Key Points
-
Transcription is associated with elevated recombination and mutagenesis in bacteria and in eukaryotes and thereby alters the genetic landscape.
-
Concurrent transcription and replication of the same DNA template result in conflicts that lead to elevated chromosome fragility and recombination. In general, head-on collisions between the two machineries are more detrimental than co-directional collisions.
-
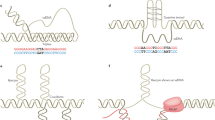
The formation of stable hybrids between the nascent RNA and its DNA template (R-loops) destabilizes the underlying template. Rapid engagement of the RNA discourages R-loop formation, but if formed, R-loops can be removed by RNase H or RNA–DNA helicases.
-
Transcription generates twin domains of positive and negative supercoiling, which are removed by topoisomerase 1 (Top1). The persistence of negative supercoils promotes R-loop formation.
-
Transcription facilitates the formation of non-B-DNA structures, which have been implicated in human trinucleotide repeat diseases.
-
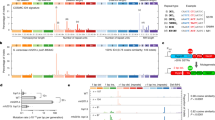
Active genes suffer more DNA damage than inactive genes in bacteria and yeast, and damage preferentially accumulates on the non-transcribed strand. When DNA repair mechanisms are intact, most transcription-associated mutations in yeast are due to the activity of Top1.
-
Transcription in yeast can alter the base composition of the underlying DNA template, and uracil specifically replaces thymine.
Abstract
Alterations in genome sequence and structure contribute to somatic disease, affect the fitness of subsequent generations and drive evolutionary processes. The crucial roles of highly accurate replication and efficient repair in maintaining overall genome integrity are well-known, but the more localized stability costs that are associated with transcribing DNA into RNA molecules are less appreciated. Here we review the diverse ways in which the essential process of transcription alters the underlying DNA template and thereby modifies the genetic landscape.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Savic, D. J. & Kanazir, D. T. The effect of a histidine operator-constitutive mutation on UV-induced mutability within the histidine operon of Salmonella typhimurium. Mol. Gen. Genet. 118, 45–50 (1972).
Herman, R. K. & Dworkin, N. B. Effect of gene induction on the rate of mutagenesis by ICR-191 in Escherichia coli. J. Bacteriol. 106, 543–550 (1971).
Datta, A. & Jinks-Robertson, S. Association of increased spontaneous mutation rates with high levels of transcription in yeast. Science 268, 1616–1619 (1995). This work, using budding yeast as a model system, was the first demonstration of transcription-associated mutagenesis in eukaryotic cells.
Saxowsky, T. T. & Doetsch, P. W. RNA polymerase encounters with DNA damage: transcription-coupled repair or transcriptional mutagenesis? Chem. Rev. 106, 474–488 (2006).
Keil, R. L. & Roeder, G. S. Cis-acting, recombination-stimulating activity in a fragment of the ribosomal DNA of S. cerevisiae. Cell 39, 377–386 (1984).
Voekel-Meiman, K., Keil, R. L. & Roeder, G. S. Recombination-stimulating sequences in yeast ribosomal DNA correspond to sequences regulating transcription by RNA polymerase I. Cell 48, 1071–1079 (1987). This paper identifies the recombination-stimulating sequence HOT1 as the rDNA promoter in budding yeast, thereby providing the first link between transcription and mitotic recombination.
Kim, N., Abdulovic, A. L., Gealy, R., Lippert, M. J. & Jinks-Robertson, S. Transcription-associated mutagenesis in yeast is directly proportional to the level of gene expression and influenced by the direction of DNA replication. DNA Repair 6, 1285–1296 (2007).
Reimers, J. M., Schmidt, K. H., Longacre, A., Reschke, D. K. & Wright, B. E. Increased transcription rates correlate with increased reversion rates in leuB and argH Escherichia coli auxotrophs. Microbiology 150, 1457–1466 (2004).
Nickoloff, J. A. Transcription enhances intrachromosomal homologous recombination in mammalian cells. Mol. Cell. Biol. 12, 5311–5318 (1992).
Postow, L. et al. Positive torsional strain causes the formation of a four-way junction at replication forks. J. Biol. Chem. 276, 2790–2796 (2001).
Rudolph, C. J., Dhillon, P., Moore, T. & Lloyd, R. G. Avoiding and resolving conflicts between DNA replication and transcription. DNA Repair 6, 981–993 (2007).
