Abstract
Somatic cells can be reprogrammed into induced pluripotent stem cells (iPSCs) by ectopic expression of transcription factors Oct4, Sox2, Klf4 and cMyc. Recent advancements have shown that small-molecule compounds can induce pluripotency, indicating that cell fate can be regulated by direct manipulation of intrinsic cell signaling pathways, thereby innovating our current understanding of reprogramming. The fact that lineage specifiers can induce pluripotency suggests that the pluripotent state is a fine balance between competing differentiation forces. Dissection of pluripotent roadmaps indicates that reprogramming is a process of reverse development, involving a series of complicated and distinct reprogramming stages. Evidence from mouse iPSC transplantation studies demonstrated that some certain but not all cells derived from iPSCs are immunogenic. These studies provide new ways to minimize reprogramming-induced abnormalities and maximize reprogramming efficiency to facilitate clinical development and use of iPSCs.
Similar content being viewed by others
Facts
-
Reprogramming is not only a simple process of reverse development but also a very complicated procedure with different reprogramming stages.
-
Binding of both facilitators and inhibitors by reprogramming factors simultaneously at early stage directly contributes to low reprogramming efficiency.
-
The fact that lineage specifiers can induce iPSC production suggests that pluripotency is a balance between competing differentiation forces.
-
Induced pluripotency by small-molecule compounds indicates that cell fate decision is regulated by intracellular signaling pathways.
-
Evidence from different research groups supports that some certain but not all cells derived from iPSCs are immunogenic.
Open Questions
-
What are the critical molecular events involved in reprogramming of somatic cells to iPSCs?
-
How do reprogramming factors orchestrate such a complicated de-differentiation process?
-
Are there any ways to selectively activate facilitator genes without co-activation of inhibitory genes during initiation of reprogramming?
-
How do the complicated signal transduction networks inside a cell control its fate?
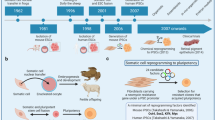
The concept of totipotent differentiated vertebrate cells was first proposed by the German embryologist Spemann in 1938, who reported that the nucleus from an embryo retained the ability to develop into a salamander after undergoing four divisions.1 In 1952, Briggs and King2 successfully generated tadpoles by transferring a cell nucleus derived from an embryo in the blastocyst stage into an enucleated oocyte using a technique called somatic cell nuclei transfer (SCNT; Figure 1). Using this breakthrough technology, Briggs and King3 tried to determine whether aging cells are still totipotent and found that as cells develop they become more difficult to clone. Gurdon extended these experiments by using nuclei from matured intestinal and keratinized skin cells of frogs as donors.4, 5, 6 His research indicated that differentiated and even matured cells do indeed retain the genetic information needed to develop into a life, and the cytoplasm of oocytes contains certain factors that can reprogram mature nuclei into pluripotency.
