Abstract
MicroRNAs are short non-coding molecules that play a major role in regulating gene expression. Peripheral levels of miR-1202 have been shown to predict and mediate antidepressant response. However, it is not clear to what extent these peripheral measures reflect central neural changes in vivo. We approached this problem with the combined use of peripheral miR-1202 measures and neuroimaging. At baseline and after 8 weeks of desvenlafaxine (50–100 mg die), 20 patients were scanned with 3T magnetic resonance imaging, first at rest then during the Go/NoGo task, a classical test of response inhibition. Blood samples were collected at both time points. During resting state, lower baseline miR-1202 levels were predictive of increased connectivity from T0 to T8 between the posterior cingulate and the prefrontal, parietal, and occipital cortices. Changes in miR-1202 levels following desvenlafaxine treatment were negatively correlated with changes in activity in right precuneus within the default-mode network, and in connectivity between the posterior cingulate and the temporal and prefrontal cortices, and the precuneus. During the Go/NoGo task, baseline miR-1202 levels and changes in these levels were correlated with activity changes in different regions, including bilateral prefrontal, insular, cingulate, and temporal cortices, and left putamen and claustrum. Finally, secondary analyses in a subset of patients showed a trend for a significant correlation between miR-1202 levels and glutamate levels measured by spectroscopy. Changes in peripheral miR-1202 levels were therefore associated with changes in brain activity and connectivity in a network of brain regions associated with depression and antidepressant response. These effects may be mediated by the glutamatergic system.
Similar content being viewed by others
Introduction
Major depressive disorders rank among the top five diseases with the highest global burden in North America and Europe (Murray et al, 2012). Although pharmacological treatments exist, several trials lasting each several weeks are often necessary before finding the most efficient therapy for a given individual, and more than 10% of depressed individuals will not respond to any correctly conducted interventions (Rush et al, 2006). These findings partly reflect our lack of understanding of the biological mechanisms underlying depression and antidepressant response.
Micro-RNAs (MiRNAs) are small, single-stranded, 21-base RNA transcripts that bind to messengers RNAs (mRNA) and target them for degradation, and as such play an important role in control of available mRNA in the cell (Mellios et al, 2008). MiRNAs circulate in blood and other fluids and are stably transported in small double lipid layered vesicles of 40–100 nm called exosomes (Valadi et al, 2007) or by lipoproteins such as HDL and LDL (Vickers et al, 2011). In addition, circulating miRNAs reflect underlying physiological and pathological states, and their levels may change as a result of treatment effects (Gilad et al, 2008; Chen et al, 2009; Zhou et al, 2009; O’Connor et al, 2012). As a consequence, there has been growing interest in the role of miRNAs as biomarkers of disease and treatment response. MiRNAs may notably play a role in response to psychotropic drugs, including lithium, valproate, and antidepressants (Chen et al, 2009; Zhou et al, 2009; Baudry et al, 2010; Bocchio-Chiavetto et al, 2013; Issler et al, 2014).
Using post-mortem human brain samples, cellular assays and blood samples from clinical trials of depressed patients, we have recently shown that miR-1202, an miRNA specific to primates, is differentially expressed in the prefrontal cortex of depressed individuals, predicts clinical response, and is upregulated in vitro and in vivo in response to antidepressant treatment (Lopez et al, 2014). Moreover, the effect of miR-1202 on antidepressant response may be related to its regulation of the glutamatergic system via its effect of the expression of the metabotropic receptor GRM4 (Lopez et al, 2014). An important question that follows from our previous study is the relationship between the peripheral variations of miR-1202 levels and changes in brain activity in vivo. Answering this question is important to elucidate potential mechanisms whereby miR-1202 may mediate antidepressant response, and to validate its role as a biomarker. The main objective of the present study was therefore to investigate this relationship using functional neuroimaging.
Specifically, we measured both miR-1202 and brain activity at rest and during a task before and after an 8-week antidepressant treatment. Impaired cognitive inhibition has been associated with significant depressive symptoms like ruminations (Lau et al, 2007), and with dysfunctional prefrontal and parietal brain activity during depression (Richard-Devantoy et al, 2016). Here, we used the classical version of the Go/NoGo task, as performance at this task has been shown to be impaired in depressed individuals (Kaiser et al, 2003). We additionally measured resting-state activity, as perturbed brain activity and connectivity at rest have been reported in depressed patients (Mulders et al, 2015; Murrough et al, 2016), and were found to be modulated by efficient antidepressant treatment (Dichter et al, 2015; Fu et al, 2015).
