Abstract
T cell antigen receptor (TCR) signaling drives distinct responses depending on the differentiation state and context of CD8+ T cells. We hypothesized that access of signal-dependent transcription factors (TFs) to enhancers is dynamically regulated to shape transcriptional responses to TCR signaling. We found that the TF BACH2 restrains terminal differentiation to enable generation of long-lived memory cells and protective immunity after viral infection. BACH2 was recruited to enhancers, where it limited expression of TCR-driven genes by attenuating the availability of activator protein-1 (AP-1) sites to Jun family signal-dependent TFs. In naive cells, this prevented TCR-driven induction of genes associated with terminal differentiation. Upon effector differentiation, reduced expression of BACH2 and its phosphorylation enabled unrestrained induction of TCR-driven effector programs.
Similar content being viewed by others
Main
After infection or immunization, naive CD8+ T cells undergo burst-like clonal proliferation and differentiation to generate a population of effector cells reactive against pathogen-associated antigens. After resolution of infection, the majority of responding cells are eliminated, allowing brisk restoration of immune homeostasis. A fraction of cells escape this fate and persist as memory cells1,2,3,4,5,6. The presence of greater numbers of antigen-specific memory cells enables more efficient pathogen clearance upon secondary infection. Thus, dynamic regulation of T cell differentiation, proliferation and survival is required to generate then curtail effector responses while maintaining a subset of pathogen-specific memory cells after withdrawal of antigen.
TCR signaling is critical to both initiation and diversification of CD8+ T cell responses. Strong or repeated TCR signaling drives progressive changes in gene expression that result in loss of lymphoid homing potential, acquisition of effector cell functions and, ultimately, terminal effector differentiation and apoptosis7,8. Conversely, memory cells differentiate in response to weak antigen signals that are insufficient to drive full effector differentiation1,5,9. Consequently, memory cells manifest only a subset of transcriptional changes accompanying effector differentiation, and their intermediate state of differentiation enables them to remain functionally quiescent, survive and circulate among secondary lymphoid tissues, where they can be efficiently recruited into secondary responses10,11,12. TCR signaling not only has a role in diversification of CD8+ T cell responses but also induces functionally distinct outcomes within specific subpopulations of CD8+ T cells. TCR stimulation of naive cells predominantly results in proliferation and differentiation, stimulation of effector cells drives rapid induction of effector cytokines and cytotoxic molecules, and stimulation of terminally differentiated effector cells induces apoptosis1,8,9.
AP-1 family TFs have a central role in transducing TCR-driven effector programs. AP-1 TFs, including c-Jun, JunD and JunB; c-Fos, Fosb, Fosl1 and Fosl2; and BATF1, BATF2 and BATF3, contain basic leucine zipper (bZip) domains that enable them to form heterodimeric complexes at palindromic 12-O-tetradecanoylphorbol-13-acetate (TPA) response elements (TREs; 5′-TGA(C/G)TCA-3′)13,14. Members of the Jun family of AP-1 TFs are phosphorylated in response to TCR signaling and recruited to TREs within the enhancers of multiple genes involved in effector differentiation, where they predominantly activate gene expression15,16,17,18,19,20. We hypothesized that modulation of the availability of AP-1 sites to Jun TFs allows TCR-driven effector programs to be modulated in a stage-specific and contextual manner in CD8+ T cells, allowing for generation of transcriptionally intermediate memory cells.
BACH2 is a 92-kDa transcriptional repressor of the bZip TF family21. BACH2 promotes the differentiation of Foxp3+ regulatory T (Treg) cells, and this function is required under homeostatic conditions to prevent lethal inflammation22. In B cells, BACH2 is critical for somatic hypermutation and class-switch recombination, and its absence leads to impaired generation of class-switched antibody responses23,24. BACH2, like the AP-1 TFs, contains a bZip domain and binds to Maf recognition elements (MAREs), which embed a TRE sequence21. Silencing of Bach2 mRNA after activation of CD8+ T cells in vitro results in reduced cellular persistence25. These observations led us to explore whether BACH2 regulates CD8+ T cell differentiation in vivo by controlling access of AP-1 family TFs to the regulatory elements of TCR-induced genes.
Results
BACH2 is required for CD8+ T cell memory
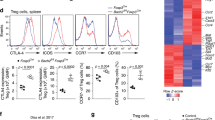
Defective generation of Foxp3+ Treg cells in Bach2−/− mice results in unrestrained effector differentiation among conventional T cells22. To evaluate the cell-intrinsic function of BACH2 in CD8+ T cells, we reconstituted C57BL/6 mice with 1:1 mixtures of congenically distinct CD45.1+ wild-type (WT) and Thy-1.1+ Bach2−/− mature lineage-depleted (Lin−) bone marrow (BM) cells (Supplementary Fig. 1a) and evaluated their CD8+ T cells. We observed low frequencies of both effector (CD62L−) and central memory (CD62L+CD44+) cells within the Bach2−/− compartment, suggesting a cell-intrinsic function of BACH2 in either the generation or maintenance of antigen-primed CD8+ T cell responses (Supplementary Fig. 1b,c). We therefore sought to test whether BACH2 controls responses to defined antigens. We used an acute viral infection model wherein recombinant vaccinia virus expressing chicken egg ovalbumin (VV-OVA) is recognized by minimal numbers of adoptively transferred TCR-transgenic CD8+ T cells specific for the OVA257–264 epitope (OT-I cells). First, mice were reconstituted with mixtures of congenically distinct CD45.1+ WT and Thy-1.1+ Bach2−/− OT-I transgenic BM cells, and naive CD44−CD62L+ OT-I cells of both genotypes were isolated from reconstituted mice. Naive WT and Bach2−/− cells were cotransferred at a 1:1 ratio into recipient C57BL/6 mice (Fig. 1a and Supplementary Fig. 1d), which were then infected with VV-OVA. Bach2−/− CD8+ T cells showed impaired expansion and a near-complete failure to establish long-lived memory responses (Fig. 1b,c). The reduced ratio of Bach2−/− to WT cells in spleens of immunized mice was similar to that in the lungs and liver, but there was a further reduction in the frequency of Bach2−/− cells in lymph nodes (Fig. 1d). Thus, BACH2 is required for maintenance of CD8+ T cell responses after primary infection in vivo.
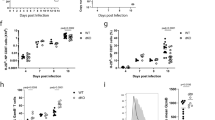
(a) Pre-transfer flow cytometry analysis of CD45.1+ WT and Thy1.1+ Bach2−/− (KO) naive OT-I cells mixed at a ∼1:1 ratio. (b,c) Kinetic analysis of cells in a after transfer into recipient mice and infection with VV-OVA. Representative flow cytometry analysis (b) and replicate measurements (c) are shown. (d) Ratio of WT and KO CD8+ T cells in various organs at day 21 after infection, normalized to ratios in spleen. iLN, inguinal lymph nodes. (e) Bacterial titers after LM-OVA infection of mice previously administered individual transfers of naive WT or KO OT-I cells and infected with VV-OVA. Primary infection with VV-OVA occurred 60 d before secondary infection with LM-OVA. (f,g) Secondary recall responses mediated by WT and KO TCM (f) and TEM (g) cells isolated at day 14 after primary infection as in c upon ∼1:1 mixed transfer into infection-naive recipients subsequently infected with VV-OVA. Numbers in flow cytometry plots (a,b) indicate percentages of cells in outlined areas. NS, not significant; *P < 0.05; **P < 0.01; ***P < 0.005; ****P < 0.001 (unpaired two-tailed Student's t-test). Data (a–g) are representative of two independent experiments with 4–6 mice per genotype and time point (error bars represent mean ± s.e.m. in c–g).
To test the function of BACH2 in mediating secondary protective immunity after primary infection, we infected mice with a recombinant strain of Listeria monocytogenes expressing the OVA257–264 epitope (LM-OVA) 60 d after individual transfer of normalized numbers of naive WT or Bach2−/− OT-I cells and primary infection with VV-OVA. Consistent with decreased maintenance of CD8+ T cell responses in the absence of BACH2, mice that had initially received Bach2−/− OT-I cells showed significant impairment in their ability to clear LM-OVA compared with mice that had received WT cells (Fig. 1e). We also asked whether BACH2 is required for secondary recall responses. We cotransferred 1:1 ratios of WT and Bach2−/− central memory T (TCM) cells (CD44+CD62L+) or effector memory (TEM) cells (CD44+CD62L−) isolated at day 14 after primary infection into infection-naive recipients then infected these mice with VV-OVA. Notably, both Bach2−/− TCM and TEM cells showed defective secondary response kinetics, although the defect was more pronounced in TCM than TEM cells (Fig. 1f,g). Collectively, these findings indicate that BACH2 has a nonredundant cell-autonomous role in maintenance of primary and secondary CD8+ T cell responses in vivo and in generation of protective immunity after primary viral infection.