Pomerantz, R. T. & O'Donnell, M. What happens when replication and transcription complexes collide? Cell Cycle 9, 2537–2543 (2010).
Gottipati, P. & Helleday, T. Transcription-associated recombination in eukaryotes: link between transcription, replication and recombination. Mutagenesis 24, 203–210 (2009).
Ellwood, M. & Nomura, M. Chromosomal locations of the genes for rRNA in Escherichia coli K-12. J. Bacteriol. 149, 458–468 (1982).
Guy, L. & Roten, C. A. Genometric analyses of the organization of circular chromosomes: a universal pressure determines the direction of ribosomal RNA genes transcription relative to chromosome replication. Gene 340, 45–52 (2004).
Wang, J. D., Berkmen, M. B. & Grossman, A. D. Genome-wide coorientation of replication and transcription reduces adverse effects on replication in Bacillus subtilis. Proc. Natl Acad. Sci. USA 104, 5608–5613 (2007).
Srivatsan, A., Tehranchi, A., MacAlpine, D. M. & Wang, J. D. Co-orientation of replication and transcription preserves genome integrity. PLoS Genet. 6, e1000810 (2010).
Brewer, B. J. & Fangman, W. L. A replication fork barrier at the 3′ end of yeast ribosomal RNA genes. Cell 55, 637–643 (1988).
Huvet, M. et al. Human gene organization driven by the coordination of replication and transcription. Genome Res. 17, 1278–1285 (2007).
Mirkin, E. V. & Mirkin, S. M. Mechanisms of transcription-replication collisions in bacteria. Mol. Cell. Biol. 25, 888–895 (2005).
Deshpande, A. M. & Newlon, C. S. DNA replication fork pause sites dependent on transcription. Science 272, 1030–1033 (1996).
Prado, F. & Aguilera, A. Impairment of replication fork progression mediates RNA polII transcription-associated recombination. EMBO J. 24, 1267–1276 (2005).
Takeuchi, Y., Horiuchi, T. & Kobayashi, T. Transcription-dependent recombination and the role of fork collision in yeast rDNA. Genes Dev. 17, 1497–1506 (2003).
Azvolinsky, A., Giresi, P. G., Lieb, J. D. & Zakian, V. A. Highly transcribed RNA polymerase II genes are impediments to replication fork progression in Saccharomyces cerevisiae. Mol. Cell 34, 722–734 (2009). In this paper, the ChIP–chip technique was used to identify sites of DNA Pol II occupancy, which are presumed to reflect replication pausing, throughout the budding yeast genome. Highly transcribed ORFs were enriched among these sites, indicating that transcription can impede replication.
Cox, M. M. et al. The importance of repairing stalled replication forks. Nature 404, 37–41 (2000).
Gottipati, P., Cassel, T. N., Savolainen, L. & Helleday, T. Transcription-associated recombination is dependent on replication in mammalian cells. Mol. Cell. Biol. 28, 154–164 (2008).
de la Loza, M. C., Wellinger, R. E. & Aguilera, A. Stimulation of direct-repeat recombination by RNA polymerase III transcription. DNA Repair 8, 620–626 (2009).
Sikdar, N., Banerjee, S., Zhang, H., Smith, S. & Myung, K. Spt2p defines a new transcription-dependent gross chromosomal rearrangement pathway. PLoS Genet. 4, e1000290 (2008).
Vilette, D., Ehrlich, S. D. & Michel, B. Transcription-induced deletions in Escherichia coli plasmids. Mol. Microbiol. 17, 493–504 (1995).
Dutta, D., Shatalin, K., Epshtein, V., Gottesman, M. E. & Nudler, E. Linking RNA polymerase backtracking to genome instability in E. coli. Cell 146, 533–543 (2011).
Blobel, G. Gene gating: a hypothesis. Proc. Natl Acad. Sci. USA 82, 8527–8529 (1985).
Bermejo, R. et al. The replication checkpoint protects fork stability by releasing transcribed genes from nuclear pores. Cell 146, 233–246 (2011).
Waters, L. S. et al. Eukaryotic translesion polymerases and their roles and regulation in DNA damage tolerance. Microbiol. Mol. Biol. Rev. 73, 134–154 (2009).