In this pilot study, we hypothesized that (1) baseline miR-1202 levels would predict change in at-rest and task-related brain activity/connectivity occurring during an 8-week antidepressant treatment and (2) changes in miR-1202 levels from baseline to week 8 would be correlated with changes in brain activity/connectivity in regions previously associated with depression.
Materials and methods
Population and Assessment
All participants were recruited from an outpatient clinic at the Douglas Mental Health University Institute in Montréal, Canada, and assessed by an experienced psychiatrist using the SCID-I (First et al, 2012) following DSM-IV criteria. Patients were all suffering from a current major depressive episode as part of a major depressive disorder. Exclusion criteria included comorbidity with other major psychiatric disorders, bipolar disorder, alcohol or substance abuse over the last 6 months, and a severe medical condition. All were right-handed as confirmed by the Edinburgh handedness inventory (Oldfield, 1971) and had no counter-indication to magnetic resonance imaging (MRI). None of the participants were medicated at baseline. None had received fluoxetine or lithium over the last month, or any psychotropic medication over the last week. Five patients (25%) had no lifetime history of psychotropic medication. Informed written consent was obtained from all participants. This study was approved by the local ethics committee.
Twenty-three participants were recruited and completed two MRI sessions, at baseline before any medication (T0) and after 8 weeks of an antidepressant treatment (T8). Blood samples were drawn at both time points for miR-1202 analyses. Three patients had to be excluded due to technical issues so final imaging analyses were conducted in 20 patients (mean age=42.3 (SD=9.3); 13 (65%) females).
All patients were treated for 8 weeks with desvenlafaxine, a serotonin–norepinephrine reuptake inhibitor, started at 50 mg and increased to 100 mg die if needed. Response was defined as more than 50% improvement in Hamilton Depression Rating Scale (HAMD-21; Hamilton, 1960) scores between T0 and T8, and remission as a T8 score below 8. In our sample, mean HAMD scores were 30.3 (4.8) at T0 and 14.8 (6.9) at T8. Three patients reached remission (15%) and 14 (70%) were responders.
Statistical Analyses
As most variables showed a close to normal distribution, parametric tests were used. Comparison of quantitative values between T0 and T8 were conducted with paired t-tests. ANOVA or t-tests was used for between-group comparisons. Correlations between quantitative values and miR-1202 levels were calculated with a Pearson’s test. A χ2 test was used to compare qualitative values. Finally, a General Linear Model was used to investigate the effect of covariates.
In line with what was previously done (Lopez et al, 2014), two measures were used for analyses with miR-1202: The first one was the baseline miRNA levels before any treatment (T0), whereas the second was the ratio between miRNA levels at the end of the 8-week treatment on levels at baseline (T8/T0) as a measure of miR level changes. An α threshold of 0.05 was set a priori for all analyses. SPSS 23.0 (SPSS, Chicago, IL) was used for statistical analyses.
Blood Samples Processing and RNA Extraction
Blood samples were collected in PAXgene blood RNA tubes (PreAnalytix) at T0 and T8. PAXgene tubes were frozen using a sequential freezing process. This involved storing tubes at room temperature for 3 h, transferring to 4 °C overnight, −20 °C for 6–8 h and then finally to −80 °C. Total RNA (including miRNA fraction) was isolated using the miRNeasy Mini Kit protocol (Qiagen, Canada) with no modifications. RNA and miRNA yield and quality were determined using the Nanodrop 1000 (Thermo Scientific, USA) and Agilent 2100 Bioanalizer (Agilent Technologies, USA), respectively. The RNA Integrity Number value was >8 for all our samples.
Quantitative Real-Time Polymerase Chain Reaction
MicroRNA was reverse transcribed using TaqMan RT-PCR microRNA assays (Applied Biosystems) according to the manufacturer’s instructions. Real-time PCR reactions were run in triplicates using the ABI 7900HT Fast Real-Time PCR System and data were collected using the Sequence Detection System (SDS) software (Applied Biosystems). Expression levels were calculated using miRNA TaqMan probes and the Absolute Quantitation (AQ) standard curve method. RNU6B was used as the endogenous control, as it showed expression levels that remained relatively constant with low variance and high abundance across human samples tested. The following TaqMan probes were used for miRNA quantification: miR-1202 (Assay# 002858); RNU6B (Assay # 001093).
In our sample, mean miR-1202 levels were 0.57 (0.30) at T0 and 0.53 (0.27) at T8.