Different strategies to induce pluripotency. (a) Somatic cell (SC) nuclei transfer (SCNT). Pluripotency can be achieved by transfer of somatic nuclei to enucleated oocytes. (b) Cell fusion. Fusion of ESCs with SCs generates pluripotent fusion hybrids. (c) Transcription factor induction. Ectopic expression of four transcription factors (Oct4, Sox2, Klf4 and cMyc) in SCs can reverse them to an ESC-like state; these cells are called iPSCs. (d) Small-molecule treatment. SC treatment with a combination of small molecules (Valproic acid, Tranvlcypromine, 616452, CHIR, Forskolin, TTNPB, DZNep) can reprogram differentiated cells into iPSCs
By fusion of mouse pluripotent embryonic carcinoma cells with thymocytes, Miller and Ruddle7 generated hybrids that form carcinomas after transplantation into nude mice, indicating that the differentiated thymocytes were reprogrammed into a pluripotent state. Two further reports showed that fusion of embryonic stem cells (ESCs) with somatic cells resulted in the formation of pluripotent hybrids, with characteristics similar to parental ESCs (Figure 1).8, 9 These proof-of-concept experiments together with successful nuclear reprogramming by SCNT indicate the presence of reprogramming factors in the ESCs and/or oocytes.10
In 2006, by screening a panel of genes specifically expressed in ESCs, Takahashi and Yamanaka11 showed that Oct4, Sox2, Klf4 and cMyc could reprogram mouse somatic cells into pluripotency, called induced pluripotent stem cells (iPSCs; Figure 1). This same molecular cocktail can successfully reprogram human somatic cells to iPSCs.8, 9 Using a similar strategy, Thomson’s group found that Oct4, Sox2, Lin28 and Nanog were able to reprogram human somatic cells into iPSCs.12 Like ESCs, iPSCs can undergo self-renewal and differentiate into all three germ layers in vitro. They can also contribute to chimeric mice and go germ line transmission. Most importantly, iPSCs can directly develop into mice by 4n complementation assay, indicating that iPSCs are totipotent.13, 14, 15
Advancements in Reprogramming Methods
Since the discovery of iPSCs, reprogramming technology has developed rapidly. To date, they are three generation of iPSCs in general. First, iPSCs can be generated by overexpression of transcription factors in target cells infected by retrovirus or lentivirus.11, 12, 16 However, viral integration into the host genome poses serious cancer risks, which significantly hinders the clinical development of this type of iPSC technology.10, 11, 17
To resolve potential cancer risks associated with viral integration, integration-free iPSC production techniques were widely developed as the secondary generation of reprogramming approaches. So far, iPSCs without exogenous DNA integration have been generated using different strategies, including piggyBac transposition,18, 19 episomal vectors,20, 21, 22, 23, 24, 25 and microRNA, which were initially demonstrated to enhance reprogramming efficiency and later used to generate iPSCs.26, 27 Furthermore, by delivering reprogramming proteins directly, mouse and human somatic cells were shown to be successfully reprogrammed but with extremely low efficiency.28, 29, 30 All of these reprogramming methods avoid using viral delivery of transcriptional factors, thereby significantly improving their safety and use in clinical settings. Most recently, a third method to generate iPSCs was successfully developed, by addition of small-molecule compounds into mouse fibroblast cultures, raising hopes of generating human iPSC for therapeutics without tedious genetic manipulations (Figure 1).31
Understanding the Mechanisms of Reprogramming
Induced pluripotency was traditonally achieved by SCNT and cell fusion before the discovery of iPSC. It is now widely accepted that different cells have distinct transcriptional repertoires correlated with their own epigenetic signatures. The identification of master transcription factors, which execute reprogramming, represents significant progress in the understanding of mechanisms of induced pluripotency. However, how reprogramming factors orchestrate epigenetic remodeling is still largely unknown.
Facilitators and inhibitors of efficient reprogramming