Aguilera, A. The connection between transcription and genomic instability. EMBO J. 21, 195–201 (2002).
Li, X. & Manley, J. L. Cotranscriptional processes and their influence on genome stability. Genes Dev. 20, 1838–1847 (2006).
Gowrishankar, J. & Harinarayanan, R. Why is transcription coupled to translation in bacteria? Mol. Microbiol. 54, 598–603 (2004).
Wu, H. Y., Shyy, S. H., Wang, J. C. & Liu, L. F. Transcription generates positively and negatively supercoiled domains in the template. Cell 53, 433–440 (1988). This paper proposed the 'twin domain' model of transcription-associated DNA helical stress.
Tuduri, S. et al. Topoisomerase I suppresses genomic instability by preventing interference between replication and transcription. Nature Cell Biol. 11, 1315–1324 (2009).
Cerritelli, S. M. & Crouch, R. J. Ribonuclease H: the enzymes in eukaryotes. FEBS J. 276, 1494–1505 (2009).
Baaklini, I., Hraiky, C., Rallu, F., Tse-Dinh, Y. C. & Drolet, M. RNase HI overproduction is required for efficient full-length RNA synthesis in the absence of topoisomerase I in Escherichia coli. Mol. Microbiol. 54, 198–211 (2004).
Mischo, H. E. et al. Yeast Sen1 helicase protects the genome from transcription-associated instability. Mol. Cell 41, 21–32 (2011).
Rudolph, C. J., Upton, A. L., Briggs, G. S. & Lloyd, R. G. Is RecG a general guardian of the bacterial genome? DNA Repair 9, 210–223 (2010).
Skourti-Stathaki, K., Proudfoot, N. J. & Gromak, N. Human senataxin resolves RNA/DNA hybrids formed at transcriptional pause sites to promote Xrn2-dependent termination. Mol. Cell 42, 794–805 (2011).
Chavez, S. & Aguilera, A. The yeast HPR1 gene has a functional role in transcriptional elongation that uncovers a novel source of genome instability. Genes Dev. 11, 3459–3470 (1997).
Huertas, P. & Aguilera, A. Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol. Cell 12, 711–721 (2003). This paper highlights the role of co-transcriptionally formed RNA–DNA hybrids (R-loops) in TAR by showing that degradation of the nascent mRNA by RNase H1 or a self-cleaving hammerhead ribozyme can suppress hyper-recombination in budding yeast hpr1 mutants.
Rondon, A. G., Jimeno, S. & Aguilera, A. The interface between transcription and mRNP export: from THO to THSC/TREX-2. Biochim. Biophys. Acta 1799, 533–538 (2010).
Gomez-Gonzalez, B. et al. Genome-wide function of THO/TREX in active genes prevents R-loop-dependent replication obstacles. EMBO J. 30, 3106–3119 (2011).
Wahba, L., Amon, J. D., Koshland, D. & Vuica-Ross, M. RNase H and multiple RNA biogenesis factors cooperate to prevent RNA:DNA hybrids from generating genome instability. Mol. Cell 44, 978–988 (2011).
French, S. L. et al. Distinguishing the roles of Topoisomerases I and II in relief of transcription-induced torsional stress in yeast rRNA genes. Mol. Cell. Biol. 31, 482–494 (2011).
El Hage, A., French, S. L., Beyer, A. L. & Tollervey, D. Loss of Topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev. 24, 1546–1558 (2010).
Christman, M. F., Dietrich, F. S. & Fink, G. R. Mitotic recombination in the rDNA of S. cerevisiae is suppressed by the combined action of DNA topoisomerases I and II. Cell 55, 413–425 (1988).
Li, X. & Manley, J. L. Inactivation of the SR protein splicing factor ASF/SF2 results in genomic instability. Cell 122, 365–378 (2005). This paper demonstrated the role of mRNA splicing complex assembly in suppressing TAR in chicken DT40 and mammalian cells.
Paulsen, R. D. et al. A genome-wide siRNA screen reveals diverse cellular processes and pathways that mediate genome stability. Mol. Cell 35, 228–239 (2009).
Dominguez-Sanchez, M. S., Barroso, S., Gomez-Gonzalez, B., Luna, R. & Aguilera, A. Genome instability and transcription elongation impairment in human cells depleted of THO/TREX. PLoS Genet. 7, e1002386 (2011).