Go/NoGo Task for Functional MRI
A classical version of the Go/NoGo task was implemented in E-Prime 2.0.10.182 (Sharpsburg, PA, USA) as a measure of cognitive inhibition. Stimuli were displayed in an MRI-compatible liquid crystal display at the rear of the scanning bore, viewable via a mirror by the participant. Each task was composed of six blocks: three Go and three NoGo blocks were presented in an ABBAAB order interleaved with 20-second blank-screen resting periods and 5 s instructions prior to the start of each block. In the Go block, participants were instructed to respond to all letters (black letters on a white screen) indiscriminately by pressing the button with their right index finger as quickly as possible. In the No-Go block, participants were instructed to respond as quickly as possible to target letters (ie all letters other than X) but NOT to equally frequent non-target letters (letter X).
Each block lasted 62 s and consisted of 24 trials. Each trial was composed of a fixation cross followed by a letter. The duration of the fixation cross varied between 700, 900, 1100 or 1300 ms, randomized to prevent habituation, six trials of each duration resulting in an average fixation cross duration of 1000 ms across the entire block and experiment. The target/non-target letters were always displayed for 500 ms. All blocks had 12 predetermined pseudo-randomly distributed target letters (50%) and 12 non-target letters (50%). In all sequences, reaction times were recorded, as were omission errors (ie, not pressing a target letter) in both conditions and commission errors (ie, responding to a non-target letter) in the NoGo conditions. Omission scores are usually interpreted as reflecting attention abilities, while commission scores are a more direct measure of inhibitory processes.
Image Acquisition
Scans were conducted at the Douglas Mental Health University Institute’s Cerebral Imaging Centre using a Siemens Magnetom Trio (Tim System 3T, MR B17) MRI scanner with a 12-channel head coil. Blood-oxygen-level dependent functional scans were acquired during one single session for the resting state sequence and the Go/NoGo task. For the resting state, 285 volumes consisting of 38 contiguous 3.5-mm transversal slices were acquired with a T2-weighted gradient echo-planar imaging sequence (repetition time: 2.09 ms; echo time: 30 ms; field of view: 24 mm; base resolution: 64 × 64; in-plane resolution: 3.5 × 3.5 mm; GRAPPA acceleration: 2; descending sequential acquisition). During the Go/NoGo task, 175 volumes were acquired with the same parameters as described above. A structural sequence was also acquired consisting of a high-resolution, whole-brain T1-weighted acquisition using a Magnetization Prepared Rapid Gradient Echo (MPRAGE) sequence with TR/TE/flip angle=2300 ms/2.98 ms/9°, and a base resolution of 256 × 256, with 1 mm3 isotropic voxels resulting in acquisition time of 9.25 min.
Image Analyses
Pre-processing
Robust skull stripping algorithms (Iglesias et al, 2011) were applied on structural images before registering them into MNI standard space. The spatial nonlinear transformation, estimated on registrations stage, was stored to be used on fMRI data set. Functional images were ‘pre-processed’ following Pereira et al (2010): (1) slice-timing correction was employed on each time-point using Fourier function and the middle axial slice (18th) was selected as the reference section; (2) corrections for head motion were performed on successive time points using Fourier interpolation. The parameters of rotation and translation, regarding the first time-point, were stored to be used as co-factors on further analysis; (3) remaining images were averaged across time-points and registered into the native-space of the structural image. The linear transformation estimated here was concatenated with the prior spatial transformation and applied on the aligned functional data set.
Default-mode network and correlations with miR-1202
Independent spatial maps (IC) were computed using Probabilistic Independent Component Analysis (Beckmann and Smith, 2004). This procedure basically seeks independent sources in the spatial dimension that are mutually associated to the similar fluctuations. The set of ICs was obtained using MELODIC v3.14, an FSL toolbox (Jenkinson et al, 2012). It was pulled out as many ICs as necessary to achieve 99% of the variability in the pre-processed data. Then, the set of ICs was masked with the Default-Mode Network (DMN) functional atlas (Shirer et al, 2012) and the z-values within and outside the mask were extracted. To determine the single component that best fits the DMN atlas, we employed the ‘Goodness-of-Fit’ (GoF) approach (Balthazar et al, 2014) as following:


The component with the highest GoF was picked up to represent the individual DMN map in each time-point (eg T0 and T8). The DMN maps were used to perform two distinct analyses. First, a paired group comparisons were carried out and permutation test was applied to control false-positive outcomes. Significant clusters (p-corrected<0.05) were then binarized to serve as a ‘seed’ on the Seed-Based functional-connectivity analysis (see below). Second, individual pairwise differences were computed and correlated with both miR-1202 scores (T0 and T8/T0) independently. Correction for multiple comparisons was carried out using uncorrected p-value of 0.001 and permutation analyses using 3dClustSim, a toolbox of AFNI (Cox, 2012). Only clusters at p<0.05 and minimum cluster size of 36 voxels are reported.
