Abstract
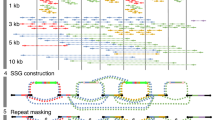
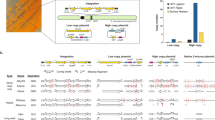
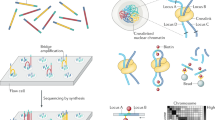
Long-read sequencing technologies have become increasingly popular due to their strengths in resolving complex genomic regions. As a leading model organism with small genome size and great biotechnological importance, the budding yeast Saccharomyces cerevisiae has many isolates currently being sequenced with long reads. However, analyzing long-read sequencing data to produce high-quality genome assembly and annotation remains challenging. Here, we present a modular computational framework named long-read sequencing data analysis for yeasts (LRSDAY), the first one-stop solution that streamlines this process. Starting from the raw sequencing reads, LRSDAY can produce chromosome-level genome assembly and comprehensive genome annotation in a highly automated manner with minimal manual intervention, which is not possible using any alternative tool available to date. The annotated genomic features include centromeres, protein-coding genes, tRNAs, transposable elements (TEs), and telomere-associated elements. Although tailored for S. cerevisiae, we designed LRSDAY to be highly modular and customizable, making it adaptable to virtually any eukaryotic organism. When applying LRSDAY to an S. cerevisiae strain, it takes ∼41 h to generate a complete and well-annotated genome from ∼100× Pacific Biosciences (PacBio) running the basic workflow with four threads. Basic experience working within the Linux command-line environment is recommended for carrying out the analysis using LRSDAY.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Goffeau, A. et al. Life with 6000 genes. Science 274, 546–567 (1996).
Gordon, D. et al. Long-read sequence assembly of the gorilla genome. Science 352, aae0344 (2016).
VanBuren, R. et al. Single-molecule sequencing of the desiccation-tolerant grass Oropetium thomaeum. Nature 527, 508–11 (2015).
Bickhart, D.M. et al. Single-molecule sequencing and chromatin conformation capture enable de novo reference assembly of the domestic goat genome. Nat. Genet. 49, 643–650 (2017).
Badouin, H. et al. The sunflower genome provides insights into oil metabolism, flowering and asterid evolution. Nature 546, 148–152 (2017).
Jain, M. et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat. Biotechnol. 36, 338–345 (2018).
Yue, J.-X. et al. Contrasting evolutionary genome dynamics between domesticated and wild yeasts. Nat. Genet. 49, 913–924 (2017).
Goodwin, S. et al. Oxford nanopore sequencing, hybrid error correction, and de novo assembly of a eukaryotic genome. Genome Res. 25, 1750–1756 (2015).
McIlwain, S.J. et al. Genome sequence and analysis of a stress-tolerant, wild-derived strain of Saccharomyces cerevisiae used in biofuels research. G3 (Bethesda) 6, 1757–66 (2016).
Istace, B. et al. De novo assembly and population genomic survey of natural yeast isolates with the Oxford Nanopore MinION sequencer. Gigascience 6, 1–13 (2017).
Giordano, F. et al. De novo yeast genome assemblies from MinION, PacBio and MiSeq platforms. Sci. Rep. 7, 3935 (2017).
Koren, S. et al. Hybrid error correction and de novo assembly of single-molecule sequencing reads. Nat. Biotechnol. 30, 693–700 (2012).
Chin, C.-S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–9 (2013).
Berlin, K. et al. Assembling large genomes with single-molecule sequencing and locality-sensitive hashing. Nat. Biotechnol. 33, 623–630 (2015).
Chin, C.-S. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods 13, 1050–1054 (2016).
Li, H. Minimap and miniasm: fast mapping and de novo assembly for noisy long sequences. Bioinformatics 32, 2103–2110 (2016).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive κ-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Salamov, A.A. & Solovyev, V.V. Ab initio gene finding in Drosophila genomic DNA. Genome Res. 10, 516–522 (2000).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–W312 (2004).
Besemer, J. & Borodovsky, M. GeneMark: web software for gene finding in prokaryotes, eukaryotes and viruses. Nucleic Acids Res. 33, W451–W454 (2005).
