Abstract
Chromatin-associated fumarase (FH) affects histone methylation via its metabolic activity. However, whether this effect is involved in gene transcription remains to be clarified. In this study, we show that under glucose deprivation conditions, AMPK phosphorylates FH at Ser75, which in turn forms a complex with ATF2 and participates in promoter activation. FH-catalysed fumarate in promoter regions inhibits KDM2A demethylase activity, and thus maintains the H3K36me2 profile and facilitates gene expression for cell growth arrest. On the other hand, FH is found to be O-GlcNAcylated at the AMPK phosphorylation site; FH–ATF2-mediated downstream events are impeded by FH O-GlcNAcylation, especially in cancer cells that display robust O-GlcNAc transferase (OGT) activity. Consistently, the FH-Ser75 phosphorylation level inversely correlates with the OGT level and poor prognosis in pancreatic cancer patients. These findings uncover a previously uncharacterized mechanism underlying transcription regulation by FH and the linkage between dysregulated OGT activity and growth advantage of cancer cells under glucose deficiency.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Schulze, A. & Harris, A. L. How cancer metabolism is tuned for proliferation and vulnerable to disruption. Nature 491, 364–373 (2012).
The American Association for Cancer Research Human Epigenome Task Force & European Union, Network of Excellence, Scientific Advisory Board. Moving AHEAD with an international human epigenome project. Nature 454, 711–715 (2008).
Boukouris, A. E., Zervopoulos, S. D. & Michelakis, E. D. Metabolic enzymes moonlighting in the nucleus: metabolic regulation of gene transcription. Trends Biochem. Sci. 41, 712–730 (2016).
Sciacovelli, M. et al. Fumarate is an epigenetic modifier that elicits epithelial-to-mesenchymal transition. Nature 537, 544–547 (2016).
Buren, S. et al. Regulation of OGT by URI in response to glucose confers c-MYC-dependent survival mechanisms. Cancer Cell 30, 290–307 (2016).
Sharma, U. & Rando, O. J. Metabolic inputs into the epigenome. Cell Metab. 25, 544–558 (2017).
Kornberg, H. L. & Krebs, H. A. Synthesis of cell constituents from C2-units by a modified tricarboxylic acid cycle. Nature 179, 988–991 (1957).
Yogev, O. et al. Fumarase: a mitochondrial metabolic enzyme and a cytosolic/nuclear component of the DNA damage response. PLoS Biol. 8, e1000328 (2010).
Jiang, Y. et al. Local generation of fumarate promotes DNA repair through inhibition of histone H3 demethylation. Nat. Cell Biol. 17, 1158–1168 (2015).
Lees-Miller, S. P. Fumarate in DNA repair. Nat. Cell Biol. 17, 1096–1097 (2015).
Gozdecka, M. & Breitwieser, W. The roles of ATF2 (activating transcription factor 2) in tumorigenesis. Biochem. Soc. Trans. 40, 230–234 (2012).
Yu, T. et al. The regulatory role of activating transcription factor 2 in inflammation. Mediat. Inflamm. 2014, 950472 (2014).
Lau, E. & Ronai, Z. A. ATF2—at the crossroad of nuclear and cytosolic functions. J. Cell Sci. 125, 2815–2824 (2012).
Berger, A. J. et al. Subcellular localization of activating transcription factor 2 in melanoma specimens predicts patient survival. Cancer Res. 63, 8103–8107 (2003).
Maekawa, T. et al. Reduced levels of ATF-2 predispose mice to mammary tumors. Mol. Cell Biol. 27, 1730–1744 (2007).
Bhoumik, A. et al. Suppressor role of activating transcription factor 2 (ATF2) in skin cancer. Proc. Natl Acad. Sci. USA 105, 1674–1679 (2008).
Gozdecka, M. et al. JNK suppresses tumor formation via a gene-expression program mediated by ATF2. Cell Rep. 9, 1361–1374 (2014).
Efeyan, A., Comb, W. C. & Sabatini, D. M. Nutrient-sensing mechanisms and pathways. Nature 517, 302–310 (2015).
Nagel, A. K. & Ball, L. E. O-GlcNAc transferase and O-GlcNAcase: achieving target substrate specificity. Amino Acids 46, 2305–2316 (2014).
Hardiville, S. & Hart, G. W. Nutrient regulation of gene expression by O-GlcNAcylation of chromatin. Curr. Opin. Chem. Biol. 33, 88–94 (2016).