iPSCs are generated by forced overexpression of transcription factors in targeted somatic cells, followed by multiple divisions over a long period of time during the reprogramming process. However, SCNT and cell fusion can induce pluripotency in a shorter time by using natural proteins inside the cytoplasm of eggs or ESCs.32 This process is similar to mammalian fertilization: soon after entering the egg, sperm chromatin rapidly switch their conformation and coordinately undergo mitosis together with the chromosomes of eggs, suggesting that the cytosolic proteins of eggs can efficiently change the epigenetic state of sperm. After introduction of a sizeable amount of reprogramming factors (Oct4, Sox2, Klf4 and cMyc; that is, OSKM), somatic cell chromosomes are forced to bind these foreign factors in either ESC physiologically relevant or irrelevant regions.33 The extensive regionally irrelevant binding of OSKM to targeted somatic cell chromatin during initial stages impedes successful reprogramming and might be a major cause of low reprogramming efficiency. Although Oct4, Sox2 and Klf4 (that is, OSK) bind targeted chromatin, cMyc enhances chromatin binding by OSK. This feedback loop is thought to facilitate successful reprogramming. On the other hand, a large scale of chromatin domains spanned by H3K9me3 that inhibit OSKM binding are known to hinder efficient reprogramming.33
Moreover, the tumor suppressor p53 is proposed to prevent induced pluripotency, as p53 knockouts have been shown to significantly increase reprogramming efficiency.34, 35, 36, 37, 38 p53 has also been shown to bind the Nanog promoter and regulate its expression;39 suppressing Nanog expression leads to differentiation of ESCs. Most interestingly, knockdown of p53 downstream effectors Puma and/or p21 can significantly increase iPSC production efficiency, indicating that apopotosis and/or cell-cycle arrest function of p53 significantly inhibits efficient reprogramming.40
Reprogramming roadmaps
It has been shown that a mesenchymal-to-epithelial transition (MET) is required for reprogramming,41, 42 and Vitamin C can enhance reprogramming efficiency throughh H3K36 demethylation.43 A recent study identified an unexpected sequential epithelial-to-mesenchymal (EMT)–MET transition during the initiation of reprogramming,44 while another suggested that reprogramming is not simply a process of reversed development.45 By analysis of the expression of novel cell-surface markers CD44 and ICAM1, Malley et al.45 defined four stages of reprogramming: (1) mesenchymal, characterized by CD44+/ICAM1+ and high expression of N-Cadherin, Snail, Slug, Zeb1 and 2; (2) epidermal, characterized by CD44+/ICAM−, with high expression of Krt6a, Krt17, Ehf, Ngfr, Sfn and Evp1; (3) early pluripotent, characterized by CD44-/ICAM1− and expression of Oct4, Sall1, Sall4, Zfp296, Tcfcp2l1 and Etv5; and (4) late pluripotent, characterized by CD44−/ICAM+, with high expression of Nanog, Dppa4, Dppa5a, Sox2, Esrrb and Klf2.
More and more studies aimed to embody reprogramming roadmaps. Using genome-wide analysis, Polo et al.46 defined an intermediate cell population poised to become iPSCs and showed that two transcriptional waves were elicited by reprogramming factors; the first wave was driven by cMyc/Klf4 and the second by Oct4/Sox2/Klf4. Cells experiencing the first transcriptional wave were refractory to reprogramming and could be rescued by elevated expression of all four OSKM factors (Figure 2).46 Meanwhile, low efficiency reprogramming (normally <3% cells expressing OSKM give rise to iPSCs) complicates the dissection of its molecular mechanisms. Furthermore, a study profiling the expression of 48 pluripotent genes at the single-cell level during the reprogramming process revealed an early stochastic and a late deterministic phase.47 Intriguingly, a recent study proposed that cellular reprogramming is a deterministic process, as nearly 100% reprogramming efficiency was achieved when using Mbd3 knockout cells as the initiating somatic cells.48
Roadmaps to induced pluripotency. (a) A MET is required for reprogramming of somatic cells to iPSCs. (b) An epithelial-to-mesenchymal transition (EMT) before MET is required during reprogramming. (c) Induced pluripotency is a balance between competing differentiation forces. (d) Reprogramming elicits two transcriptional waves, which are driven by c-Myc/Klf4 (first wave) and Oct4/Sox2/Klf4 (second wave)
Seesaw model for reprogramming