Proux-Wéra, E., Armisén, D., Byrne, K.P. & Wolfe, K.H. A pipeline for automated annotation of yeast genome sequences by a conserved-synteny approach. BMC Bioinformatics 13, 237 (2012).
Strope, P.K. et al. The 100-genomes strains, an S. cerevisiae resource that illuminates its natural phenotypic and genotypic variation and emergence as an opportunistic pathogen. Genome Res. 25, 762–774 (2015).
Almeida, P. et al. A population genomics insight into the Mediterranean origins of wine yeast domestication. Mol. Ecol. 24, 5412–5427 (2015).
Gallone, B. et al. Domestication and divergence of Saccharomyces cerevisiae beer yeasts. Cell 166, 1397–1410.e16 (2016).
Peter, J. et al. Genome evolution across 1,011 Saccharomyces cerevisiae isolates. Nature. http://dx.doi.org/10.1038/s41586-018-0030-5 (2018).
Bergström, A. et al. A high-definition view of functional genetic variation from natural yeast genomes. Mol. Biol. Evol. 31, 872–888 (2014).
Drillon, G., Carbone, A. & Fischer, G. SynChro: a fast and easy tool to reconstruct and visualize synteny blocks along eukaryotic chromosomes. PLoS One 9 (2014).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40 (2012).
Louis, E.J. & Haber, J.E. The structure and evolution of subtelomeric Y repeats in Saccharomyces cerevisiae. Genetics 131, 559–574 (1992).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Smit, A., Hubley, R. & Green, P. RepeatMasker Open-4.0. 2013-2015 http://www.repeatmasker.org (2013).
Kolmogorov, M., Raney, B., Paten, B. & Pham, S. Ragout - a reference-assisted assembly tool for bacterial genomes. Bioinformatics 30, i302–i309 (2014).
Robinson, J.T. et al. Integrative Genomics Viewer. Nat. Biotechnol. 29, 24–26 (2011).
Pereira, V. Automated paleontology of repetitive DNA with REANNOTATE. BMC Genomics 9, 614 (2008).
Walker, B.J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9 (2014).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12, 491 (2011).
Haas, B.J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the program to assemble spliced alignments. Genome Biol. 9, R7 (2008).
Lowe, T.M. & Eddy, S.R. A computational screen for methylation guide snoRNAs in yeast. Science 283, 1168–71 (1999).
Minkin, I., Patel, A., Kolmogorov, M., Vyahhi, N. & Pham, S. Sibelia: a scalable and comprehensive synteny block generation tool for closely related microbial genomes. in Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) 8126 LNBI, 215–229 (2013).
Acknowledgements
We thank the developers of the MAKER software for allowing us to incorporate MAKER into the LRSDAY auto-installation process. We thank G. Fischer, S. O'Donnell, and L. Tattini for testing LRSDAY and providing valuable feedback. This work was supported by ATIP-Avenir (CNRS/INSERM), Fondation ARC pour la Recherche sur le Cancer (PJA20151203273), Marie Curie Career Integration Grants (322035), Agence Nationale de la Recherche (ANR-16-CE12-0019, ANR-13-BSV6-0006-01, and ANR-11-LABX-0028-01), Cancéropôle PACA (AAP émergence 2015), and a DuPont Young Professor Award to G.L. J.-X.Y. was supported by a postdoctoral fellowship from Fondation ARC pour la Recherche sur le Cancer (PDF20150602803).
Author information
Authors and Affiliations
Contributions
J.-X.Y. designed, implemented, and tested the LRSDAY workflow. G.L. coordinated the work. J.-X.Y. and G.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Yue, JX., Liti, G. Long-read sequencing data analysis for yeasts. Nat Protoc 13, 1213–1231 (2018). https://doi.org/10.1038/nprot.2018.025
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2018.025
This article is cited by
-
Convergent genomic signatures associated with vertebrate viviparity
BMC Biology (2024)
-
Unlocking the functional potential of polyploid yeasts
Nature Communications (2022)
-
Aborting meiosis allows recombination in sterile diploid yeast hybrids
Nature Communications (2021)
-
A 19-isolate reference-quality global pangenome for the fungal wheat pathogen Zymoseptoria tritici
BMC Biology (2020)
-
A yeast living ancestor reveals the origin of genomic introgressions
Nature (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.