Ferrer, C. M., Sodi, V. L. & Reginato, M. J. O-GlcNAcylation in cancer biology: linking metabolism and signaling. J. Mol. Biol. 428, 3282–3294 (2016).
Baldini, S. F. & Lefebvre, T. O-GlcNAcylation and the metabolic shift in high-proliferating cells: all the evidence suggests that sugars dictate the flux of lipid biogenesis in tumor processes. Front. Oncol. 6, 6 (2016).
Ferrer, C. M. & Reginato, M. J. Sweet connections: O-GlcNAcylation links cancer cell metabolism and survival. Mol. Cell Oncol. 2, e961809 (2015).
Taylor, R. P. et al. Glucose deprivation stimulates O-GlcNAc modification of proteins through up-regulation of O-linked N-acetylglucosaminyltransferase. J. Biol. Chem. 283, 6050–6057 (2008).
Klaus, A. et al. A two-dimensional screen for AMPK substrates identifies tumor suppressor fumarate hydratase as a preferential AMPKalpha2 substrate. J. Proteomics 75, 3304–3313 (2012).
Steentoft, C. et al. Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J. 32, 1478–1488 (2013).
Xiao, M. et al. Inhibition of alpha-KG-dependent histone and DNA demethylases by fumarate and succinate that are accumulated in mutations of FH and SDH tumor suppressors. Genes Dev. 26, 1326–1338 (2012).
Tsukada, Y. et al. Histone demethylation by a family of JmjC domain-containing proteins. Nature 439, 811–816 (2006).
Olive, K. P. et al. Inhibition of Hedgehog signaling enhances delivery of chemotherapy in a mouse model of pancreatic cancer. Science 324, 1457–1461 (2009).
Kamphorst, J. J. et al. Human pancreatic cancer tumors are nutrient poor and tumor cells actively scavenge extracellular protein. Cancer Res. 75, 544–553 (2015).
Carnero, A. & Lleonart, M. The hypoxic microenvironment: a determinant of cancer stem cell evolution. Bioessays 38, S65–S74 (2016).
Ma, Z., Vocadlo, D. J. & Vosseller, K. Hyper-O-GlcNAcylation is anti-apoptotic and maintains constitutive NF-κB activity in pancreatic cancer cells. J. Biol. Chem. 288, 15121–15130 (2013).
Perera, R. M. et al. Transcriptional control of autophagy-lysosome function drives pancreatic cancer metabolism. Nature 524, 361–365 (2015).
Krall, A. S. & Christofk, H. R. Cancer: a metabolic metamorphosis. Nature 496, 38–40 (2013).
Phoomak, C. et al. High glucose levels boost the aggressiveness of highly metastatic cholangiocarcinoma cells via O-GlcNAcylation. Sci. Rep. 7, 43842 (2017).
Balbach, S. T. & Orkin, S. H. An Achilles’ heel for MLL-rearranged leukemias: writers and readers of H3 lysine 36 dimethylation. Cancer Discov. 6, 700–702 (2016).
Tanaka, Y. et al. Mild glucose starvation induces KDM2A-mediated H3K36me2 demethylation through AMPK to reduce rRNA transcription and cell proliferation. Mol. Cell Biol. 35, 4170–4184 (2015).
Tanaka, Y. et al. JmjC enzyme KDM2A is a regulator of rRNA transcription in response to starvation. EMBO J. 29, 1510–1522 (2010).
Gut, P. & Verdin, E. The nexus of chromatin regulation and intermediary metabolism. Nature 502, 489–498 (2013).
Zheng, Y. et al. Secreted and O-GlcNAcylated MIF binds to the human EGF receptor and inhibits its activation. Nat. Cell Biol. 17, 1348–1355 (2015).
Lu, Z. et al. Activation of protein kinase C triggers its ubiquitination and degradation. Mol. Cell Biol. 18, 839–845 (1998).
Xia, Y. et al. c-Jun downregulation by HDAC3-dependent transcriptional repression promotes osmotic stress-induced cell apoptosis. Mol. Cell 25, 219–232 (2007).
Hayakawa, J. et al. Identification of promoters bound by c-Jun/ATF2 during rapid large-scale gene activation following genotoxic stress. Mol. Cell 16, 521–535 (2004).
Buckel, L. et al. Tumor radiosensitization by monomethyl auristatin E: mechanism of action and targeted delivery. Cancer Res. 75, 1376–1387 (2015).