Seed-based functional connectivity and correlations with miR-1202
We extracted the time-series within the seed of pre-processed data and averaged them. The resulting mean time-series was regressed with pre-processed images using motion correction parameters as nuisance factor. β-Values estimated in this approach represent the individual Function Connectivity (FC) map, which was used to perform two distinct second-level analyses. Similarly to the DMN analyses, FC of at T0 and T8 were compared using paired t-test and permutation tests to correct for multiple comparisons. In addition, individual FC differences were correlated with both miR-1202 scores.
Go/No-Go contrast and correlation with miR-1202
A first-level fixed-effect block-design model was constructed for each participant at both time-points (before and after drug treatment). Gamma function with its linear and quadratic orders were used to model the block design. Head motion parameters as well as the linear and the quadratic drifts were regressed as nuisance variables whereas the ‘No-Go versus Go’ contrast was designed as contrast of interest. A second-level analysis was performed on the contrast images to evaluate main effects changes before and after the drug treatment. Correction for multiple comparisons was estimated using permutation tests implemented in 3dClustSim. Only clusters with corrected p<0.05 and minimum 36 voxels were considered significant. A pairwise difference of the contrast-of-interest (T8−T0) was then computed and independently correlated with the two miRNA parameters. Similarly, correction for multiple comparisons was employed and survival clusters had p<0.05.
Results
Resting State Analyses (Default-Mode Network and Functional Connectivity) and Correlations with miR-1202
Activation at T0 and T8, and contrasts between T8 and T0 within the default-mode network in the 20 participants are presented in Supplementary Figure S1. From T0 to T8, there was decreased activity in two clusters localized in the right and left posterior cingulate cortex (90 voxels; peak voxels with MNI coordinates: x=16, y=−56, z=8; Z-value=−3.8; and 61 voxels; −20, −58, 22; Z=−3.6). These two clusters were then used as one seed region for connectivity analyses with the whole brain. Functional connectivity results at T0, T8 and changes from T0 to T8 are presented in Supplementary Figure S2. Over time, there was decreased connectivity between this seed region and two clusters in right and left uncus/inferior temporal gyrus/parahippocampal gyrus (102 voxels; −26, −12, −42; Z=−3.7 and 36 voxels; 36, −18, −36; Z=−4.0). Decreased connectivity with the posterior default-mode network after antidepressant treatment is in line with the literature (Dichter et al, 2015).
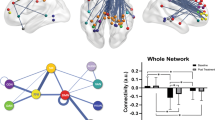
T0 miR-1202 was not associated with changes in activity within the default-mode network. However, it was negatively correlated with changes in connectivity between the seed region and seven clusters localized in occipital, parietal and anterior and dorsal prefrontal cortices (Table 1 and Figure 1).
Brain regions associated with miR-1202 levels during resting state over the 8-week antidepressant treatment. Top: Significant correlations between changes in miR-1202 levels (T8/T0 ratio) and changes in activity within the default-mode network (DMN); middle: significant correlations between baseline miR-1202 levels (T0) and changes in whole-brain functional connectivity (FC) with posterior cingulate; below: significant correlations between changes in miR-1202 levels (T8/T0 ratio) and changes in whole-brain FC with posterior cingulate.
T8/T0 miR-1202 was negatively correlated with changes in activity in a cluster localized in the right precuneus within the default-mode network. Moreover, changes in miR-1202 levels were negatively correlated with changes in the connectivity between the seed region and three clusters localized in left middle temporal and right anterior prefrontal cortices, and in left precuneus (Table 1 and Figure 1).
Go/NoGo Task Analyses and Correlation with miR-1202
Contrasts between NoGo and Go conditions at T0 and T8, and the difference between T8 and T0 for this contrast in the 20 patients are presented in Supplementary Figure S3. At corrected level, three large clusters showed significant changes over time, including two neighboring clusters (191 voxels; 2, −74, 20; Z=−3.9; and 169 voxels; 0, −62, 14; Z=−4.2) in bilateral precuneus (Brodmann Area (BA) 31) extending to occipital cortex (BA18) and posterior cingulate (BA23), and one smaller cluster (64 voxels; −6, −4, 52; Z=−4.6) in left medial frontal gyrus (BA6) extending to left cingulate gyrus (BA23). These regions have previously been associated with activation changes during an antidepressant treatment (Delaveau et al, 2011). Post-hoc explorations showed larger difference in activation between Go and NoGo conditions at T8 relative to T0, due to a larger deactivation during NoGo at T8. Moreover, there was a trend for decreased number of commission (t=1.9; DoF=19; p=0.08) but not omission errors with time.