BACH2 restrains terminal differentiation
Progressive acquisition of effector characteristics is accompanied by loss of lymphoid homing potential and reduction in cellular lifespan1,7,11. We asked whether defective maintenance and lymphoid homing of Bach2−/− CD8+ T cells responding to viral infection are accompanied by changes in differentiation state. Among antigen-experienced CD8+ T cells, CD62L+ central memory cells, CD62L−KLRG1− effector cells and KLRG1+ terminally differentiated effector cells have been described1. We measured expression of CD62L and KLRG1 on adoptively transferred naive WT and Bach2−/− OT-I CD8+ T cells after viral infection. We observed excessive induction of terminally differentiated KLRG1+ cells and near-complete loss of CD62L+ TCM cells among surviving Bach2−/− cells at late time points after infection (Fig. 2a–d). However, the decline in total CD8+ T cell numbers among Bach2−/− cells resulted in a decrease in the absolute number of terminally differentiated effector cells (Supplementary Fig. 2a) in addition to memory cells. Consistent with augmented effector differentiation of Bach2−/− cells, we noted higher expression of the effector cytokine interferon-γ (IFN-γ) upon brief restimulation ex vivo (Fig. 2e). Conversely, retroviral overexpression of BACH2 in OT-I CD8+ T cells prevented acquisition of terminally differentiated effector cell characteristics upon transfer into mice subsequently infected with VV-OVA, as indicated by increased expression of CD62L and lower expression of CD44 and KLRG1 (Fig. 2f). Additionally, cells transduced with BACH2 expressed reduced amounts of IFN-γ and the effector cytokine tumor necrosis factor (TNF) (Fig. 2g).
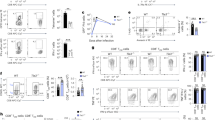
(a–d) Surface phenotype of cells before (a) or at indicated time points (b) after mixed transfer of naive WT and Bach2−/− (KO) OT-I CD8+ T cells into recipient mice and infection with VV-OVA. Representative flow cytometry (a,b) and replicate measurements of CD62L (c) and KLRG1 (d) on the surface of transferred cells are shown. Too few KO cells were detected at day 90 for phenotypic analysis. (e) Intracellular cytokine staining of IFN-γ expression by WT and KO CD8+ T cells from spleens of mice 5 d after infection. (f) Overexpression of BACH2 prevents terminal effector differentiation during secondary responses to infection in vivo. Expression of CD62L, CD44 and KLRG1 on the surface of CD8+ T cells 5 d after transfer of GFP+ (transduced) OT-I CD8+ T into mice subsequently infected with VV-OVA. (g) Expression of IFN-γ and TNF by transferred cells after brief restimulation ex vivo. Numbers in flow cytometry plots (a,b,g) indicate percentages of cells in outlined areas. *P < 0.05; **P < 0.01; ***P < 0.005; ****P < 0.001 (unpaired two-tailed Student's t-test). Data (a–g) are representative of two independent experiments with 4–6 mice per genotype and time point (error bars represent mean ± s.e.m. in c–g). EV, empty vector.
Terminal differentiation is associated with loss of memory precursor potential among effector cells26. Accordingly, we observed a loss of CD127+KLRG1− memory precursor effector cells (MPECs) and an increase in the frequency of CD127−KLRG1+ short-lived effector cells (SLECs) among Bach2−/− cells at day 7 after infection (Fig. 3a). We also noted a reduction in the frequency of CD127+ cells within the CD62L−KLRG1− effector population that predominates at day 7 (Supplementary Fig. 2b). We did not, however, observe differences in CD43 or CD27 expression27,28,29 (Supplementary Fig. 2c,d). We also observed increased acquisition of effector cell characteristics after stimulation of Bach2−/− CD8+ T cells in vitro, and this was antigen dependent (Supplementary Fig. 3a), proliferation independent (Supplementary Fig. 3b) and associated with high expression of lymphocyte activation markers CD25 and CD69 (Supplementary Fig. 3c).
(a) Expression of CD127 and KLRG1 on the surface of OT-I cells at day 7 after mixed transfer of naive FACS-sorted WT and KO OT-I CD8+ T cells into recipient mice and infection with VV-OVA. **P < 0.01, ****P < 0.001. (b) BrdU incorporation by WT and Bach2−/− (KO) CD8+ T cells at the indicated time points after infection. Mice were injected intraperitoneally with BrdU 16 h before harvest at each time point. *P < 0.05; ***P < 0.005 (unpaired two-tailed Student's t-test). (c) Annexin V and propidium iodide staining of cells at indicated time points after infection. **P < 0.01; ****P < 0.001 (unpaired two-tailed Student's t-test). NS, not significant. Numbers in gates (a,c) or adjacent to outlined areas (b) indicate percentages of cells. Data are representative of two independent experiments with 4–6 mice per genotype and time point (a–c; error bars represent mean ± s.e.m.).
Terminal differentiation of CD8+ T cells is accompanied by induction of apoptosis30,31. We measured rates of proliferation and apoptosis among WT and Bach2−/− CD8+ T cells responding to viral infection. Whereas BACH2 was dispensable for sustaining early proliferation (Fig. 3b), we observed substantially increased rates of apoptosis in Bach2−/− cells at days 5, 7 and 10 after infection (Fig. 3c), coinciding with their marked decrease in cell number. Defective survival was not attributable merely to induction of apoptosis at the naive cell stage, as we observed high rates of apoptosis in successive rounds of cell division after stimulation of Bach2−/− naive CD8+ T cells in vitro (Supplementary Fig. 3d). Terminal differentiation of CD8+ T cells is accompanied by depletion of antiapoptotic Bcl-2 proteins26,32,33. We observed decreased expression of Bcl-2 family members Bcl-xL and Mcl1 in Bach2−/− cells activated in vitro (Supplementary Fig. 3e). Corresponding to widespread differences in the phenotypes of WT and Bach2−/− RNA sequencing (RNA-seq) of these cells isolated at day 7 after acute infection showed widespread differences in gene expression (Supplementary Fig. 4a and Supplementary Table 1) and enrichment of transcripts associated with effector differentiation (Supplementary Fig. 4b and Supplementary Table 2). Gene expression differences between WT and Bach2−/− cells persisted but were substantially mitigated when fractionated populations of CD62L−KLRG1− effector cells were specifically analyzed (Supplementary Fig. 4c and Supplementary Table 3). Thus, BACH2 restrains terminal effector differentiation and apoptosis, enabling generation of long-lived memory cells after viral infection.
BACH2 controls TCR-driven transcriptional programs
AP-1 TFs bind to genomic TREs and have central roles in transducing TCR-driven effector programs. We hypothesized that BACH2 restrains TCR-driven terminal effector differentiation by binding to TREs and attenuating the activity of AP-1 factors. We mapped genome-wide BACH2 binding sites using chromatin immunoprecipitation (ChIP) coupled with massively parallel DNA sequencing (ChIP-seq). To generate sufficient numbers of cells for ChIP, we isolated naive CD8+ T cells by flow cytometry, cultured them with plate-bound anti-CD3 and anti-CD28 in the presence of IL-2 for 2 d then harvested them after an additional 3 d of culture (cells are referred to hereafter as d5 in vitro–activated CD8+ T cells). ChIP-seq of these cells showed extensive binding of BACH2 at intergenic and intronic regions (Fig. 4a). Analysis of these sites enabled identification of a BACH2 consensus motif (5′-TGA(C/G)TCAGC-3′) in CD8+ T cells, in which was embedded the canonical AP-1 recognition motif (5′-TGA(C/G)TCA-3′)34 (Fig. 4b). Notably, this consensus motif was very similar to a BACH2 recognition motif resolved by PCR-assisted binding-site selection in vitro21. Given that a majority of BACH2 binding sites were outside annotated promoter regions, we next asked whether BACH2 binds to enhancers. We measured the genome-wide distribution of the coactivator protein p300 and histone H3 acetylation at K27 (H3K27ac) or monomethylation at K4 (H3K4me1), which are characteristically located at enhancers35. We found that a majority of BACH2 binding sites colocalized with one or more of these markers (Fig. 4c). Consistent with similarity in BACH2 and AP-1 binding motifs, BACH2 binding sites were enriched at known genome-wide binding sites of AP-1 family members JunB, c-Jun and JunD17 (Fig. 4d–f and Supplementary Tables 4–6).
(a) Distribution of genome-wide BACH2 binding sites in d5 in vitro–activated CD8+ T cells relative to annotated features of known genes. Numbers indicate percentages. UTR, untranslated region. (b) Consensus motif enriched in BACH2 binding sites in CD8+ T cells (top) and AP-1 consensus motif (JASPAR; bottom). (c) Genome-wide colocalization analysis of BACH2 with indicated enhancer associated post-transcriptional histone modifications or p300 within d5 in vitro–activated CD8+ T cells. (d–f) Colocalization of JunB (d), c-Jun (e) or JunD (f) at genome-wide BACH2 binding sites in CD8+ T cells. Histograms indicate number of peaks at various distances from BACH2 peak summits. Pie charts (inset) indicate proportions of Jun binding sites (red) that directly overlap with BACH2 binding sites. Binomial tests were used to determine peak significance within ChIP-seq data, and a threshold of P < 1 × 10−5 was used for peak calling.