Acknowledgements
We thank X.-y. Liu and K.-J. Zhang at the Chinese Academy of Sciences for mice study assistance, and Z.-M. Lu at MD Anderson for his critical reading of this manuscript. This work was supported by the National Nature Science Foundation of China 81572722 (Y.J.), Shanghai Committee of Science and Technology 16QA1403200 (Y.J.), Shanghai Municipal Education Commission-Gaofeng Clinical Medicine Grant 20161319 (Y.J.), Program for eastern young scholar at Shanghai institutions of higher learning (Y.J.), National Nature Science Foundation of China 81602042 (T.W.), and National Nature Science Foundation of China 81672710 (Q.Y.).
Author information
Authors and Affiliations
Contributions
This study was conceived by Y.J.; Y.J. and T.W. designed the study; T.W., Q.Y., J.L., B.H., Q.Z., C.M., L.Z. and H.F. performed experiments; W.H., L.L. and Y.E.C. provided reagents and conceptual advice; Y.J. wrote the paper with comments from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
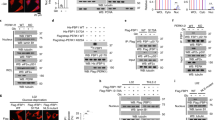
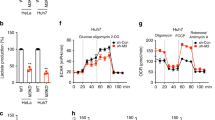
Supplementary Figure 1 Glucose deficiency induced FH-ATF2 interaction. In b–m, immunoblotting or immunoprecipitation analyses were performed using the indicated antibodies.
Data represent 1 out of 3 experiments. (a) HPDE cells expressing Flag–FH were cultured for 6 h in the glucose-free DMEM; cellular extracts subjected to immunoprecipitation with an anti-Flag antibody were analyzed by mass spectrometry. FH-interacting proteins identified by mass spectrometry were shown. (b) HPDE cells were cultured for 6 h in the DMEM with indicated concentrations of glucose (left panel). HPDE cells were cultured for 6 h in the glucose-free DMEM (right panel). (c) SW1990 cells were cultured for 6 h in the glucose-free DMEM. (d) PANC-1 (upper panel) and SW1990 cells (bottom panel) pre-treated with Compound C for 1 h were cultured for 6 h in the glucose-free DMEM. (e) HPDE cells were cultured for 6 h in the glutamine-free or glucose-free DMEM. (f,g) HPDE cells (f) and PANC-1 cells (g) were cultured for various length of time in the glucose-free DMEM. (h) PANC-1 cells were expressed with OGT shRNA and reconstituted with or without the expression of RNAi-resistant Myc-rOGT. Cells were cultured for 6 h in the glucose-free DMEM. (i) SW1990 cells expressing Flag–FH with or without OGT shRNA expression were cultured for 6 h in the glucose-free DMEM. (j) HPDE and PANC-1, and SW1990 cells were expressed with or without OGT shRNA. (k) HPDE cells were cultured for 6 h in the glucose-free DMEM. Cytosolic and nuclear fractions were prepared. (l) PANC-1 (left panel) and SW1990 cells (right panel) were expressed with OGT shRNA, and the cell lines with distinct efficiency of OGT depletion were isolated. Cell death rate was examined by PI staining and FACS analyses. PANC-1 and SW1990 cell lines with moderate silence of OGT were selected (shown as ‘P2’ in PANC-1 and ‘S1’ in SW1990 cells). The values are presented as mean ± s.e.m. (n = 3 independent experiments). ∗ represents P < 0.05 and ∗∗ represents P < 0.01 (Student’s t-test) between indicated groups (see Supplementary Table 1 for source data). (m) PANC-1 were transfected with or without the siRNA against AMPKα.
Supplementary Figure 2 FH is required for cell growth arrest under glucose deprivation.
In a,c and d, the values are presented as mean ± s.e.m. (n = 3 independent experiments). ∗ represents P < 0.05 and ∗∗ represents P < 0.01 (Student’s t-test) between indicated groups (see Supplementary Table 1 for source data). (a) SW1990 cells with or without OGT shRNA and FH shRNA expression, were cultured for 24 h under glucose deficiency condition (0.5 mM). mRNA levels of ATF2 targeted genes were analyzed by real-time PCR. (b) PANC-1 cells with or without OGT depletion were expressed with a vector for control shRNA or FH shRNA and ATF2 shRNA. Representative images of FACS analyses for BrdU positive cells were shown. The ratio of the right peak value to the total value positively correlates cell proliferation rate. Cell death rate was examined by PI staining and FACS analyses. (c,d) SW1990 cells with or without OGT depletion were expressed with a vector for control shRNA or FH shRNA and ATF2 shRNA. Cellular proliferation rate was examined by BrdU incorporation assay and FACS analyses (c). Cell death rate was examined by PI staining and FACS analyses (d).