The fact that cell fate is controlled by a series of master transcription factors (OSKM) overwhelms demonstration of reprogramming mechanisms.11 Most interestingly, a recent study showed that mesendodermal (GATA3, GATA6, SOX7, PAX1, GATA4, CEBPa, HNF4a, GRB2) and ectodermal specifiers (Sox1, Sox3, RCOR2, GMNN) can replace Oct4 and Sox2 to induce pluripotency, respectively.49 Most Oct4 substitutes are not enriched in ESCs and normally function in the early stages of mesendodermal differentiation. Oct4 and its substitutes can inhibit the upregulation of a group of ectodermal genes, such as the ectodermal lineage-specifier Dlx3, triggered by SKM during reprogramming. Conversely, Sox2 and its substitutes can attenuate expression of mesendodermal genes induced by OKM. The competition between mesendodermal and ectodermal specifiers promotes successful reprogramming without the two most critical reprogramming factors Oct4 and Sox2. These innovative findings have resulted in a completely new way in which to explain reprogramming, called the ‘see-saw’ model wherein cell fate is dependent on the balance between pluripotency factors and/or counteracting lineage specifiers.49, 50
Pluripotency by modification of intracellular signaling
Small-molecule compounds were originally shown to enhance reprogramming efficiency, introducing the possibility of generating iPSCs using only these compounds.51, 52 Inhibition of both mitogen-activated protein kinase and glycogen synthase kinase-3 (GSK3) signaling pathways have been shown to promote ground pluripotent reprogramming.53, 54 Furthermore, the concept of pluripotency being a balance between competing differentiation forces also suggests that it is possible to achieve pluripotency by modifing signaling networks. Most recently, by adding seven small-molecules (CHIR, 616452, Forskolin, DZNep, Valproic acid, Tranylcypromine, TTNPB) into mouse somatic cells, Hou et al.31 successfully generated completely compound-derived iPSCs. The small-molecule compounds used for reprogramming target GSK3,55, 56, 57 transforming growth factor-beta, cAMP, S-adenosylhomocysteine hydrolase, histone deacetylase, lysine-specific demethylase 1 and retinoic acid signaling (Figure 3, Table 1). This finding not only provides a new approach to induce pluripotency, avoiding tedious genetic manipulation, but also revolutionizes our understanding of molecular mechanisms of reprogramming, raising the possibility to modulating cell fate by simply modifying intrinsic cell signaling pathways alone. In support of this hypothesis, a recent report claimed a transient low-pH stress treatment could induce pluripotency in CD45+ cells.58 However, how small-molecule compounds and external stimuli like low pH activate and integrate intracellular signal transduction pathways to form completely new regulatory networks is still largely unknown.
Small-molecule signaling pathways in reprogramming. Seven small-molecule compounds synergistically induce pluripotency. CHIR99021 blocks GSK3 signaling, promoting ESC self-renewal and pluripotency. Compound 616452 inhibits TGF-β activity, which promotes MET and Nanog expression. Forskolin activates adenylate cyclase to increase cAMP expression, and DZNep inhibits S-adenosylhomocysteine hydrolase (SAH) activity and trimethylation of lysine 27 on histone H3, but how these molecules promote reprogramming is unclear. Valproic acid (VPA) looses chromatin through histone deacetylase (HDAC) inhibition and activates gene expression, while tranylcypromine inhibits lysine-specific demethylase 1 (LSD1) activity and blocks differentiation. Finally TTNPB competitively binds retinoic acid (RA) receptors and enhances proliferation
Immunogenicity of iPSC Derivatives
A major reason for the wide-spread increase in iPSC studies is their clinical application, providing potentially unlimited autologous cells for regenerative medicine. Patient-specific iPSCs can conceivably provide autologous cells that do not induce an immune response underlying rejection in human body. Unexpectedly, a previous report using a teratoma transplantation model showed that some certain iPSC-derived cells can elicit immune rejection responses.25 However, as opposed to allografts, only some certain but not all iPSC-derived cells caused immune rejection responses, due to the differential presentation of abnormalities induced by reprogramming.25, 59 For example, Abe’s group recently found that iPSC-derived cardiomyocytes, but not skin cells, can induce significant immune rejection responses.60 Although, compared with allogeneic iPSC derivatives, rejection responses induced by syngeneic iPSC-derived cells are significantly lower, endodermal cells differentiated from syngeneic iPSCs can induce immune rejection, whereas cells derived from syngeneic ESCs do not.61 Thus, these data support that only certain cell types differentiated from iPSCs are immunogenic (Figure 4).