Helmrich, A., Ballarino, M. & Tora, L. Collisions between replication and transcription complexes cause common fragile site instability at the longest human genes. Mol. Cell 44, 966–977 (2011).
Yu, K., Chedin, F., Hsieh, C. L., Wilson, T. E. & Lieber, M. R. R-loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells. Nature Immunol. 4, 442–451 (2003).
Maizels, N. Dynamic roles for G4 DNA in the biology of eukaryotic cells. Nature Struct. Mol. Biol. 13, 1055–1059 (2006).
Duquette, M. L., Handa, P., Vincent, J. A., Taylor, A. F. & Maizels, N. Intracellular transcription of G-rich DNAs induces formation of G-loops, novel structures containing G4 DNA. Genes Dev. 18, 1618–1629 (2004). This work demonstrated that a highly transcribed immunoglobulin switch region sequence forms G-loop structures visible by electron microscope both in vitro and in cells in E. coli.
Belotserkovskii, B. P. et al. Mechanisms and implications of transcription blockage by guanine-rich DNA sequences. Proc. Natl Acad. Sci. USA 107, 12816–12821 (2010).
Shinkura, R. et al. The influence of transcriptional orientation on endogenous switch region function. Nature Immunol. 4, 435–441 (2003).
Kim, N. & Jinks-Robertson, S. Guanine repeat-containing sequences confer transcription-dependent instability in an orientation-specific manner in yeast. DNA Repair 10, 953–960 (2011).
Gan, W. et al. R-loop-mediated genomic instability is caused by impairment of replication fork progression. Genes Dev. 25, 2041–2056 (2011).
Lopes, J. et al. G-quadruplex-induced instability during leading-strand replication. EMBO J. 30, 4033–4046 (2011).
Paeschke, K., Capra, J. A. & Zakian, V. A. DNA replication through G-quadruplex motifs is promoted by the Saccharomyces cerevisiae Pif1 DNA helicase. Cell 145, 678–691 (2011).
Mirkin, S. M. Expandable DNA repeats and human disease. Nature 447, 932–940 (2007).
McMurray, C. T. DNA secondary structure: a common and causative factor for expansion in human disease. Proc. Natl Acad. Sci. USA 96, 1823–1825 (1999).
Swami, M. et al. Somatic expansion of the Huntington's disease CAG repeat in the brain is associated with an earlier age of disease onset. Hum. Mol. Genet. 18, 3039–3047 (2009).
Petruska, J., Arnheim, N. & Goodman, M. F. Stability of intrastrand hairpin structures formed by the CAG/CTG class of DNA triplet repeats associated with neurological diseases. Nucleic Acids Res. 24, 1992–1998 (1996).
Salinas-Rios, V., Belotserkovskii, B. P. & Hanawalt, P. C. DNA slip-outs cause RNA polymerase II arrest in vitro: potential implications for genetic instability. Nucleic Acids Res. 39, 7444–7454 (2011).
Lin, Y., Dion, V. & Wilson, J. H. Transcription promotes contraction of CAG repeat tracts in human cells. Nature Struct. Mol. Biol. 13, 179–180 (2006). This work showed that contraction of CAG triplet repeats in human cells requires transcription and components of defined DNA repair pathways.
Lin, Y. & Wilson, J. H. Transcription-induced CAG repeat contraction in human cells is mediated in part by transcription-coupled nucleotide excision repair. Mol. Cell. Biol. 27, 6209–6217 (2007).
Jung, J. & Bonini, N. CREB-binding protein modulates repeat instability in a Drosophila model for polyQ disease. Science 315, 1857–1859 (2007). Using a D. melanogaster model system, this work demonstrated that CAG repeat expansion requires transcription and is mediated by TC-NER.
Wells, R. D., Dere, R., Hebert, M. L., Napierala, M. & Son, L. S. Advances in mechanisms of genetic instability related to hereditary neurological diseases. Nucleic Acids Res. 33, 3785–3798 (2005).
Grabczyk, E., Mancuso, M. & Sammarco, M. C. A persistent RNA•DNA hybrid formed by transcription of the Friedreich ataxia triplet repeat in live bacteria, and by T7 RNAP in vitro. Nucleic Acids Res. 35, 5351–5359 (2007).