The extensive colocalization of BACH2 at known AP-1 sites in CD8+ T cells suggests that BACH2 may function to limit TCR-driven gene expression in CD8+ T cells. A prediction of this hypothesis is that the majority of TCR-driven genes repressed by BACH2 will not be differentially expressed in Bach2−/− and WT cells before stimulation. To test this, we measured global changes in gene expression induced by brief (6 or 18 h) ex vivo stimulation of naive CD44−CD62L+ WT and Bach2−/− CD8+ T cells using RNA-seq. We identified 192 TCR-driven genes that were upregulated in Bach2−/− cells at 0, 6 or 18 h after stimulation (fold change (FC) > 2; P < 0.05) (Fig. 5a). Most of these genes (163, 84%) did not show significant differences in expression at 0 h (i.e., before stimulation) (Fig. 5b,c and Supplementary Table 7). However, as early as 6 h after stimulation, we observed a notable increase in induction of genes encoding proteins associated with effector differentiation in Bach2−/− cells, including Ifng, Cd44, Ccl1, Ccl9, Icam1, Lif, Lta, Gadd45b and Ttc39c. A majority (72.3%) of genes upregulated in Bach2−/− cells contained mapped BACH2 binding sites within 15 kb upstream and 3 kb downstream of their gene bodies, suggesting that BACH2 directly represses their TCR-induced expression. The genes identified were highly enriched among transcripts induced in terminally differentiated KLRG1+ effector cells responding to VV-OVA infection compared with naive precursors (Fig. 5d and Supplementary Table 8).
(a) Genes upregulated in naive Bach2−/− (KO) CD8+ T cells compared with WT cells before (0 h) or after stimulation for 6 or 18 h with plate-bound anti-CD3 and anti-CD28 (P < 0.05, log2 FC > 1). Of these, 192 TCR-induced genes (upregulated by stimulation of WT or KO cells at 6 or 18 h) were identified. (b) Hierarchical cluster analysis showing expression of TCR-induced genes upregulated in KO cells. Fragments per kilobase of transcript per million mapped reads (FPKM) values are normalized to row maxima. (c) Fraction of genes in b differentially expressed in KO compared with WT cells at 0, 6 and 18 h (P < 0.05, log2 FC > 1). NS, not significant. (d) Enrichment of genes in b among transcriptional differences between terminally differentiated KLRG-1+ OT-I cells and naive cells. Normalized enrichment score (NES) = 1.9; Q < 0.0001. (e) Alignments showing mRNA expression in WT and KO naive CD8+ T cells (top) and mapped BACH2, JunD and H3K4me1 binding sites (bottom). (f) mRNA expression relative to Actb in naive WT and KO CD8+ T cells pretreated for 1 h with JNKi (12.5 μM) or vehicle (veh) at 37 °C and stimulated with plate-bound anti-CD3 and anti-CD28. *P < 0.01; **P < 0.001 (unpaired two-tailed Student's t-test). (g) mRNA expression at indicated time points after anti-CD3 restimulation of GFP+ (transduced) cells infected with retrovirus encoding BACH2 or BACH2 lacking its bZip domain (ΔZip) or empty vector (EV) during primary stimulation. AU, arbitrary units. Average FPKM values (b) and analyses of two replicate measurements per condition (a,c–e) are shown. Data in f and g represent mean ± s.e.m. of two independent experiments.
BACH2-mediated repression of TCR-driven gene expression is exemplified at the Ifng and Gp49a (Lilr4b) loci (Fig. 5e), where we observed colocalization of BACH2 and AP-1 factors at known36 or putative regulatory elements (Fig. 5e). To test whether excessive TCR-induced gene expression in Bach2−/− cells was driven by Jun factors, we used JNKi, a well-characterized pharmacological inhibitor of Jun N-terminal kinase (Jnk)37, which activates Jun family TFs in response to TCR signaling38. Pretreatment with JNKi partially reversed excessive induction of Gp49a and Ifng mRNA in naive Bach2−/−cells after stimulation (Fig. 5f). Moreover, BACH2-mediated repression of TCR-driven Gp49a and Ifng expression was abrogated in cells overexpressing BACH2 with a disrupted bZip domain (ΔZip BACH2), indicating that this suppression is dependent on the bZip domain (Fig. 5g). Collectively, these results indicate that BACH2 restrains TCR-driven transcriptional programs associated with terminal effector differentiation in naive CD8+ T cells.
BACH2 limits access of Jun family TFs to its binding sites
We hypothesized that BACH2 represses TCR-driven gene expression by competing with AP-1 factors for DNA binding. A prediction of this hypothesis is that loss of BACH2 will lead to increased binding of Jun family TFs at BACH2 binding sites. BACH2 was expressed at readily detectable levels in WT d5 in vitro–activated CD8+ T cells (Fig. 6a and Supplementary Fig. 5a). We restimulated WT and Bach2−/− in vitro–activated CD8+ T cells with plate-bound anti-CD3 for 2 h and subjected them to ChIP-seq analysis of JunD binding or RNA-seq analysis. Notably, Bach2−/− cells showed increased global binding of JunD at BACH2 binding sites compared with WT cells (Fig. 6a) despite having similar levels of total JunD (Fig. 6a). This JunD enrichment was restricted to shared JunD–BACH2 binding sites, as average JunD binding was not significantly increased at JunD sites where BACH2 does not bind (Supplementary Fig. 5b). Moreover, a proportion of JunD binding sites identified in Bach2−/− cells was not detected in WT cells, indicating that BACH2 prevents JunD binding with extremely high efficiency at a subset of loci (Supplementary Fig. 5c,d and Supplementary Table 9). In contrast to increased JunD binding, we observed little difference in chromatin accessibility at BACH2 binding sites between WT and Bach2−/− d5 in vitro–activated CD8+ T cells (Fig. 6b). Similarly, we found no correlation between the differences in JunD binding observed at BACH2 binding sites and levels of chromatin accessibility in WT and Bach2−/− cells (Supplementary Fig. 6a). Thus, within in vitro–activated CD8+ T cells, BACH2 inhibits JunD binding without affecting chromatin accessibility. This behavior is consistent with a function of BACH2 as a passive transcriptional repressor39.
(a) Histogram of JunD enrichment centered around genome-wide BACH2 binding sites in WT and Bach2−/− (KO) d5 in vitro–activated CD8+ T cells. Inset, immunoblot analysis of BACH2 and JunD expression. Numbers below blot indicate densitometric values of JunD abundance relative to β-actin. (b) Average assay for transposase-accessible chromatin (ATAC)-seq signal around BACH2 binding sites in WT and KO d5 in vitro–activated CD8+ T cells. RPM, reads per million. (c) Comparison of differences between average JunD binding at BACH2 binding sites with differences in mRNA expression of associated genes. TCR-induced genes (induced by 2 h anti-CD3 stimulation of d5 in vitro–activated CD8+ T cells) and noninduced genes are shown separately. Statistical significance was evaluated by two-sample Kolmogorov–Smirnov test. (d) Representative alignments of ChIP-, ATAC- and RNA-seq measurements in d5 in vitro–activated CD8+ T cells at selected loci showing concordantly increased mRNA expression and JunD binding in Bach2−/− compared with WT cells. Arrowheads indicate increased JunD binding at BACH2 binding sites in Bach2−/− cells. (e) mRNA expression at specific time points after restimulation of transduced (GFP+) d5 in vitro–activated CD8+ T cells infected with retrovirus expressing BACH2 or BACH2 ΔZip or empty vector (EV) during primary stimulation. AU, arbitrary units. (f) mRNA expression WT and KO CD8+ T cells activated in vitro in the presence of JNKi (12.5 μM) or vehicle (veh). *P < 0.001 (unpaired two-tailed Student's t-test). (g) Ratio of KO to WT cells in the spleens of recipient mice after transfer of congenically distinct GFP+ (transduced) WT and KO OT-I cells (∼1:1 ratio) infected with various retroviruses during primary stimulation and infection with VV-OVA for 5 d. AU, arbitrary units; NS, not significant; *P < 0.01 (unpaired two-tailed Student's t-test). Data and analyses are based on one (a), four (b) or two (c) measurements or represent two repeated experiments (e–g; error bars represent mean ± s.e.m.).
We next asked whether differences in JunD binding were associated with differences in gene expression. We compared average differences in JunD binding at BACH2 binding sites with differences in expression of associated genes in WT and Bach2−/− d5 in vitro–activated CD8+ T cells. Increased JunD binding in Bach2−/− cells was positively correlated with increased expression of associated genes, especially among TCR-induced genes (Fig. 6c,d and Supplementary Table 10). We confirmed increased JunD binding by qPCR-based ChIP quantification at select loci and extended our analysis by measuring occupancy of JunB and c-Jun, in addition to JunD, at these sites (Supplementary Fig. 6b). We observed increased JunD binding at all loci examined, increased c-Jun binding at a majority of loci and no increases in JunB binding in Bach2−/− cells. Loss of BACH2 ChIP signals in Bach2−/− cells confirmed specificity of the ChIP assay (Supplementary Fig. 6b).
Because passive transcriptional repressors work through competition with transcriptional activators, they often share DNA-binding domains with the transcriptional activators they antagonize39. BACH2 and AP-1 TFs share DNA-binding bZip domains. Consistent with a requirement for the bZip domain in repression of TCR-induced genes, BACH2 inhibition of TCR-induced gene expression was abrogated in cells transduced with BACH2 ΔZip (Fig. 6e). Additionally, and similarly to our observations in naive cells, excessive induction of TCR-driven genes in Bach2−/− d5 in vitro–activated cells was partially reversed upon pharmacological inhibition of Jun activity with JNKi (Fig. 6f). This led us to ask whether maintenance of CD8+ T cell responses to acute infection is dependent on the BACH2 bZip domain. Retroviral overexpression of BACH2 partially rescued the secondary response of d5 in vitro–activated Bach2−/− OT-I cells upon transfer and VV-OVA infection in vivo, whereas overexpression of ΔZip BACH2 did not (Fig. 6g). Collectively, these results indicate that BACH2 functions as a passive repressor of TCR-driven transcriptional programs, competing for access with AP-1 factors at a subset of its binding sites.