T0 miR-1202 levels were correlated with activity changes in seven clusters on both hemispheres, overlapping the temporal, occipital, frontal, dorsolateral and dorsomedial prefrontal, and anterior cingulate cortices, in addition to some left subcortical nuclei (lentiform nucleus, including putamen and claustrum) (Table 2 and Figure 2). All correlations were negative at the exception of the smallest cluster.
Brain regions associated with miR-1202 levels during the Go/NoGo task over the 8-week antidepressant treatment. Top: Significant correlations between baseline miR-1202 levels (T0) and changes in NoGo vs Go contrast; below: significant correlations between changes in miR-1202 levels (T8/T0 ratio) and changes in NoGo vs. Go contrast.
Changes in miR-1202 levels were correlated with activity changes in nine clusters on both hemispheres, overlapping the temporal, occipital, ventrolateral and ventromedial prefrontal, and subcallosal cingulate cortices, in addition again to the same left subcortical nuclei (Table 2 and Figure 2). Both positive and negative correlations were observed. Of note, regional overlap between T0 and T8/T0 correlational analyses was very limited at the exception of the left subcortical nuclei.
Secondary Analyses: Correlation between miR-1202 and Glutamate
In order to investigate the mechanisms underlying miR-1202 effects, we took the opportunity of available spectroscopy data collected at T0 in five of our participants. Data were acquired in the right dorsomedial prefrontal cortex roughly corresponding to region BA6 (Jollant et al, 2016). We found a trend for a significant positive correlation between baseline miR-1202 and glutamate levels measured during the same session at rest (r=0.86; p=0.06) but not GABA, inositol, or NAA (all p>0.1). Of note, these five patients had low miR-1202 levels, a factor predictive of antidepressant response (Lopez et al, 2014).
Discussion
Baseline levels of miR-1202 have previously been shown to be predictive of antidepressant response and to increase during response to antidepressant treatment. The present study showed that peripheral miR-1202 levels are also associated with significant changes in both task-based and resting-state activity following 8 weeks of antidepressant treatment. These findings suggest that variation in peripheral miR-1202 levels may reflect biochemical changes occurring at the central level, and miR-1202 may be involved in the modulation of central circuitry functioning. Our results therefore add to the previous investigations of this miRNA and represent an advance in the attempt to link different biological levels of understanding of the antidepressant response.
Brain regions associated with miR-1202 level changes include ventral and dorsal prefrontal cortex, insula, temporal cortex, posterior cingulate, parietal (notably precuneus), and occipital cortices, and subcortical nuclei notably the putamen and claustrum. These regions have previously associated with antidepressant response (Delaveau et al, 2011). Research in depressed patients has shown structural and functional alterations in corticolimbic regions, associated with emotional and cognitive disturbances (Seminowicz et al, 2004; Fitzgerald et al, 2008; Dutta et al, 2014; Phillips et al, 2015), and functional connectivity alterations between the prefrontal cortex, anterior and posterior cingulate cortices, and parietal cortex, among others (Mulders et al, 2015). Conversely, antidepressant drugs tend to restore a normal brain activity while improving depressive symptoms (Fu et al, 2004; Fitzgerald et al, 2008; Delaveau et al, 2011). Our findings suggest that miR-1202 may be implicated in the regulation of the activity and connectivity of several of these regions and networks, and consequently contribute to the clinical response.
To our knowledge, very few studies have to date investigated the link between miRNAs and brain activity in vivo, and none in depression. One study previously found a significant association between levels of miR-29c and functional brain connectivity with the ventromedial prefrontal cortex during a social stress task in healthy males (Vaisvaser et al, 2016). Another study showed that rats overexpressing miR-30e, an miRNA associated with cognitive impairment, present aberrant regional resting-state synchronization in the hippocampus and striatum (Xu et al, 2016). These results and ours therefore suggest a significant association between miRNA variation and brain functioning at the circuit level in vivo.
Much remains to be done to understand the mechanisms linking miR-1202 and brain activity. However, preliminary hypotheses can be put forward. MiR-1202 negatively regulates the expression of GRM4 (Lopez et al, 2014), a metabotropic glutamatergic receptor whose expression is increased during depression (Lopez et al, 2014; Gray et al, 2015) and decreased during an antidepressant treatment (Cruceanu et al, 2016). MiR-1202 may therefore modulate the activity of the glutamatergic system. In the present study, we report preliminary evidence for an association between glutamate levels in the dorsomedial prefrontal cortex and peripheral miR-1202 levels in vivo. Interestingly, recent reports combining spectroscopy and resting-state analyses showed that prefrontal glutamate levels in depressed women were positively correlated with spontaneous fluctuations during resting state (Zhang et al, 2016). Moreover, glutamate levels were correlated with the activity of brain regions outside of the location where glutamate was measured (Duncan et al, 2013), suggesting a significant role in long-distance functional connectivity. Prefrontal glutamate was also found to modulate the relationship between striatal dopamine and response inhibition (Lorenz et al, 2015). Overall, one hypothesis may be that variations in central miR-1202 levels may modulate the glutamatergic system, which in turn may modulate resting-state activity and response inhibition, through the modulation of long-distance functional connectivity.