Ditch, S., Sammarco, M. C., Banerjee, A. & Grabczyk, E. Progressive GAA•TTC repeat expansion in human cell lines. PLoS Genet. 5, e1000704 (2009).
Rindler, P. M. & Bidichandani, S. I. Role of transcript and interplay between transcription and replication in triplet-repeat instability in mammalian cells. Nucleic Acids Res. 39, 526–535 (2011).
Yelin, R. et al. Widespread occurrence of antisense transcription in the human genome. Nature Biotech. 21, 379–386 (2003).
Lin, Y. & Wilson, J. H. Transcription-induced DNA toxicity at trinucleotide repeats: double bubble is trouble. Cell Cycle 10, 611–618 (2011).
Lin, Y., Leng, M., Wan, M. & Wilson, J. H. Convergent transcription through a long CAG tract destabilizes repeats and induces apoptosis. Mol. Cell. Biol. 30, 4435–4451 (2010).
Nakamori, M., Pearson, C. E. & Thornton, C. A. Bidirectional transcription stimulates expansion and contraction of expanded (CTG)•(CAG) repeats. Hum. Mol. Genet. 20, 580–588 (2011).
Morey, N. J., Greene, C. N. & Jinks-Robertson, S. Genetic analysis of transcription-associated mutation in Saccharomyces cerevisiae. Genetics 154, 109–120 (2000).
Garcia-Rubio, M., Huertas, P., Gonzalez-Barrera, S. & Aguilera, A. Recombinogenic effects of DNA-damaging agents are synergistically increased by transcription in Saccharomyces cerevisiae. New insights into transcription-associated recombination. Genetics 165, 457–466 (2003).
Lippert, M. J. et al. Role for topoisomerase 1 in transcription-associated mutagenesis in yeast. Proc. Natl Acad. Sci. USA 108, 698–703 (2011).
Takahashi, T., Burguiere-Slezak, G., Van der Kemp, P. A. & Boiteux, S. Topoisomerase 1 provokes the formation of short deletions in repeated sequences upon high transcription in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 108, 692–697 (2011). The above two references describe transcription-dependent short deletions at tandem repeats and demonstrate that these events require Top1 activity.
Kim, N. & Jinks-Robertson, S. Abasic sites in the transcribed strand of yeast DNA are removed by transcription-coupled nucleotide excision repair. Mol. Cell. Biol. 30, 3206–3215 (2010). This paper showed that uracil-derived AP sites and the ensuing mutagenesis are highly stimulated by transcription in budding yeast. The uracil reflects direct incorporation of dUTP in place of dTTP rather than cytosine deamination.
Hudson, R. E., Bergthorsson, U. & Ochman, H. Transcription increases multiple spontaneous point mutations in Salmonella enterica. Nucleic Acids Res. 31, 4517–4522 (2003).
Klapacz, J. & Bhagwat, A. S. Transcription-dependent increase in multiple classes of base substitution mutations in Escherichia coli. J. Bacteriol. 184, 6866–6872 (2002).
Beletskii, A. & Bhagwat, A. S. Transcription-induced mutations: increase in C to T mutations in the nontranscribed strand during transcription in Escherichia coli. Proc. Natl Acad. Sci. USA 93, 13919–13924 (1996). This work demonstrated that in E. coli , C to T mutations resulting from spontaneous deamination of cytosines occur more frequently on the NTS of an active gene.
Beletskii, A. & Bhagwat, A. S. Correlation between transcription and C to T mutations in the non-transcribed DNA strand. Biol. Chem. 379, 549–551 (1998).
Klapacz, J. & Bhagwat, A. S. Transcription promotes guanine to thymine mutations in the non-transcribed strand of an Escherichia coli gene. DNA Repair 4, 806–813 (2005).
Frederico, L. A., Kunkel, T. A. & Shaw, B. R. A sensitive genetic assay for the detection of cytosine deamination: determination of rate constants and the activation energy. Biochemistry 29, 2532–2537 (1990).
Gomez-Gonzalez, B. & Aguilera, A. Activation-induced cytidine deaminase action is strongly stimulated by mutations of the THO complex. Proc. Natl Acad. Sci. USA 104, 8409–8414 (2007).
Green, P., Ewing, B., Miller, W., Thomas, P. J. & Green, E. D. Transcription-associated mutational asymmetry in mammalian evolution. Nature Genet. 33, 514–517 (2003).