Stage-specific control of TCR-driven programs by BACH2
Given that effector differentiation proceeds in cells harboring intact Bach2 loci, we asked whether BACH2 expression is attenuated to permit effector differentiation in a subset of cells. Analyzing known genome-wide distributions of histone modifications in CD8+ T cells40, we noted decreased promoter-bound H3K4me3 at the Bach2 locus in memory and effector cells compared with naive cells (Fig. 7a,b). Consistently, Bach2 mRNA expression was highest in naive OT-I CD8+ T cells and was progressively downregulated in central memory (CD62L+KLRG1−), effector (CD62L−KLRG1−) and terminally differentiated effector (CD62L−KLRG1+) cells 21 d after VV-OVA infection (Fig. 7c). We observed corresponding changes in expression of Bach2 mRNA and the encoded protein in bulk populations of CD8+ T cells at serial time points after infection, with early decreased expression at days 5 and 7, when effector cells predominate, and increased expression at day 30, when memory cells recrudesce (Supplementary Fig. 7). Thus, expression of Bach2 is dynamically regulated in CD8+ T cells at distinct stages of differentiation.
(a,b) Genomic alignments (a) and quantification (b) of H3K4me3 in naive, memory (mem) and effector (eff) CD8+ T cells at the Bach2 gene. (c) mRNA expression in indicated subsets of OT-I cells. Naive cells prior to transfer or antigen-experienced cells sorted 30 d after transfer into WT recipient mice and infection with VV-OVA were analyzed. *P < 0.05, **P < 0.01 (unpaired two-tailed Student's t-test). (d) mRNA expression normalized to Actb (left) and protein expression (right) in naive and CD62L−KLRG1− d7 Eff cells. Numbers below blot indicate densitometric values of BACH2 abundance relative to β-actin. ****P < 0.001 (unpaired two-tailed Student's t-test). (e) Comparison of TCR-induced gene expression in KO and WT naive and d7 Eff cells. Sorted cells were subjected to 6 h stimulation ex vivo with plate-bound anti-CD3 and anti-CD28. TCR-induced genes (in stimulated naive WT (blue) or KO (red) cells; P < 0.05, log2 FC > 1) significantly upregulated in KO cells at 0 or 6 h (P < 0.05, log2 FC > 1) were selected for further analysis. (f) Comparison of expression of genes in e in naive and d7 Eff cells at 0 or 6 h after stimulation. Color scale of heat map normalized to row maxima. (g) Fraction of genes in f that were significantly differentially expressed between KO and WT cells at 0 or 6 h after stimulation (P < 0.05, log2 FC > 1). AU, arbitrary units. Data are representative of two independent experiments (c,d; mean ± s.e.m.) or average fragments per kilobase of transcript per million mapped reads (FPKM) values (f) or (e,g) two replicate measurements per genotype and time point.
We asked whether modulation of Bach2 mRNA expression enables stage-specific control of its function. CD62L−KLRG1− cells isolated at day 7 after infection (d7 Eff) expressed lower levels of Bach2 mRNA and the encoded protein than naive cells (Fig. 7d). Both WT and Bach2−/− d7 Eff cells are present in sufficient numbers for transcriptional analysis. We hypothesized that the effect of Bach2 deficiency on TCR-driven gene expression would be less substantial in d7 Eff cells than in naive cells, corresponding to its lower expression in these cells. We conducted RNA-seq analysis of TCR-induced gene expression in WT and Bach2−/− naive and d7 Eff cells. We identified TCR-driven genes that were upregulated in naive Bach2−/− cells compared with WT cells at 0 or 6 h after stimulation (Fig. 7e). We then compared TCR-induced expression of these genes in WT and Bach2−/− naive and d7 Eff cells (Fig. 7f,g and Supplementary Table 11). Notably, most of the genes differentially expressed in naive cells were not significantly differentially expressed between WT and Bach2−/− d7 Eff cells at 0 or 6 h after stimulation, corresponding with reduced BACH2 expression in this subset (Fig. 7f,g). Thus, dynamic regulation of BACH2 expression enables stage-specific control of TCR-driven transcriptional programs.
Contextual control of BACH2 function by phosphorylation
Incomplete extinguishment of Bach2 mRNA expression in KLRG1+ terminally differentiated CD8+ T cells prompted us to explore whether BACH2's function is further regulated by post-translational modification. We noted decreased mobility of BACH2 upon denaturing gel electrophoresis of lysates from cells that were briefly restimulated with anti-CD3, which was reversed upon treatment with lambda phosphatase (Fig. 8a). This suggested that BACH2 is phosphorylated upon TCR stimulation. A highly conserved serine residue (S520) of murine BACH2 corresponds to a previously reported phosphoresidue of human BACH2 in chronic myeloid leukemia cell lines41 (Fig. 8b). The residues surrounding S520 form a consensus motif of the serine–threonine kinase Akt. To test whether BACH2 S520 is phosphorylated by Akt after TCR stimulation, we transduced CD8+ T cells with retroviruses encoding WT BACH2 or a substitution mutant BACH2(S520A) after primary stimulation in vitro. Four days after transduction, cells were restimulated by crosslinking of surface-bound anti-CD3 monoclonal antibodies. Using an Akt-substrate motif-specific antibody that detects RXXpS, we tested whether inhibition of Akt with a well-characterized and highly specific allosteric Akt inhibitor (AKTi)42 prevents acute phosphorylation of BACH2 S520. Pharmacological inhibition of Akt prevented acute phosphorylation of BACH2, whereas inhibition of downstream mTORC1 activation with rapamycin did not (Fig. 8c). Moreover, the S520A substitution prevented detectable phosphorylation. Thus, S520 phosphorylation of BACH2 after stimulation of CD8+ T cells is prevented by pharmacological inhibition of Akt.
(a) Denaturing electrophoresis and immunoblot analysis of BACH2 migration in cellular lysates from d5 in vitro–activated CD8+ T cells restimulated for 30 min with crosslinked anti-CD3, treated with or without lambda phosphatase (LP). (b) Alignment of BACH2 amino acid sequences in various vertebrates. Conserved residues corresponding to mouse S520 and the S520A substitution mutation are shown in red. (c) Detection of phosphorylated BACH2 (p-BACH2) using an RXXpS-specific antibody. CD8+ T cells were transduced with retroviruses expressing Flag-tagged BACH2 (Flag-BACH2) or Flag-BACH2(S520A) during primary stimulation and restimulated with crosslinked anti-CD3 4 d after transduction, preincubating with AKTi or rapamycin for 1 h. IP, immunoprecipitation; lys, cell lysate. (d) Detection of p-BACH2 after addition of purified Akt1 and ATP to Flag-BACH2 or Flag-BACH2(S520A) immunoprecipitates. (e) Representative flow cytometry (top) and quantification (bottom) of cytokine expression and cell size in CD8+ T cells transduced with retrovirus expressing BACH2 or BACH2(S520A) or empty vector (EV) during primary stimulation in IL-2 and IL-12 and restimulated on anti-CD3-coated plates in the presence of brefeldin A. FSC, forward scatter; NS, not significant; *P < 0.05, **P < 0.01, ***P < 0.005 (unpaired two-tailed Student's t-test). (f,g) Localization of BACH2-GFP in CD8+ T cells transduced with various retroviruses and cultured as in e. Cells were briefly restimulated with plate-bound anti-CD3 (2 h) and stained for Thy-1.1 during the last 30 min of culture. Representative cells (f) and the ratio of nuclear to cytoplasmic mean pixel intensity (MPI) in replicate cells (g) are shown. ***P < 0.005. Data are representative of three (a,c,d) or two (e,g; mean ± s.e.m.) independent experiments and three culture replicates per group (e,g).
This observation led us to ask whether BACH2 is a direct substrate of Akt. We performed an in vitro kinase reaction combining purified Akt1 and BACH2 in the presence of ATP in vitro. Addition of purified Akt in the presence of ATP to BACH2 caused S520-dependent phosphorylation (Fig. 8d), indicating that BACH2 S520 is a substrate of Akt kinase in vitro. Because Akt drives effector differentiation in CD8+ T cells43,44,45 and effector differentiation is antagonized by its substrate BACH2, we hypothesized that phosphorylation of BACH2 promotes its functional inactivation, enabling effector differentiation to proceed. This hypothesis was supported by the observation that Akt inhibition suppresses effector differentiation in a partially BACH2-dependent manner (Supplementary Fig. 8). To test whether S520 phosphorylation functionally inactivates BACH2, we overexpressed BACH2 or BACH2(S520A) in CD8+ T cells stimulated in vitro in the presence of IL-2 and IL-12. Despite gating on cells expressing similar amounts of the transduction marker Thy-1.1, we noted enhanced suppression of effector cytokines and cell size when BACH2(S520A) was overexpressed (Fig. 8e). We used strongly activating in vitro stimulation conditions, as overexpression of WT BACH2 caused near-complete suppression of effector cytokines among transferred cells after VV-OVA infection (Fig. 3b), precluding assessment of a gain-of-function phenotype using this overexpression-based experimental system in vivo.