This pilot study presents several limitations that have to be underlined. First, the sample is relatively small, increasing the risk of both type-I and II errors. Results will therefore have to be replicated in independent and larger samples. This small sample size notably prevented us from running analyses in subgroups of patients, for example, according to clinical response, age, or gender. Second, the lack of a placebo group prevents us from discriminating the specific effect of antidepressants from the global effect of care. While this was not an issue here, it may be interesting in future studies to distinguish these specific contributions. Third, we only focused on one miRNA. Interaction between several miRNA may be in play to modulate the antidepressant effect. Similarly, while we examined two different functional imaging sequences, it is very plausible that different miRNA regulate different regions and connections, and ultimately different cognitive and emotional processes. Fourth, antidepressants with different actions may modulate different miRNA and brain networks. Fifth, analyses were correlational, limiting the interpretation of causal chains. Sixth, while patients had withdrawn from medication at least 7 days before and a quarter of them had never received any medication in their life, it cannot be excluded that past pharmacological treatments had long-term effects in several participants. As patients were severely depressed, it was difficult to maintain a longer washout period. Finally, results of the association between miR-1202 and metabolites measured by spectroscopy were conducted in a very small subsample, and while results are in line with previous findings linking the same miRNA and the glutamatergic system as discussed above, they should be interpreted with extreme caution.
In conclusion, our study showed that peripheral miR-1202 levels, an miRNA associated with response to antidepressant treatment, are correlated with changes in brain activity and connectivity occurring during an 8-week antidepressant treatment. These changes affected a large network of regions mainly encompassing the prefrontal, temporal, and parietal cortices. It is hypothesized that MiR-1202 levels modulate the antidepressant response through its modulation of brain circuitry functioning, possibly via the glutamatergic system.
Funding and disclosure
This study was funded by two operating grants to GT (MOP111260) and to FJ (119288) from the Canadian Institutes of Health Research (CIHR). JPL received PhD awards from the Fond de Recherche du Québec—Santé (FRQS) and CIHR, SR-D a CIHR fellowship grant, FJ a ‘chercheur-boursier clinicien’ FRQS salary grant at the time of the study. GT is supported by grants from the CIHR (MOP93775, MOP11260, MOP119429, and MOP119430), from the US National Institutes of Health (NIH) (1R01DA033684), by the FRQS through a Chercheur National salary award and through the Quebec Network on Suicide, Mood Disorders and Related Disorders, and through an investigator-initiated research grant from Pfizer. The authors declare no conflict of interest.
References
Balthazar ML, Pereira FR, Lopes TM, da Silva EL, Coan AC, Campos BM et al (2014). Neuropsychiatric symptoms in Alzheimer’s disease are related to functional connectivity alterations in the salience network. Hum Brain Mapp 35: 1237–1246.
Baudry A, Mouillet-Richard S, Schneider B, Launay JM, Kellermann O (2010). miR-16 targets the serotonin transporter: a new facet for adaptive responses to antidepressants. Science 329: 1537–1541.
Beckmann CF, Smith SM (2004). Probabilistic independent component analysis for functional magnetic resonance imaging. IEEE Trans Med Imaging 23: 137–152.
Bocchio-Chiavetto L, Maffioletti E, Bettinsoli P, Giovannini C, Bignotti S, Tardito D et al (2013). Blood microRNA changes in depressed patients during antidepressant treatment. Eur Neuropsychopharmacol 23: 602–611.
Chen H, Wang N, Burmeister M, McInnis MG (2009). MicroRNA expression changes in lymphoblastoid cell lines in response to lithium treatment. Int J Neuropsychopharmacol 12: 975–981.
Cox RW (2012). AFNI: what a long strange trip it’s been. Neuroimage 62: 743–747.
Cruceanu C, Lopez JP, Tsai WT, Turecki G (2016). Dysregulation of the glutamatergic receptors after antidepressant treatment in human neural progenitor cells. Mol Psychiatry (epub ahead of print).
Delaveau P, Jabourian M, Lemogne C, Guionnet S, Bergouignan L, Fossati P (2011). Brain effects of antidepressants in major depression: a meta-analysis of emotional processing studies. J Affect Disord 130: 66–74.