Majewski, J. Dependence of mutational asymmetry on gene-expression levels in the human genome. Am. J. Hum. Genet. 73, 688–692 (2003). The above two papers together demonstrated a strand-specific mutation bias in mammalian genes that correlates with expression level.
Rubin, A. F. & Green, P. Mutation patterns in cancer genomes. Proc. Natl Acad. Sci. USA 106, 21766–21770 (2009).
Wright, B. E., Reschke, D. K., Schmidt, K. H., Reimers, J. M. & Knight, W. Predicting mutation frequencies in stem-loop structures of derepressed genes: implications for evolution. Mol. Microbiol. 48, 429–441 (2003).
Schmidt, K. H., Reimers, J. M. & Wright, B. E. The effect of promoter strength, supercoiling and secondary structure on mutation rates in Escherichia coli. Mol. Microbiol. 60, 1251–1261 (2006).
Wright, B. E. et al. The roles of transcription and genotoxins underlying p53 mutagenesis in vivo. Carcinogenesis 32, 1559–1567 (2011).
Hanawalt, P. C. & Spivak, G. Transcription-coupled DNA repair: two decades of progress and surprises. Nature Rev. Mol. Cell Biol. 9, 958–970 (2008).
Iannone, R. et al. Mutation spectra analysis suggests that N-(2-chloroethyl)-N′-cyclohexyl-N-nitrosourea-induced lesions are subject to transcription-coupled repair in Escherichia coli. Mol. Carcinog. 19, 39–45 (1997).
Li, B. H., Ebbert, A. & Bockrath, R. Transcription-modulated repair in Escherichia coli evident with UV-induced mutation spectra in supF. J. Mol. Biol. 294, 35–48 (1999).
Moriya, M. et al. TP53 Mutational signature for aristolochic acid: an environmental carcinogen. Int. J. Cancer 129, 1532–1536 (2011).
Hendriks, G. et al. Gene transcription increases DNA damage-induced mutagenesis in mammalian stem cells. DNA Repair 7, 1330–1339 (2008).
Hendriks, G. et al. Transcription-dependent cytosine deamination is a novel mechanism in ultraviolet light-induced mutagenesis. Curr. Biol. 20, 170–175 (2010).
Kim, N. et al. Mutagenic processing of ribonucleotides in DNA by yeast topoisomerase I. Science 332, 1561–1564 (2011).
Kim, N. & Jinks-Robertson, S. dUTP incorporation into genomic DNA is linked to transcription in yeast. Nature 459, 1150–1153 (2009).
Neuberger, M. S. et al. Somatic hypermutation at A.T pairs: polymerase error versus dUTP incorporation. Nature Rev. Immunol. 5, 171–178 (2005).
Roche, B., Claes, A. & Rougeon, F. Deoxyuridine triphosphate incorporation during somatic hypermutation of mouse VkOx genes after immunization with phenyloxazolone. J. Immunol. 185, 4777–4782 (2010).
Maul, R. W. et al. Uracil residues dependent on the deaminase AID in immunoglobulin gene variable and switch regions. Nature Immunol. 12, 70–76 (2011).
Cho, R. J. et al. A genome-wide transcriptional analysis of the mitotic cell cycle. Mol. Cell 2, 65–73 (1998).
Pybus, C. et al. Transcription-associated mutation in Bacillus subtilis cells under stress. J. Bacteriol. 192, 3321–3328 (2010).
Maldonado, E. et al. A human RNA polymerase II complex associated with SRB and DNA-repair proteins. Nature 381, 86–89 (1996).
Deem, A. et al. Break-induced replication is highly inaccurate. PLoS Biol. 9, e1000594 (2011).
Hicks, W. M., Kim, M. & Haber, J. E. Increased mutagenesis and unique mutation signature associated with mitotic gene conversion. Science 329, 82–85 (2010).
Strathern, J. N., Shafer, B. & McGill, C. B. DNA synthesis errors associated with double-strand-break repair. Genetics 140, 965–972 (1995).
Gonzalez-Barrera, S., Garcia-Rubio, M. & Aguilera, A. Transcription and double-strand breaks induce similar mitotic recombination events in Saccharomyces cerevisiae. Genetics 162, 603–614 (2002).
Saxe, D., Datta, A. & Jinks-Robertson, S. Stimulation of mitotic recombination events by high levels of RNA polymerase II transcription in yeast. Mol. Cell. Biol. 20, 5404–5414 (2000).