Phosphorylation of human BACH2 at a residue homologous to murine BACH2 S520 in chronic myeloid leukemia cells leads to its functional inactivation through nuclear exclusion41. To test whether phosphorylation of BACH2 causes its nuclear exclusion in T cells, we transduced CD8+ T cells with retroviral vectors expressing BACH2-GFP and BACH2(S520A)-GFP fusion proteins. The subcellular localization of GFP within transduced (Thy-1.1+) cells was resolved by imaging flow cytometry after brief restimulation to enable quantification of nuclear and cytoplasmic localization. WT BACH2 showed weaker GFP signal in the nucleus than BACH2(S520A) (Fig. 8f,g), indicating that phosphorylation of BACH2 at S520 favors its nuclear exclusion in CD8+ T cells. Thus, S520 phosphorylation provides a mechanism for extrinsic modulation of BACH2 function and contextual control of CD8+ T cell differentiation.
Discussion
Here we report a cell-autonomous role for BACH2 in regulating CD8+ T cell differentiation and function after primary and secondary infection in vivo. BACH2 restrains terminal effector differentiation and apoptosis within a subset of cells responding to viral infection, allowing generation of long-lived memory cells that mediate secondary protection upon pathogen reencounter. BACH2 binds to AP-1-like sites within enhancers and modulates their availability to Jun family signal-dependent TFs. In less differentiated cells, this enables BACH2 to prevent TCR-driven induction of genes associated with terminal differentiation; in effector cells, reduction of Bach2 mRNA expression and phosphorylation of BACH2 protein permit TCR-driven induction of effector cell transcriptional programs.
The observation that BACH2 modulates binding of Jun factors without affecting chromatin accessibility in activated CD8+ T cells suggests that BACH2 engages in passive transcriptional repression through competition with AP-1 factors for DNA binding. This is consistent with published observations that BACH2 exerts chromatin-independent repression of TPA-induced luciferase reporter gene expression driven by nonchromatinized MARE sequences46. These results do not exclude the possibility that BACH2 modulates chromatin structure in other cell types or CD8+ T cell subsets. The localized function of BACH2 in modulating access of AP-1 factors at enhancers is characteristic of short-range repression, which permits multiple enhancers to function autonomously within a promoter47; this function is compatible with a requirement for complex regulation of genes involved in lymphocyte differentiation. IRF4 is known to assemble with AP-1 factors at AP-1–IRF composite elements48. We did not observe statistically significant differences in Irf4 mRNA expression between WT and Bach2−/− cells. Because Irf4 is therefore unlikely to be a transcriptional target of BACH2 in CD8+ T cells, we conclude that differences in AP-1 factor binding in Bach2−/− cells are not attributable to an IRF4-dependent mechanism. It is also notable that only a subset of TCR-induced gene loci contributed strongly to the correlation between differences in JunD binding and gene expression in WT and Bach2−/− cells. These findings suggest that the extent of competition between BACH2 and AP-1 and its effect on gene expression may vary in a site-specific fashion. It will be important for future investigations to dissect the molecular mechanisms and sequence-level determinants of site-specific regulation of BACH2 function.
We found that reduced transcription of Bach2 in effector cells corresponded to a decrease in its ability to restrain TCR-induced gene expression, enabling stage-dependent control of its function. BACH2 was also post-translationally modified to enable further contextual regulation of its function. Our finding that BACH2 phosphorylated at S520 is functionally inactivated are consistent with published observations in chronic myeloid leukemia cells that phosphorylation at a homologous residue in human BACH2 causes nuclear exclusion and functional inactivation41. Further investigations will be required to precisely quantify the effect of changes in BACH2 expression and nuclear exclusion on binding of BACH2 and AP-1 factors at enhancers. We used TCR stimulation to study BACH2 phosphorylation; however, the finding that BACH2 is phosphorylated by Akt provides a potential mechanism by which a variety of extrinsic cues modulate T cell differentiation. The PI3K–Akt pathway integrates diverse antigenic, inflammatory, costimulatory and metabolic signals and regulates multiple aspects of lymphocyte biology44,45,49,50. It will be important to investigate the range of signals that regulate immune function via phosphorylation of BACH2 and how phosphorylation affects BACH2 function at endogenous levels of expression.
Collectively, our results show that BACH2 regulates transcriptional responses of CD8+ T cells to TCR signaling, enabling appropriate control of responses to viral infection. Our findings provide a basis for further investigation into how BACH2 controls the differentiation and function of CD8+ T cells and other lymphocyte lineages. Additionally, our results identify a new target for therapeutic manipulation of lymphocyte function in the context of chronic infection, autoimmunity and cancer.
Methods
Mice and reagents.
Experiments were approved by the Institutional Animal Care and Use Committee of the NCI and were performed in accordance with NIH guidelines. C57BL/6J, OT-I transgenic, B6-Ptprca Pepcb/BoyJ (Ly5.1) and B6.PL-Thy1a/CyJ (Thy-1.1) mice were purchased from the Jackson Laboratory. Bach2−/− mice, which have been described24, were backcrossed >16 times with C57BL/6 mice. Female mice, age 8–12 weeks, were used for experiments unless otherwise specified. Congenically distinct WT and Bach2−/− naive CD62L+CD44− CD8+ T cells were isolated by flow cytometric sorting from spleens and lymph nodes of mixed BM chimeric mice 6–12 weeks after reconstitution. For in vitro stimulation assays, naive CD8+ T cells were activated with plate-bound anti-CD3 (2C11; 1 μg/ml; eBioscience) and soluble anti-CD28 (37.51; 1 μg/ml; eBioscience) in medium containing 100 IU recombinant human IL-2 for 5 d. Where indicated, AKTi was added to culture medium at a concentration of 1 μM (AKT inhibitor VIII, Calbiochem), JNKi was added at a concentration of 12.5 μM (JNK Inhibitor I, L-Form, Calbiochem), and rapamycin (Sigma-Aldrich) was added at a concentration of 100 ng/ml.
Viral infection and kinetic analysis.
For experiments assessing the response of CD8+ T cells to acute viral infection, 2 × 104 congenically distinct WT or Bach2−/− naive CD44−CD62L+ OT-I CD8+ T cells were transferred into recipient mice. Immediately after transfer, mice were infected with 1 × 106 plaque-forming units (PFU) VV-OVA. Naive cells for transfer were isolated by flow cytometry from mice reconstituted with 1:1 mixtures of bone marrow (BM) cells from WT CD45.1+ and Bach2−/− Thy-1.1+ OT-I TCR-transgenic mice. Where indicated, WT and Bach2−/− naive cells were mixed at a 1:1 ratio before transfer.
Retroviral transduction.
pMIG- and pMIT-BACH2 constructs have been described previously21,22. S520A substitution mutations were generated by site-directed mutagenesis (Agilent). pMIT-BACH2-GFP and BACH2(S520A)-GFP were generated by cloning synthesized BACH2-GFP and BACH2(S520A)-GFP fusion constructs into the NotI and ClaI restriction sites within pMIT. Platinum-E ecotropic packaging cells (Cell Biolabs) were plated 1 d before transfection on poly-D-lysine–coated 10-cm plates (Becton Dickinson) at a concentration of 6 × 106 cells per plate. Packaging cells were transfected with 20 μg of retroviral plasmid DNA along with 6 μg pCL-Eco plasmid DNA using 60 μl Lipofectamine 2000 in 3 ml OptiMEM (Invitrogen) for 8 h in antibiotic-free medium. Medium was replaced 8 h after transfection, and cells were incubated for a further 48 h. Retroviral supernatants were collected and spun at 2,000 × g for 2 h at 32 °C into 24-well non-tissue-culture-treated plates coated overnight in RetroNectin (20 μg/ml; Takara Bio), 1 μg/ml anti-CD3 (2C11) and 1 μg/ml anti-CD28 (37.51) (eBioscience). Cells were cultured with 100 IU recombinant human IL-2. For experiments involving functional assessment and subcellular localization of BACH2(S520A), medium was additionally supplemented with 0.2–20 ng/ml IL-12 (Peprotech).
Antibodies and flow cytometry.
The following fluorochrome-conjugated antibodies to surface and intracellular antigens were used at 0.25–1 × 10−3 mg/ml: anti-Thy-1.1 (OX-7), anti-CD45.1 (A20), anti-KLRG1 (2F1), anti-CD25 (PC61), anti-CD62L (MEL-14), anti-IFN-γ (XMG1.2), anti-TNF (MP6-XT22), anti-CD44 (IM7), and anti-CD8α clone 53-6.7 (BD Biosciences). Cells were incubated with specific antibodies for 30 min on ice in the presence of 2.4G2 monoclonal antibody (mAb) to block FcγR binding. All samples were acquired with a FACS Canto II flow cytometer (Becton Dickinson) and analyzed by FlowJo software (TreeStar). To determine cytokine expression, cellular suspensions containing T cells were stimulated with phorbol 12-myristate 13-acetate (PMA), ionomycin and brefeldin A (Leukocyte Activation Cocktail with Golgiplug; BD Biosciences) for 4 h. After stimulation, cells were stained an amine-reactive exclusion-based viability dye (Invitrogen) and antibodies against cell-surface antigens, fixed and permeabilized and intracellularly stained with specific anti-cytokine antibodies. CountBright beads (Invitrogen) were added for the flow cytometric quantification of absolute cell number.
Imaging flow cytometric detection of BACH2 subcellular localization.