Dichter GS, Gibbs D, Smoski MJ (2015). A systematic review of relations between resting-state functional-MRI and treatment response in major depressive disorder. J Affect Disord 172: 8–17.
Duncan NW, Wiebking C, Tiret B, Marjanska M, Hayes DJ, Lyttleton O et al (2013). Glutamate concentration in the medial prefrontal cortex predicts resting-state cortical-subcortical functional connectivity in humans. PLoS One 8: e60312.
Dutta A, McKie S, Deakin JF (2014). Resting state networks in major depressive disorder. Psychiatry Res 224: 139–151.
First MB, Spitzer RL, Gibbon M, Williams JBW (2012) Structured Clinical Interview for DSM-IV® Axis I Disorders (SCID-I), Clinician Version, Administration Booklet. American Psychiatric Publishing: New York.
Fitzgerald PB, Laird AR, Maller J, Daskalakis ZJ (2008). A meta-analytic study of changes in brain activation in depression. Hum Brain Mapp 29: 683–695.
Fu CH, Williams SC, Cleare AJ, Brammer MJ, Walsh ND, Kim J et al (2004). Attenuation of the neural response to sad faces in major depression by antidepressant treatment: a prospective, event-related functional magnetic resonance imaging study. Arch Gen Psychiatry 61: 877–889.
Fu CH, Costafreda SG, Sankar A, Adams TM, Rasenick MM, Liu P et al (2015). Multimodal functional and structural neuroimaging investigation of major depressive disorder following treatment with duloxetine. BMC Psychiatry 15: 82.
Gilad S, Meiri E, Yogev Y, Benjamin S, Lebanony D, Yerushalmi N et al (2008). Serum microRNAs are promising novel biomarkers. PLoS One 3: e3148.
Gray AL, Hyde TM, Deep-Soboslay A, Kleinman JE, Sodhi MS (2015). Sex differences in glutamate receptor gene expression in major depression and suicide. Mol Psychiatry 20: 1057–1068.
Hamilton M (1960). A rating scale for depression. J Neurol Neurosurg Psychiatry 23: 56–62.
Iglesias JE, Liu CY, Thompson PM, TU Z (2011). Robust brain extraction across datasets and comparison with publicly available methods. IEEE Trans Med Imaging 30: 1617–1634.
Issler O, Haramati S, Paul ED, Maeno H, Navon I, Zwang R et al (2014). MicroRNA 135 is essential for chronic stress resiliency, antidepressant efficacy, and intact serotonergic activity. Neuron 83: 344–360.
Jenkinson M, Beckmann CF, Behrens TE, Woolrich MW, Smith SM (2012). FSL. Neuroimage 62: 782–790.
Jollant F, Richard-Devantoy S, Ding Y, Turecki G, Bechara A, Near J (2016). Prefrontal inositol levels and implicit decision-making in healthy individuals and depressed patients. Eur Neuropsychopharmacol 26: 1255–1263.
Kaiser S, Unger J, Kiefer M, Markela J, Mundt C, Weisbrod M (2003). Executive control deficit in depression: event-related potentials in a Go/Nogo task. Psychiatry Res 122: 169–184.
Lau MA, Christensen BK, Hawley LL, Gemar MS, Segal ZV (2007). Inhibitory deficits for negative information in persons with major depressive disorder. Psychol Med 37: 1249–1259.
Lopez JP, Lim R, Cruceanu C, Crapper L, Fasano C, Labonte B et al (2014). miR-1202 is a primate-specific and brain-enriched microRNA involved in major depression and antidepressant treatment. Nat Med 20: 764–768.
Lorenz RC, Gleich T, Buchert R, Schlagenhauf F, Kühn S, Gallinat J (2015). Interactions between glutamate, dopamine, and the neuronal signature of response inhibition in the human striatum. Hum Brain Mapp 36: 4031–4040.
Mellios N, Huang HS, Grigorenko A, Rogaev E, Akbarian S (2008). A set of differentially expressed miRNAs, including miR-30a-5p, act as post-transcriptional inhibitors of BDNF in prefrontalcortex. Hum Mol Genet 17: 3030–3042.
Mulders PC, van Eijndhoven PF, Schene AH, Beckmann CF, Tendolkar I (2015). Resting-state functional connectivity in major depressive disorder: a review. Neurosci Biobehav Rev 56: 330–344.
Murray CJ, Vos T, Lozano R, Naghavi M, Flaxman AD, Michaud C et al (2012). Disability-adjusted life years (DALYs) for 291 diseases and injuries in 21 regions, 1990-2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet 380: 2197–2223.