Supplementary Figure 3 AMPK-mediated FH phosphorylation is required for FH-ATF2 interaction.
In a–j, immunoblotting, immunoprecipitation analyses (a–i) and ChIP (j and k) analyses were performed using the indicated antibodies. Data represent 1 out of 3 experiments. In j–m, the values are presented as mean ± s.e.m. (n = 3 independent experiments). ∗ represents P < 0.05 and ∗∗ represents P < 0.01 (Student’s t-test) between indicated groups (see Supplementary Table 1 for source data). (a) In vitro phosphorylation analyses were performed by mixing the purified AMPK complex with the indicated purified His-ATF2 proteins in the presence of [γ-32P]ATP. (b) HPDE cells expressing Flag-FH were cultured for 6 h in the glucose-free DMEM. Immunoprecipitation analysis was performed using the Flag antibody, and the extracts were analyzed by mass spectrometry. The results of a mass spectrometric analysis of a tryptic fragment at m/z 404.16 (mass error, −3.873 ppm) matched those of the doubly charged peptide 111-2-120, suggesting that S75 was phosphorylated. The Sequest score for this match was Xcorr = 2.233. The Mascot score was 281.71. (c) HPDE cells were pretreated with Compound C (left panel) or transfected with or without the siRNA against AMPKα (right panel). Cells were cultured for 6 h in the glucose-free DMEM. (d) PANC-1 (left panel) and SW1990 cells (right panel) expressed with or without OGT shRNA were cultured for 6 h in the glucose-free DMEM. (e,f) HPDE cells with expression of indicated Flag–FH (e), or SW1990 cells with depletion of OGT and expression of indicated Flag–FH (f) were cultured for 6 h in the glucose-free DMEM. Immunoprecipitation analyses were performed. (g) PANC-1 cells were expressed with a vector for control shRNA or FH shRNA and reconstituted with expression of rFH(N)WT or rFH(N) S75A. (h) SW1990 cells were expressed with a vector for control shRNA or FH shRNA and reconstituted with expression of rFH(N) WT or rFH(N) S75A. (i,j) PANC-1 cells with depleted OGT, FH and reconstituted expression of rFH(N)WT or rFH(N) S75A were cultured for 6 h in the glucose-free DMEM. Immunoprecipitation analyses were performed (i). ChIP analyses were performed. The primers covering ATF2 binding site of c-Jun gene promoter region were used for the real-time PCR. The Y axis shows the value normalized to the input (j). (k) SW1990 cells with depleted OGT, FH and reconstituted expression of WT FH or FH S75A were cultured for 6 h in the glucose-free DMEM. ChIP analyses were performed. The primers covering ATF2 binding site of c-Jun gene promoter region were used for the real-time PCR. The Y axis shows the value normalized to the input. (l,m) SW1990 cells with depleted OGT, FH and reconstituted expression of indicated Flag–rFH(N) were cultured for 24 h in the DMEM with low glucose (0.5 mM). mRNA levels of ATF2 targeted genes were analyzed by real–time PCR (l). Cellular proliferation rate were examined by BrdU incorporation assay and analyzed by FACS analyses (m).
Supplementary Figure 4 FH was O-GlcNAcylated at phosporylation site of AMPK.
(a) HPDE cells expressing Flag-FH were treated with or without PUGNAc (20 μM) for 24 h. Immunoprecipitation analyses and immunoblotting analyses were performed using the indicated antibodies. Data represent 1 out of 3 experiments (see Supplementary Table 1 for source data). (b) NetOGlyc. 4-software analysis of O-GlcNAcylation site within FH amino acid sequence. (c) Flag-FH was expressed in PANC-1 cells. Immunoprecipitation analysis was performed using the Flag antibody, and the extracts were analyzed by mass spectrometry. The results of a mass spectrometric analysis of a tryptic fragment at m/z 404.16 (mass error, −2.835 ppm) matched those of the doubly charged peptide 111-2-120, suggesting that S75 was O-GlcNAcylated. The Sequest score for this match was Xcorr = 1.486. The Mascot score was 907.59. (d) PANC-1 cells were expressed with indicated Flag-FH. The O-GlcNAc modified proteins modified by azide were labeled with biotin and isolated with streptavidin beads for immunoblotting analyses. Data represent 1 out of 3 experiments.
