melanoma and accelerates its onset. Nature 471, 513–517 (2011).
Rodriguez-Paredes, M. et al. Gene amplification of the histone methyltransferase SETDB1 contributes to human lung tumorigenesis. Oncogene 33, 2807–2813 (2014).
Sarraf, S. A. & Stancheva, I. Methyl-CpG binding protein MBD1 couples histone H3 methylation at lysine 9 by SETDB1 to DNA replication and chromatin assembly. Mol. Cell 15, 595–605 (2004).
Fritsch, L. et al. A subset of the histone H3 lysine 9 methyltransferases Suv39h1, G9a, GLP, and SETDB1 participate in a multimeric complex. Mol. Cell 37, 46–56 (2010).
Guo, X. D. J.et al. Methylation by SETDB1 promotes Akt kinase activity and oncogenic functions. Nat. Cell Biol. (2018).
Kim, T. D. et al. Histone demethylase JMJD2A drives prostate tumorigenesis through transcription factor ETV1. J. Clin. Invest. 126, 706–720 (2016).
Mallette, F. A. & Richard, S. JMJD2A promotes cellular transformation by blocking cellular senescence through transcriptional repression of the tumor suppressor CHD5. Cell Rep. 2, 1233–1243 (2012).
Berry, W. L., Shin, S., Lightfoot, S. A. & Janknecht, R. Oncogenic features of the JMJD2A histone demethylase in breast cancer. Int. J. Oncol. 41, 1701–1706 (2012).
Mallette, F. A. et al. RNF8- and RNF168-dependent degradation of KDM4A/JMJD2A triggers 53BP1 recruitment to DNA damage sites. EMBO J. 31, 1865–1878 (2012).
Cederquist, C. T. et al. Systemic insulin sensitivity is regulated by GPS2 inhibition of AKT ubiquitination and activation in adipose tissue. Mol. Metab. 6, 125–137 (2017).
Acknowledgements
We are grateful to the members of Lin’s lab for critical inputs and suggestions. We thank Z.-P. Liu, Y. Shinkai, F. J. Rauscher III, B. Vogelstein, D. Bohmann, M.C. Hung and R. Janknecht for providing mice, cell lines or plasmids. We thank E. Spooner for performing mass spectrometry assays. We acknowledge the support of Cellular Imaging & flow cytometry Shared of Resource, the Wake Forest Baptist Comprehensive Cancer Center, supported by the National Cancer Institute’s Cancer Center Support Grant (P30CA012197). This work is supported by start-ups from Wake Forest School of Medicine, NIH grants (R01CA182424 and R01CA193813) to H.K.L., NIH grants (RO1CA194094 and RO1CA197178) to H.L. and NIH grants (R01CA207098 and R01CA207109) to M.G.L.
Author information
Authors and Affiliations
Contributions
G.W. and H.K.L. designed the research and analysed the data. G.W. carried out the major experiments. J.L. performed the immunohistological characterization. J.L., Y.G., W.Z., F.H., C.X., L.S., Z. H. and Z.C. assisted with the experiments. S.C.Y., J.L., G.J., C.C.H., Y.H.W., and J.H. assisted with experimental design, T.Y.C. and M.G.L. assisted with in vitro methylation assay, G.W. and W.Z. developed the work models. M.G.L. and H.L. provided scientific inputs. G.W. and H.K.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 SETDB1 interacts with Akt, induces Akt K64 methylation and is required for Akt activation.
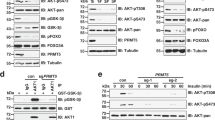
(a) Mass spectrometry analysis shows Akt binding proteins, including SETDB1 and JMJD2A. The mass spectrometry was performed once. (b) Setdb1± and Setdb1-/- MEFs were serum-starved, treated with EGF for 5 mins, and WCE were collected for IB analysis. (c) HEK293 cells transfected with Vector (Vec), Flag-SETDB1 (WT) and Flag-SETDB1 H1224K (HK) along with HA-Akt were serum-starved for 1 day, treated with 50 ng/ml IGF-1 for various times, and WCE were collected for IP with HA, followed by IB analysis indicated proteins. (d) HEK293 cells transfected with HA-Akt were serum-starved for 1 day, treated with 50 ng/ml IGF-1 for various times, and Whole cell extracts (WCE) were collected for IP with HA, followed by IB analysis indicated proteins. Data in b-d represent results from 3 independent experiments. (e) We immunoprecipitated endogenous Akt and performed mass spectrometry analysis, which reveals Akt methylation sites in normal condition, including K64, K112, K183, K268 and K400. The mass spectrometry was performed once. (f) We immunoprecipitated endogenous Akt in shLuc or shSETDB1 HEK293 cells and performed the Time-of-flight mass spectrometry (TOFMS) analysis, which identified Akt methylation sites that are different between Shluc and ShSETDB1 cells. TOFMS was performed once. Unprocessed blots are shown in Supplementary Figure 8.