Total CD8+ T cells were isolated by negative magnetic selection (Miltenyi Biotec), stimulated using plates coated overnight with 1 μg/ml anti-CD3 (2C11) and 1 μg/ml anti-CD28 (37.51) in the presence of 100 IU IL-2 and 0.2–20 ng/ml IL-12 and transduced as described above with either pMIT-BACH2-GFP or pMIT-BACH2(S520A)-GFP. Four days after transduction, cells were briefly restimulated using plates coated with 1 μg/ml anti-CD3, stained with Thy-1.1-APC during the last 30 min of culture, fixed using 2% PFA in PBS, stained with DAPI and analyzed using an Amnis ImageStreamX MKII imaging flow cytometer. Data were analyzed with Amnis IDEAS. Briefly, single cells were selected on the basis of cell size, focus attributes and Thy-1.1, GFP and DAPI positivity. Cellular masks were defined as bounded by the Thy-1.1–APC cytoplasmic membrane signal, nuclear masks were defined using the DAPI signal, and a cytoplasmic mask was defined by subtracting the nuclear mask from the cytoplasmic mask. Mean pixel intensity of the BACH2-GFP signal was computed within the nuclear and cytoplasmic masks separately, and the ratio of the two values was analyzed to assess nuclear localization.
Immunoblotting and immunoprecipitation.
CD8+ T cells and HEK293T cells were lysed in IP lysis buffer (Pierce). The following antibodies were used for immunoblotting: anti-BACH2 (ab83364, lot GR40626-2, Abcam); anti-RXXpS (110B7E), anti-pAkt T308 (49D7), anti-pS6K1 T389 (9205) and anti-S6K1 (9202) (Cell Signaling); and anti-β-actin (AC-74; Sigma-Aldrich). For immunoprecipitation experiments Flag-BACH2/BACH2(S520A) was immunoprecipitated after overnight incubation with M-2 anti-Flag monoclonal antibodies (Sigma-Aldrich) using protein A/G magnetic beads (Dynal). After several washes, immunoprecipitated proteins were released from beads by competitive elution with Flag peptide according to manufacturer's instructions (Sigma-Aldrich) or by boiling in 1× Laemmli buffer. Proteins similarly immunoprecipitated from HEK293T cells transfected with WT or BACH2(S520A) were subjected to kinase assays while bound to beads and in the presence of ATP (Sigma-Aldrich) and kinase reaction buffer according to the manufacturer's instructions (Cell Signaling).
Quantitative reverse-transcription polymerase chain reaction (qRT-PCR).
Cells were isolated by flow cytometric sorting or harvested from cell cultures and transferred into RNALater solution (Ambion) for storage at −80 °C. Total RNA from pelleted cells was isolated using the RNAeasy Mini Kit (Qiagen). First-strand cDNA synthesis was performed using the high-capacity cDNA synthesis kit (Applied Biosystems) in the presence of SuperaseIn RNase inhibitor (Ambion). cDNA was used as a template for qPCR reactions using Taqman primer-probes against specified mRNA transcripts (Applied Biosystems). Reactions were performed using Fast Universal PCR Mastermix (Applied Biosystems) and thermocycled in quadruplicate 10 μl reactions in 384-well plates. Signals in the FAM channel were normalized to ROX intensity, and threshold cycle (Ct) values were calculated using automatically determined threshold values using SDS software (Applied Biosystems).
Bone-marrow chimera experiments and measurement of antiviral response kinetics.
For bone marrow (BM) reconstitution experiments, C57BL/6 mice were administered 1,000 Gy total-body γ-radiation from a 137Cs source before intravenous injection of BM cells with or without depletion of mature lineages from single-cell BM preparations from 6- to 10-week-old mice using antibody-coupled magnetic beads (Miltenyi Biotec). To measure antiviral responses, 2 × 104 OT-I cells were transferred intravenously into recipients before infection with 1 × 106 PFU VV-OVA. Where indicated, WT and Bach2 KO naive cells were sorted from mixed-BM-chimeric mice and mixed at a 1:1 ratio before transfer.
In vivo BrdU labeling.
For analysis of cellular proliferation in vivo, mice were administered 2 mg BrdU suspended in 500 μl PBS at 16 h (antiviral kinetic experiment) or every day for 5 d (steady-state proliferation in mixed-BM chimeras) before analysis. Incorporated BrdU was detected using the APC BrdU staining kit (BD Biosciences).
RNA sequencing and analysis.
RNA-seq analyses were performed using ≥2 biological replicates. RNA sequencing was performed and analyzed as described previously22. Briefly, total RNA was prepared from cells using the RNeasy Plus Mini Kit (Qiagen). 200 ng total RNA was subsequently used to prepare RNA-seq library by using TruSeq RNA sample prep kit (FC-122-1001, Illumina) according to the manufacturer's instructions. Sequenced reads were aligned to the mouse genome (NCBI37/mm9) with Tophat 2.0.11 (ref. 51), and uniquely mapped reads were used to calculate gene expression. RefSeq gene database (mm9) was downloaded from the UCSC genome browser for RNA-seq analysis. Raw counts that fell on exons of each gene were calculated, and differentially expressed genes were identified with the statistical R package edgeR52 or Cuffdiff53. Fisher's exact test or t-test were used to evaluate significance with indicated P value and fold-change thresholds. Expression heat maps were generated with the R package pheatmap.
Chromatin immunoprecipitation and accessibility assays.
T cells were chemically crosslinked and sonicated to generate fragmented genomic DNA. Chromatin immunoprecipitation was performed using the following antibodies: anti-BACH2 (N-2; Tohoku University), anti-JunD (sc-74), anti-c-Jun (sc-45), anti-JunB (sc-73) and anti-p300 (sc-586; Santa Cruz) anti-H3K4me1 (ab8895) and anti-H3K27Ac (ab4729; Abcam). For PCR-based confirmation of BACH2 binding, chromatin immunoprecipitation was performed as described above, and qPCR reactions were carried out on input and immunoprecipitated DNA using the Power SYBR Green kit (Applied Biosystems). Genome-wide measurement of chromatin accessibility and computational alignment of generated data were performed using ATAC-Seq on 50,000 d5 in vitro–activated CD8+ T cells as described54. Subsequent bioinformatic analyses were performed as described below.
ChIP-seq analysis.
For sequencing of immunoprecipitated DNA, DNA fragments were blunt-end ligated to Illumina adaptors, amplified, and sequenced by using the HiSeq 2000 platform (Illumina). Sequenced reads of 50–125 bp were obtained by using the Illumina Pipeline. Sequenced reads were aligned to the mouse genome (NCBI37/mm9) with Bowtie 0.12.9 (ref. 55); only uniquely mapped reads were retained. The output of Bowtie was converted to BAM files, which represent the genomic coordinates of each read. Reads were mapped into nonoverlapping 10-bp windows, and the location of reads on positive (negative) strand was shifted ±75 bp from its 5′ start to determine the approximate center of the DNA fragment associated with the reads. Genomic graphs were generated and viewed using the IGV (Integrative Genomics Viewer)56, and we aligned the BACH2, JunD, cJun, p300, H3K4me1, H3K27ac and input control in CD8+ T cells.
Peak calling, motif discovery and genome-wide distribution analysis.
We used MACS 1.4.2 (ref. 57) to call binding sites (peaks) relative to input control libraries. The binomial distribution test was used to assess peak significance, and a P value threshold of 1 × 10−5 was used unless otherwise specified. For motif discovery, the most significant 1,000 peaks ranked by lowest P value were used, and 100-bp DNA sequences centered on each peak summit were analyzed using MEME to characterize the BACH2 consensus binding motif in mouse CD8+ T cells. The discovered motifs were compared with the JASPAR motif database to evaluate motif similarities. 5′ untranslated regions (UTRs), 3′ UTRs, introns, exons and intergenic regions were defined according to the RefSeq database. Peaks were identified to be associated with genes if they were found between 15 kb upstream of the transcription start site and 3 kb downstream of the transcription end site of that gene. The distance between summits of two peaks was calculated and plotted as histograms using the R package.
Genome-wide binding profiles.
Genome-wide binding profiles were established by counting the reads into binned windows (20 bp/bin) relative to peak summits. Normalized read densities were plotted in the range [−1 kb, 1 kb] centered on peak summits. The Kolmogorov–Smirnov test was used to evaluate the significance between different distributions.
Statistical testing.
Unless otherwise specified, statistical tests were performed using unpaired two-tailed Student's t-test. Where necessary, the Shapiro–Wilk test was used to test for normality of the underlying sample distribution. No blinding was done, as objective quantitative assays such as flow cytometry, were used. Experimental sample sizes were chosen using power calculations with preliminary experiments or were based on previous experience of variability in similar experiments. Samples that had undergone technical failure during processing were excluded from analyses. Where relevant, recipient mice were randomized before adoptive transfer.
Accession codes.
Gene Expression Omnibus: Microarray data have been deposited under accession code GSE77857.
Accession codes
References
Kaech, S.M. & Cui, W. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 12, 749–761 (2012).
Belz, G.T. & Kallies, A. Effector and memory CD8+ T cell differentiation: toward a molecular understanding of fate determination. Curr. Opin. Immunol. 22, 279–285 (2010).
D'Cruz, L.M., Rubinstein, M.P. & Goldrath, A.W. Surviving the crash: transitioning from effector to memory CD8+ T cell. Semin. Immunol. 21, 92–98 (2009).