Supplementary Figure 5 The local activity of FH at promoter region promotes ATF2-mediated cell proliferation arrest.
In a–b, immunoblotting analyses were performed using the indicated antibodies. Data represent 1 out of 3 experiments. In c–i, the values are presented as mean ± s.e.m. (n = 3 independent experiments). ∗ represents P < 0.05 and ∗∗ represents P < 0.01 (Student’s t-test) between indicated groups (see Supplementary Table 1 for source data). (a) PANC-1 cells with depleted OGT, FH and reconstituted expression of indicated FH proteins were analyzed by immunoblotting. (b) HPDE cells with FH depletion and reconstituted expression of indicated FH proteins were analyzed by immunoblotting. (c) PANC-1 cells with depleted OGT, FH and reconstituted with expression of the indicated FH proteins, and were incubated with the indicated concentrations of monoethyl-fumarate. The abundance of intracellular fumarate was measured. (d–f) HPDE cells with or without ATF2 and FH depletion, were reconstituted with expression of the indicated FH proteins. Cells were cultured for 24 h in the DMEM with low glucose (0.5 mM) and indicated concentration of monomethyl fumarate. The abundance of intracellular fumarate was measured (d). The mRNA level of ATF3 was analyzed by realtime-PCR (e). Cellular proliferation rate were examined by BrdU incorporation assay and FACS (f). (g–i) PANC-1 cells with depletion of OGT, FH and reconstituted expression of indicated Flag–rFH(N) were cultured for 24 h in the DMEM with low glucose (0.5 mM) and indicated concentration of diethyl-malate. The abundance of intracellular fumarate was measured (g). The mRNA level of ATF3 was analyzed by realtime-PCR (h). Cellular proliferation rate were examined by BrdU incorporation assay and FACS (i).
Supplementary Figure 6 Fumarate produced at promoter region inhibits H3K36me2 demethylation.
In a and b, the values are presented as mean ± s.e.m. (n = 3 independent experiments) (see Supplementary Table 1 for source data). (a) PANC-1 cells with depleted OGT were cultured for 6 h in the absence of glucose. ChIP analyses were performed by using antibodies for H3K36me2, H3K9me2, H3K9me3, and H3K27me2. The primers covering ATF2 binding site of c-Jun gene promoter region were used for the real-time PCR. The Y axis shows the fold of the value without glucose normalized to the value with glucose. (b) PANC-1 cells with depleted endogenous FH and expressing WT or indicated mutant Flag–rFH(N) and OGT shRNA were cultured for 6 h in the absence of glucose. ChIP analyses were performed by using KDM2A antibody. The primers covering ATF2 binding site of c-Jun gene promoter region were used for the realtime-PCR. The y axis shows the value normalized to the input. (c) PANC-1 cells with depleted OGT were overexpressed with or without KDM2A in presence or absence of monoethyl-fumarate (5 mM) addition. Immunoblotting analyses were performed.
Supplementary Figure 7 Inhibition of FH S75 phosphorylation by OGT is required for tumorigenesis.
The FH S75 phosphorylation antibody specificities were validated using IHC analyses with specific blocking FH S75 phosphorylation-peptides. Scale bars: 50 μm.
Supplementary information
Supplementary Information
Supplementary Information (PDF 4118 kb)
Supplementary Table 1
Supplementary Information (XLSX 26 kb)
Rights and permissions
About this article
Cite this article
Wang, T., Yu, Q., Li, J. et al. O-GlcNAcylation of fumarase maintains tumour growth under glucose deficiency. Nat Cell Biol 19, 833–843 (2017). https://doi.org/10.1038/ncb3562
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3562
This article is cited by
-
Metabolic reprogramming regulated by TRAF6 contributes to the leukemia progression
Leukemia (2024)
-
Protein post-translational modifications in the regulation of cancer hallmarks
Cancer Gene Therapy (2023)
-
Regulation of tumor metabolism by post translational modifications on metabolic enzymes
Cancer Gene Therapy (2023)
-
O-GlcNAcylation of MORC2 at threonine 556 by OGT couples TGF-β signaling to breast cancer progression
Cell Death & Differentiation (2022)
-
O-GlcNAcylation suppresses TRAP1 activity and promotes mitochondrial respiration
Cell Stress and Chaperones (2022)