Supplementary Figure 2 Akt K64 methylation promotes Akt membrane translocation.
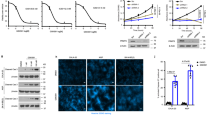
(a) Immunofluorescence (IF) assay showed the localization of indicated proteins in A549 cells treated with IGF-1 at 0 min or 10 mins. All IF representative data from 3–4 independent repeats. (b) HEK293 cells silenced with control (Shluc) or SETDB1 ShRNA (#1, #2) were serum-starved, treated with IGF-1 for 15 mins, and harvested for IF staining for indicated proteins. Scale bar represents 50 μm.All IF representative data from 3 independent repeats.
Supplementary Figure 3 SETDB1-mediated Akt k64 methylation regulates cancer development.
(a) Tumour weight of xenograft tumours derived from A549 cells silenced with control (Shluc) or SETDB1 ShRNA (#1 and #2), data shown represent mean ± s.d, n = 5 mice for each group, **** p < 0.0001, statistical significance was assessed using Student’s two-tailed t-test. (b) Tumour weight of xenograft tumours derived from A549 cells silenced with control (Shluc), SETDB1 ShRNA (#1) and SETDB1 ShRNA plus Myr-Akt overexpression, data shown represent mean ± s.d, n = 5 mice for each group, **** p < 0.0001, statistical significance was assessed using Student’s two-tailed t-test. (c) Soft agar transformation assay of 3T3 cells stable transfected with Vector (Vec), Akt1 (WT), Akt1 E17K (E17K) and Akt1 E17K/K64R (E17K/K64R). Crystal Violet staining before photograph. The average colony ( > 0,03 mm colony) number was quantitation (n = 3 independent experiments, Student’s two-tailed t-test, **** p < 0.0001). All Statistics source data are shown in Supplementary Table. 4.
Supplementary Figure 4 SETDB1 is overexpressed in cancers and correlates with overall survival (OS).
(a) cBioPortal databases examine SETDB1 gene amplification, mutation, deletion in multiple types of cancers. (b) Oncomine databases examine SETDB1 gene expression in multiple subtypes of lung cancers. (c) KM-Plot database reveals lung cancer patient overall OS with SETDB1 low or high expression. Affymetrix ID:214197-s-at. Total patients number is 1926 cases. (d) The overall survival of patients with higher and lower expressions of SETDB1 was marked in purple and blue, respectively. Overall survival in 429 Colorectal Adenocarcinoma Cancer patients (P = 2.13e-2), 541 Endometrial cancer patients (P = 2.24e-3), 501 Thyroid Cancer patients (P = 3.14e-2), 365 Liver Cancer patients (P = 8.88e-4), 877 Renal Cancer patients (P = 4.25e-4) and 102 Melanoma patients (P = 9.94e-3) was analysed, n means the sample number for each group. Log-rank P value for Kaplan-Meier plot showing results from analysis of correlation between mRNA expression level and patient survival. The P value obtained from log-rank test. The data-set that supports the findings of this study is available in http://www.cbioportal.org. The human cancer SETDB1 protein expression data were derived from the Human Protein Atlas Network and the supports the findings of this study is available in https://www.proteinatlas.org/ENSG00000143379-SETDB1/pathology.
Supplementary Figure 5 Akt K64 tri-methylation is critical for Akt binding to TRAF6 and Skp2.