Williams, M.A. & Bevan, M.J. Effector and memory CTL differentiation. Annu. Rev. Immunol. 25, 171–192 (2007).
Restifo, N.P. & Gattinoni, L. Lineage relationship of effector and memory T cells. Curr. Opin. Immunol. 25, 556–563 (2013).
Chang, J.T., Wherry, E.J. & Goldrath, A.W. Molecular regulation of effector and memory T cell differentiation. Nat. Immunol. 15, 1104–1115 (2014).
Teixeiro, E. et al. Different T cell receptor signals determine CD8+ memory versus effector development. Science 323, 502–505 (2009).
Wirth, T.C. et al. Repetitive antigen stimulation induces stepwise transcriptome diversification but preserves a core signature of memory CD8+ T cell differentiation. Immunity 33, 128–140 (2010).
Ahmed, R. & Gray, D. Immunological memory and protective immunity: understanding their relation. Science 272, 54–60 (1996).
Roychoudhuri, R. et al. Transcriptional profiles reveal a stepwise developmental program of memory CD8+ T cell differentiation. Vaccine 33, 914–923 (2015).
Willinger, T., Freeman, T., Hasegawa, H., McMichael, A.J. & Callan, M.F. Molecular signatures distinguish human central memory from effector memory CD8 T cell subsets. J. Immunol. 175, 5895–5903 (2005).
Sarkar, S. et al. Functional and genomic profiling of effector CD8 T cell subsets with distinct memory fates. J. Exp. Med. 205, 625–640 (2008).
Turner, R. & Tjian, R. Leucine repeats and an adjacent DNA binding domain mediate the formation of functional cFos-cJun heterodimers. Science 243, 1689–1694 (1989).
Glover, J.N. & Harrison, S.C. Crystal structure of the heterodimeric bZIP transcription factor c-Fos-c-Jun bound to DNA. Nature 373, 257–261 (1995).
Rincón, M. & Flavell, R.A. AP-1 transcriptional activity requires both T-cell receptor-mediated and co-stimulatory signals in primary T lymphocytes. EMBO J. 13, 4370–4381 (1994).
Macián, F., López-Rodríguez, C. & Rao, A. Partners in transcription: NFAT and AP-1. Oncogene 20, 2476–2489 (2001).
Kurachi, M. et al. The transcription factor BATF operates as an essential differentiation checkpoint in early effector CD8+ T cells. Nat. Immunol. 15, 373–383 (2014).
Li, P. et al. BATF-JUN is critical for IRF4-mediated transcription in T cells. Nature 490, 543–546 (2012).
Cippitelli, M. et al. Negative transcriptional regulation of the interferon-γ promoter by glucocorticoids and dominant negative mutants of c-Jun. J. Biol. Chem. 270, 12548–12556 (1995).
Falvo, J.V. et al. Stimulus-specific assembly of enhancer complexes on the tumor necrosis factor-α gene promoter. Mol. Cell. Biol. 20, 2239–2247 (2000).
Oyake, T. et al. Bach proteins belong to a novel family of BTB-basic leucine zipper transcription factors that interact with MafK and regulate transcription through the NF-E2 site. Mol. Cell. Biol. 16, 6083–6095 (1996).
Roychoudhuri, R. et al. BACH2 represses effector programs to stabilize Treg-mediated immune homeostasis. Nature 498, 506–510 (2013).
Muto, A. et al. Bach2 represses plasma cell gene regulatory network in B cells to promote antibody class switch. EMBO J. 29, 4048–4061 (2010).
Muto, A. et al. The transcriptional programme of antibody class switching involves the repressor Bach2. Nature 429, 566–571 (2004).
Hu, G. & Chen, J. A genome-wide regulatory network identifies key transcription factors for memory CD8+ T-cell development. Nat. Commun. 4, 2830 (2013).
Kaech, S.M. et al. Selective expression of the interleukin 7 receptor identifies effector CD8 T cells that give rise to long-lived memory cells. Nat. Immunol. 4, 1191–1198 (2003).
Hikono, H. et al. Activation phenotype, rather than central- or effector-memory phenotype, predicts the recall efficacy of memory CD8+ T cells. J. Exp. Med. 204, 1625–1636 (2007).
Boyman, O., Cho, J.H., Tan, J.T., Surh, C.D. & Sprent, J. A major histocompatibility complex class I-dependent subset of memory phenotype CD8+ cells. J. Exp. Med. 203, 1817–1825 (2006).
Hendriks, J., Xiao, Y. & Borst, J. CD27 promotes survival of activated T cells and complements CD28 in generation and establishment of the effector T cell pool. J. Exp. Med. 198, 1369–1380 (2003).
Kaech, S.M. & Wherry, E.J. Heterogeneity and cell-fate decisions in effector and memory CD8+ T cell differentiation during viral infection. Immunity 27, 393–405 (2007).
Rutishauser, R.L. et al. Transcriptional repressor Blimp-1 promotes CD8+ T cell terminal differentiation and represses the acquisition of central memory T cell properties. Immunity 31, 296–308 (2009).
Cui, W. & Kaech, S.M. Generation of effector CD8+ T cells and their conversion to memory T cells. Immunol. Rev. 236, 151–166 (2010).
Opferman, J.T. et al. Development and maintenance of B and T lymphocytes requires antiapoptotic MCL-1. Nature 426, 671–676 (2003).
Sandelin, A., Alkema, W., Engström, P., Wasserman, W.W. & Lenhard, B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res. 32, D91–D94 (2004).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
Shnyreva, M. et al. Evolutionarily conserved sequence elements that positively regulate IFN-γ expression in T cells. Proc. Natl. Acad. Sci. USA 101, 12622–12627 (2004).
Barr, R.K., Kendrick, T.S. & Bogoyevitch, M.A. Identification of the critical features of a small peptide inhibitor of JNK activity. J. Biol. Chem. 277, 10987–10997 (2002).
Dong, C. et al. JNK is required for effector T-cell function but not for T-cell activation. Nature 405, 91–94 (2000).
Thiel, G., Lietz, M. & Hohl, M. How mammalian transcriptional repressors work. Eur. J. Biochem. 271, 2855–2862 (2004).
Russ, B.E. et al. Distinct epigenetic signatures delineate transcriptional programs during virus-specific CD8+ T cell differentiation. Immunity 41, 853–865 (2014).
Yoshida, C. et al. Bcr-Abl signaling through the PI-3/S6 kinase pathway inhibits nuclear translocation of the transcription factor Bach2, which represses the antiapoptotic factor heme oxygenase-1. Blood 109, 1211–1219 (2007).
Calleja, V., Laguerre, M., Parker, P.J. & Larijani, B. Role of a novel PH-kinase domain interface in PKB/Akt regulation: structural mechanism for allosteric inhibition. PLoS Biol. 7, e17 (2009).
Macintyre, A.N. et al. Protein kinase B controls transcriptional programs that direct cytotoxic T cell fate but is dispensable for T cell metabolism. Immunity 34, 224–236 (2011).
Kim, E.H. et al. Signal integration by Akt regulates CD8 T cell effector and memory differentiation. J. Immunol. 188, 4305–4314 (2012).
Crompton, J.G. et al. Akt inhibition enhances expansion of potent tumor-specific lymphocytes with memory cell characteristics. Cancer Res. 75, 296–305 (2015).
Muto, A. et al. Activation of Maf/AP-1 repressor Bach2 by oxidative stress promotes apoptosis and its interaction with promyelocytic leukemia nuclear bodies. J. Biol. Chem. 277, 20724–20733 (2002).
Gray, S., Szymanski, P. & Levine, M. Short-range repression permits multiple enhancers to function autonomously within a complex promoter. Genes Dev. 8, 1829–1838 (1994).
Glasmacher, E. et al. A genomic regulatory element that directs assembly and function of immune-specific AP-1-IRF complexes. Science 338, 975–980 (2012).
Okkenhaug, K. Signaling by the phosphoinositide 3-kinase family in immune cells. Annu. Rev. Immunol. 31, 675–704 (2013).
Cantrell, D. Protein kinase B (Akt) regulation and function in T lymphocytes. Semin. Immunol. 14, 19–26 (2002).
Kim, D. & Salzberg, S.L. TopHat-Fusion: an algorithm for discovery of novel fusion transcripts. Genome Biol. 12, R72 (2011).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Acknowledgements
We thank S.A. Rosenberg, K. Hanada, K. Hirahara, K. Mousavi, H. Zare, V. Sartorelli, N. Van Panhuys, S. Kerkar and A. Restifo for ideas and discussion, A. Mixon and S. Farid for expertise with cell sorting, members of the NHLBI sequencing core facility for help with sequencing, L. Samsel for help with ImageStream imaging flow cytometry and G. McMullen for expertise with mouse handling. Supported by the Intramural Research Programs of the NCI and NHLBI, Wellcome Trust/Royal Society grant 105663/Z/14/Z (R.R.) and UK Biotechnology and Biological Sciences Research Council grant BB/N007794/1 (R.R. and K.O.).