Murrough JW, Abdallah CG, Anticevic A, Collins KA, Geha P, Averill LA et al (2016). Reduced global functional connectivity of the medial prefrontal cortex in major depressive disorder. Hum Brain Mapp 37: 3214–3223.
O’Connor RM, Dinan TG, Cryan JF (2012). Little things on which happiness depends: microRNAs as novel therapeutic targets for the treatment of anxiety and depression. Mol Psychiatry 17: 359–376.
Oldfield RC (1971). The assessment and analysis of handedness: the Edinburgh inventory. Neuropsychologia 9: 97–113.
Pereira FR, Alessio A, Sercheli MS, Pedro T, Bilevicius E, Rondina JM et al (2010). Asymetricla hippocampal connectivity in mesial temporal lobe epilepsy: evidence from resting state fMRI. BMC Neurosci 11: 66.
Phillips ML, Chase HW, Sheline YI, Etkin A, Almeida JR, Deckersbach T et al (2015). Identifying predictors, moderators, and mediators of antidepressant response in major depressive disorder: neuroimaging approaches. Am J Psychiatry 172: 124–138.
Richard-Devantoy S, Ding Y, Lepage M, Turecki G, Jollant F (2016). Cognitive inhibition in depression and suicidal behavior: a neuroimaging study. Psychol Med 46: 933–944.
Rush AJ, Trivedi MH, Wisniewski SR, Nierenberg AA, Stewart JW, Warden D et al (2006). Acute and longer-term outcomes in depressed outpatients requiring one or several treatment steps: a STAR*D report. Am J Psychiatry 163: 1905–1917.
Seminowicz DA, Mayberg HS, McIntosh AR, Goldapple K, Kennedy S, Segal Z et al (2004). Limbic-frontal circuitry in major depression: a path modeling metanalysis. Neuroimage 22: 409–418.
Shirer WR, Ryali S, Rykhlevskaia E, Menon V, Greicius MD (2012). Decoding subject-driven cognitive states with whole-brain connectivity patterns. Cereb Cortex 22: 158–165.
Vaisvaser S, Modai S, Farberov L, Lin T, Sharon H, Gilam A et al (2016). Neuro-epigenetic indications of acute stress response in humans: the case of microRNA-29c. PLoS One 11: e0146236.
Valadi H, Ekström K, Bossios A, Sjöstrand M, Lee JJ, Lötvall JO (2007). Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9: 654–659.
Vickers KC, Palmisano BT, Shoucri BM, Shamburek RD, Remaley AT (2011). MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat Cell Biol 13: 423–433.
Xu C, Liu X, Song X, Gao Q, Cheng L, Wang L et al (2016). Aberrant resting state in microRNA-30e rat model of cognitive impairment. Neuroreport 27: 809–817.
Zhang X, Tang Y, Maletic-Savatic M, Sheng J, Zhang X, Zhu Y et al (2016). Altered neuronal spontaneous activity correlates with glutamate concentration in medial prefrontal cortex of major depressed females: an fMRI-MRS study. J Affect Disord 201: 153–161.
Zhou R, Yuan P, Wang Y, Hunsberger JG, Elkahloun A, Wei Y et al (2009). Evidence for selective microRNAs and their effectors as common long-term targets for the actions of mood stabilizers. Neuropsychopharmacology 34: 1395–1405.
Acknowledgements
We would like to thank Mrs Iliana Lilova for her role in patient recruitment, and M Yang Ding for his role in neuroimaging data acquisition and storing.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Neuropsychopharmacology website
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Lopez, J., Pereira, F., Richard-Devantoy, S. et al. Co-Variation of Peripheral Levels of miR-1202 and Brain Activity and Connectivity During Antidepressant Treatment. Neuropsychopharmacol 42, 2043–2051 (2017). https://doi.org/10.1038/npp.2017.9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2017.9
This article is cited by
-
Stress, microRNAs, and stress-related psychiatric disorders: an overview
Molecular Psychiatry (2023)
-
Epigenetic regulation in major depression and other stress-related disorders: molecular mechanisms, clinical relevance and therapeutic potential
Signal Transduction and Targeted Therapy (2023)
-
The important roles of microRNAs in depression: new research progress and future prospects
Journal of Molecular Medicine (2021)
-
Neural extracellular vesicle-derived miR-17 in blood as a potential biomarker of subthreshold depression
Human Cell (2021)
-
MiR-1202 Exerts Neuroprotective Effects on OGD/R Induced Inflammation in HM Cell by Negatively Regulating Rab1a Involved in TLR4/NF-κB Signaling Pathway
Neurochemical Research (2020)