(a) HEK293 cells were transfected with Vector (Vec), HA-Akt1 (WT) and HA-Akt1 K8R (K8R), and WCE were collected for IB analysis. (b-c) HEK293 cells were transfected with Vector (Vec), HA-Akt1 (WT) and HA-Akt1 K64R (K64R), and WCE were collected for IP with HA, followed by IB analysis. All IB data represent results from 3 independent experiments. Unprocessed blots are shown in Supplementary Figure 8.
Supplementary Figure 6 JMJD2A promotes the phosphorylation of Akt and tumorigenesis.
(a) JMJD2Alox/lox MEFs cells were infected with retrovirus control (MSCV1) or retrovirus Cre (Cre), serum-starved for 1 day, treated with 50 ng/ml IGF-1 for various times, and WCE were collected for IB analysis. (b) HEK293 cells were silenced with JMJD2A ShRNA (#1), transfected with pBabe Vector or pBabe-JMJD2A (pBabe-2A), and WCE were collected for IB analysis. (c) A549 cells silenced with control (Shluc) or JMJD2A ShRNA (#1 and #2) were were serum-starved for 1 day, treated with 50 ng/ml IGF-1 for 15 mins, and WCE were collected for IB analysis. Data in a-c represent results from 3 independent experiments. (d) Tumour volume of xenograft tumours derived from A549 cells silenced with control (Shluc), JMJD2A ShRNA (#1) and JMJD2A ShRNA (#1) plus Myr-Akt overexpression (data shown represent mean ± s.d, n = 6 or 7 mice for each group as indicated, **** p < 0.0001, statistical significance was assessed using Student’s two-tailed t-test.). (e) Tumour weight of xenograft tumours derived from A549 cells silenced with control (Shluc), JMJD2A ShRNA (#1) and JMJD2A ShRNA (#1) plus Myr-Akt overexpression, data shown represent mean ± s.d, n = 6 or 7 mice for each group as indicated, **** p < 0.0001, statistical significance was assessed using Student’s two-tailed t-test. Statistics source data for d and e are shown in Supplementary Table. 4. Unprocessed blots are shown in Supplementary Figure 8.
Supplementary Figure 7 PI 3-Kinase did not regulate Akt K64 methylation and Akt K140 and K142 methylation regulates Akt activation.
(a) HEK293 cells were transfected with Vector (Vec), HA-Akt1 (WT) and HA-Akt1 R25C (R25C), and WCE were collected for IB analysis. (b) HEK293 cells were treated with LY294002 in different concentration for 4 h and WCE were collected for IB analysis. (c) HEK293 cells were transfected with Vector (Vec), HA-Akt1 (WT), HA-Akt1 K140R (K140R), HA-Akt1 K142R (K142R) and HA-Akt1 K64R (K64R), and WCE were collected for IP with HA, followed by IB analysis. (d) HEK293 cells were transfected with Vector (Vec), HA-Akt1 (WT), HA-Akt1 K140/K142R (K140/142R) and HA-Akt1 K64R (K64R), and WCE were collected for IB analysis. Three independent experiments were performed with similar results for the IB. Unprocessed blots are shown in Supplementary Figure 8.
Supplementary Figure 8
Unprocessed blots.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and legends for Supplementary Tables 1–4.
Rights and permissions
About this article
Cite this article
Wang, G., Long, J., Gao, Y. et al. SETDB1-mediated methylation of Akt promotes its K63-linked ubiquitination and activation leading to tumorigenesis. Nat Cell Biol 21, 214–225 (2019). https://doi.org/10.1038/s41556-018-0266-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0266-1
This article is cited by
-
SETDB1 regulates short interspersed nuclear elements and chromatin loop organization in mouse neural precursor cells
Genome Biology (2024)
-
LINC00115 promotes chemoresistant breast cancer stem-like cell stemness and metastasis through SETDB1/PLK3/HIF1α signaling
Molecular Cancer (2024)
-
SETDB1 promotes progression through upregulation of SF3B4 expression and regulates the immunity in ovarian cancer
Journal of Ovarian Research (2024)
-
AKT1 phosphorylation of cytoplasmic ME2 induces a metabolic switch to glycolysis for tumorigenesis
Nature Communications (2024)
-
PDK1 neddylation by Smurf1 drives Akt activation
Cell Research (2024)