Author information
Authors and Affiliations
Contributions
R.R., D.C. and N.P.R. wrote the manuscript and designed experiments. R.R., D.C., K.M.Q., Y.J., Z.Y., J.H.P., Y.K., Y.W., L.G. and G.F. performed experiments. P.L. and R.R. analyzed bioinformatic data. C.A.K., D.C.P., D.C.M., M.S., S.J.P., H.-Y.S., R.S., A.M., L.G., R.L.E., J.Z., K.O., J.J.O., K.I. and W.J.L. edited the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cell-intrinsic function of BACH2 in CD8+ T cells.
a-c, Mice were reconstituted with mixtures of congenically distinct Lin− WT and KO bone marrow cells (BM) (a) and CD44 and CD62L expression was measured on the surface of WT and KO CD4+ and CD8+ T cells 2 months following reconstitution (b). Replicate measurements (c) of the proportions of naïve (NAI; CD44+ CD62L+), central memory (CM; CD44+ CD62L+) and effector (EFF; CD44+ CD62L−) cells are shown. d, Experimental schema. Congenically distinct naïve WT Ly5.1+ and KO Thy-1.1+ OT-I TCR-transgenic CD8+ T cells were sorted from mixed BM chimeric mice reconstituted with WT and KO OT-I BM. Cells were mixed at ~1:1 ratios and transferred into naïve C57BL/6 recipient mice prior to infection with VV-OVA (2x10-6 pfu) and kinetic analysis. Numbers in gates indicate percentages. Bars and error (c) represent mean and s.e.m. **P<0.01. Data are representative of 3 independently repeated experiments.
Supplementary Figure 2 Phenotypic analysis of WT and Bach2−/– CD8+ T cells responding to viral infection.
a, Absolute number of KLRG1+ WT and KO CD8+ T cells at indicated timepoints following mixed transfer of naïve sorted WT and KO OT-I CD8+ T cells into recipient mice and infection with VV-OVA. b, Expression of CD127 on the surface of gated CD62L− KLRG1− OT-I CD8+ T cells at day 7 following infection with VV-OVA. Representative flow cytometry (left) and replicate measurements (right) are shown. c-d, Expression of CD43 (c) and CD27 (d) on the surface of WT and KO cells at day 7 post-infection. Representative flow cytometry (left) and replicate measurements (right) are shown. Bars and error (b-d) mean and s.e.m. *P<0.05; **P<0.01; ****P<0.001.
Supplementary Figure 3 Increased activation of Bach2−/– cells is antigen dose dependent and proliferation independent.
a, CD44 and CD62L expression 4 days after stimulation of naïve WT and KO OT-I CD8+ T cells with indicated concentrations of cognate peptide ligand (SL9) in the presence of congenically distinct feeder splenocytes. Numbers in gates indicate percentages. Data are representative of 2 independently repeated experiments. Naïve cells for assays were isolated by flow cytometric sorting. b, CFSE dilution and CD62L expression amongst CFSE-labelled WT and KO naïve CD8+ T cells stimulated in vitro for 3 days. c, Expression of CD69 and CD25 at day 4 following in vitro stimulation of naïve WT and KO CD8+ T cells with platebound anti-CD3 and anti-CD28 in the presence of IL-2. d, CFSE dilution and Annexin V staining on CFSE-labelled WT and KO naïve CD8+ T cells stimulated in vitro for 3 days with platebound anti-CD3 and anti-CD28 antibodies in the presence of 100 IU IL-2. Numbers in gates indicate percentages. e, Naïve WT and KO CD8+ T cells were stimulated with anti-CD3 and anti-CD28 antibodies in the presence of 100IU IL-2 and harvested for analysis at day 4. Expression of the indicated proteins was measured by SDS-PAGE and western blotting. Data are representative of 2 independently repeated experiments.
Supplementary Figure 4 Analysis of global transcriptional differences between WT and Bach2−/– CD8+ T cells at day 7 after infection.
a, Hierarchical clustering analysis of differentially expressed genes (p<0.05; log2 fold-change>1) between WT and KO OT-I cells isolated ex vivo 7 days following VV-OVA infection. Color scale shows FPKM values normalized to row maxima as indicated. b, Geneset enrichment analysis of the indicated geneset in global transcriptional differences between KO and WT cells. Statistical significance was tested using a weighted Kolmogorov–Smirnov test. FPKM values from 3 replicate RNA-Seq measurements per experimental condition are shown. c, Hierarchical clustering analysis of differentially expressed genes (p<0.05; log2 fold-change>1) between fractionated WT and KO CD62L− KLRG1− effector CD8+ T cells isolated ex vivo at day 7 following VV-OVA infection. Color scale shows average FPKM values normalized to row maxima as indicated. FPKM values from 2 replicate RNA-Seq measurements per experimental condition are shown.
Supplementary Figure 5 Analysis of JunD and BACH2 binding in d5 in vitro–activated CD8+ T cells.
a, Naïve CD8+ T cells were isolated by flow cytometric sorting from spleens of wildtype animals and stimulated with plate-bound anti-CD3 and anti-CD28 antibodies in the presence of IL-2 for 2 days followed by 3 further days of culture to generate d5 in vitro activated CD8+ T cells. Protein expression was measured at indicated timepoints following stimulation using SDS-PAGE and immunoblotting. Quantification of BACH2 abundance normalized to β-actin is shown. BACH2 expression in day 5 in vitro activated CD8+ T cells is ~25% of that in naïve cells. b, Histogram of JunD binding, centered around JunD peaks, in WT and Bach2 KO CD8+ T cells at sites bound exclusively by JunD (left histogram) or sites at which JunD and BACH2 binding sites colocalize (right histogram). A significant increase in JunD binding is observed in Bach2-deficient cells only at sites where BACH2 and JunD binding sites colocalize. c-d, Analysis of exclusive JunD binding in Bach2-deficient cells. Pie chart showing JunD binding sites in Bach2-deficient cells (c). A majority of sites (blue) were found to be shared with WT cells, while a minority of sites (red) were only present in Bach2 KO cells. Alignments of JunD binding in WT and Bach2 KO cells at loci where exclusive JunD binding in Bach2 KO cells is detected (d). Arrows indicate loci at which JunD peaks only pass a low significance threshold for peak calling in Bach2-deficient cells (p<1x10-3; Binomial test).
Supplementary Figure 6 Effect of BACH2 on chromatin accessibility and occupancy of Jun family AP-1 factors.
a, Differences in JunD binding in WT and KO d5 in vitro activated CD8+ T cells do not correlate with differences in chromatin accessibility. Scatterplot comparing differences in JunD binding (y-axis) with differences in ATAC-Seq signal around BACH2 binding sites in WT and Bach2−/– d5 in vitro activated CD8+ T cells. Statistical significance was evaluated using the two-sample Kolmogorov-Smirnov test. b, Effect of BACH2 on binding of c-Jun, JunB and JunD. Enrichment of BACH2, c-Jun, JunB and JunD at indicated loci in d5 in vitro activated KO and WT CD8+ T cells relative to input. Enrichment values are normalized to WT cells. Bars and error (b) represent mean and s.e.m. of replicate measurements. *P<0.05; **P<0.01; ***P<0.005; ****P<0.001 (Student’s t-test).
Supplementary Figure 7 Expression of Bach2 mRNA and the encoded protein within CD8+ T cells isolated ex vivo after viral infection.
a-b, Naïve CD45.2+ OT-1 CD8+ T cells were isolated by flow cytometric sorting and pelleted for analysis or transferred into congenically distinct CD45.1+ hosts. Recipient mice were infected with rVV-OVA and Ly5.2+ CD8+ T cells were sorted by flow cytometric sorting at the indicated timepoints following infection. Bach2 mRNA was measured in isolated cells relative to Actb (a) or the abundance of BACH2 and β-actin was assessed in whole cell lysates by western blot (b). Data are representative of two independently repeated experiments (a). Bars and error (a) represent mean and s.e.m. **P<0.01; ***P<0.005; ****P<0.001.
Supplementary Figure 8 Suppression of effector differentiation by pharmacological inhibition of Akt is partially dependent on BACH2.
a-b, Naïve CD44− CD62L+ WT and KO CD8+ T cells (a) were stimulated in the presence of 100IU rhIL-2 in the presence or absence of 1μM AKTi. Expression of IFN-γ and TNF was measured 4 days after stimulation by intracellular cytokine staining. Numbers in gates indicate percentages. Bars and error represent mean and s.e.m. ***P<0.005; ****P<0.001. Data (a-b) are representative of 2 independently repeated experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–11 (PDF 4195 kb)
Rights and permissions
About this article
Cite this article
Roychoudhuri, R., Clever, D., Li, P. et al. BACH2 regulates CD8+ T cell differentiation by controlling access of AP-1 factors to enhancers. Nat Immunol 17, 851–860 (2016). https://doi.org/10.1038/ni.3441
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3441
This article is cited by
-
Integration of ATAC-Seq and RNA-Seq reveals FOSL2 drives human liver progenitor-like cell aging by regulating inflammatory factors
BMC Genomics (2023)
-
Reprogramming T cell differentiation and exhaustion in CAR-T cell therapy
Journal of Hematology & Oncology (2023)
-
The landscape of PBMC methylome in canine mammary tumors reveals the epigenetic regulation of immune marker genes and its potential application in predicting tumor malignancy
BMC Genomics (2023)
-
Aging-associated HELIOS deficiency in naive CD4+ T cells alters chromatin remodeling and promotes effector cell responses
Nature Immunology (2023)
-
Single-cell CRISPR screens in vivo map T cell fate regulomes in cancer
Nature (2023)