Schildkraut, E., Miller, C. A. & Nickoloff, J. A. Transcription of a donor enhances its use during double-strand break-induced gene conversion in human cells. Mol. Cell. Biol. 26, 3098–3105 (2006).
Derr, L. K. & Strathern, J. N. A role for reverse transcripts in gene conversion. Nature 361, 170–173 (1993).
Fink, G. R. Pseudogenes in yeast? Cell 49, 5–6 (1987).
Storici, F., Bebenek, K., Kunkel, T. A., Gordenin, D. A. & Resnick, M. A. RNA-templated DNA repair. Nature 447, 338–341 (2007).
Sekiguchi, J. & Shuman, S. Site-specific ribonuclease activity of eukaryotic DNA topoisomerase I. Mol. Cell 1, 89–97 (1997).
Henningfeld, K. A. & Hecht, S. M. A model for topoisomerase I-mediated insertions and deletions with duplex DNA substrates containing branches, nicks, and gaps. Biochemistry 34, 6120–6129 (1995).
Maizels, N. Immunoglobulin gene diversity. Annu. Rev. Genet. 39, 23–46 (2005).
Chiarle, R. et al. Genome-wide translocation sequencing reveals mechanisms of chromosome breaks and rearrangements in B cells. Cell 147, 107–119 (2011).
Acknowledgements
This work has been supported by grants from the US National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Non-transcribed strand
-
(NTS). The NTS is complementary to the transcribed strand and has the same sequence as the RNA (except that it contains thymine instead of uracil); it is often referred to as the coding strand, whose sequence is given as standard.
- Replisome
-
The multi-protein complex that contains all of the proteins that are required for DNA replication. This includes the DNA polymerases, factors that increase the processivity of DNA synthesis and a helicase to unwind duplex DNA.
- Two-dimensional gels
-
These are used to visualize replication fork progression across a defined segment of DNA. DNA is separated by size in the first dimension and by shape in the second; the fragment of interest is visualized by Southern blot analysis. Linear fragments run on a diagonal; fragments that run off the diagonal correspond to replicating or branched molecules.
- Chromatin immunoprecipitation followed by microarray
-
(ChIP–chip). DNA that interacts with a given protein is immunoprecipitated from cell extracts ('ChIP'). The precipitated DNA is labelled and hybridized to a microarray ('chip'), where signals above background reflect sequences preferentially immunoprecipitated with the protein of interest; it is used to map locations of protein–DNA interactions.
- THO complex
-
A conserved protein complex that includes the proteins Tho2, Hpr1, Mft1 and Thp2 in yeast. It interacts with the TREX complex and functions in mRNA metabolism and export.
- Transcription-coupled nucleotide excision repair
-
(TC-NER). TC-NER is a subpathway of the NER pathway that is initiated specifically in response to an RNA polymerase arrested by damage on the DNA template. The net effect is more efficient NER-directed repair of lesions on the transcribed than on the non-transcribed strand of active genes.
- Reversion assays
-
Assays that start with a mutant allele, typically containing a change in a single base pair or the insertion or deletion of a single base pair, and that select for restoration of gene function. The change that restores gene function is usually limited to the position of the original mutation.
- Forward mutations
-
Forward mutation assays select for loss of a gene function and can detect any change in the DNA sequence that inactivates the encoded product, which is usually a protein.
Rights and permissions
About this article
Cite this article
Kim, N., Jinks-Robertson, S. Transcription as a source of genome instability. Nat Rev Genet 13, 204–214 (2012). https://doi.org/10.1038/nrg3152
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3152
This article is cited by
-
Plastid-encoded RNA polymerase variation in Pelargonium sect Ciconium
Horticulture Advances (2024)
-
The testosterone paradox of advanced prostate cancer: mechanistic insights and clinical implications
Nature Reviews Urology (2023)
-
RNase H2, mutated in Aicardi‐Goutières syndrome, resolves co-transcriptional R-loops to prevent DNA breaks and inflammation
Nature Communications (2022)
-
Recent advances in solid-state beyond lithium batteries
Journal of Solid State Electrochemistry (2022)
-
The genome-wide impact of trisomy 21 on DNA methylation and its implications for hematopoiesis
Nature Communications (2021)