Different fates of cells derived from iPSCs after transplantation. (a) Most cells derived from iPSCs are not immunogenic and can survive after transplantation. (b) Some cells differentiated from iPSCs with minor immunogenicity can be tolerated by the host. (c) Some abnormal iPSC progenies with immunogenicity will be rejected
As the guardian of genome, p53 has an important role in maitaining genomic stability in somatic cells and ESCs.10 If inhibition of p53 pathway is required for successful reprogramming, this raises concerns regarding the genomic stability of iPSCs and their derivatives.34, 35, 36, 37, 38 Recent studies have shown that reprogramming itself can induce both genetic and epigenetic abnormalities,62, 63, 64, 65, 66 fostering additional concerns regarding the safety of iPSCs in clinical use. Two minor antigens were identified to be abnormally expressed in the teratomas derived by syngeneic iPSCs, but not ESCs, leading to immune rejection, suggesting that reprogramming-induced epigenetic abnormalities can be passed to their progeny.67 To facilitate clinical application of iPSCs, it is important to develop new techniques that expedite clinical production of iPSCs. Furthermore, immunogenic evaluation of therapeutically valuable cells for improved patient tolerance is of utmost importance.
Conclusions and Perspectives
Long induction periods and only small initiating populations of reprogrammable cells are two major hurdles in understanding the detailed molecular mechanisms of reprogramming. New reports of lineage specifiers and small-molecule compounds that can induce pluripotency have begun to transform our comprehension of reprogramming mechanisms. However, the most critical molecular events are still unclear. In addition, considering suppression of Mbd3 has been shown to significantly increase reprogramming efficiency to almost 100%, manipulation of this gene/protein in future provides a very promising system to dissect reprogramming mechanisms. Meanwhile, with the quick development on single-cell profiling techniques and omics, our understanding of reprogramming will significantly expand. Furthermore, given the urgent clinical need, reprogramming techniques need to be optimized to minimize potential reprogramming-induced abnormalities in iPSCs.
Abbreviations
- iPSC:
-
induced pluripotent stem cell
- ESC:
-
embryonic stem cell
- MET:
-
mesenchymal-to-epithelial transition
- EMT:
-
epithelial -to-mesenchymal transition
- SCNT:
-
somatic cell nuclei transfer
- GSK3:
-
glycogen synthase kinase-3
- OSKM:
-
Oct4, Sox2, Klf4 and cMyc
References
Spemann H . Embryonic Development and Induction. Yale University Press: New Haven, CT, USA, 1938.
Briggs R, King TJ . Transplantation of living nuclei from blastula cells into enucleated frogs' eggs. Proc Natl Acad Sci USA 1952; 38: 455–463.
Briggs RK. TJ . J Embryol Exp Morphol 1957; 100: 269–312.
Gurdon JB . The developmental capacity of nuclei taken from intestinal epithelium cells of feeding tadpoles. J Embryol Exp Morphol 1962; 10: 622–640.
Gurdon JB . Adult frogs derived from the nuclei of single somatic cells. Dev Biol 1962; 4: 256–273.
Gurdon JB, Laskey RA, Reeves OR . The developmental capacity of nuclei transplanted from keratinized skin cells of adult frogs. J Embryol Exp Morphol 1975; 34: 93–112.
Miller RA, Ruddle FH . Pluripotent teratocarcinoma-thymus somatic cell hybrids. Cell 1976; 9: 45–55.
Tada M, Takahama Y, Abe K, Nakatsuji N, Tada T . Nuclear reprogramming of somatic cells by in vitro hybridization with ES cells. Curr Biol 2001; 11: 1553–1558.
Cowan CA, Atienza J, Melton DA, Eggan K . Nuclear reprogramming of somatic cells after fusion with human embryonic stem cells. Science 2005; 309: 1369–1373.
Zhao T, Xu Y . p53 and stem cells: new developments and new concerns. Trends Cell Biol 2010; 20: 170–175.
Takahashi K, Yamanaka S . Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006; 126: 663–676.
Yu J, Vodyanik MA, Smuga-Otto K, Antosiewicz-Bourget J, Frane JL, Tian S et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 2007; 318: 1917–1920.
Boland MJ, Hazen JL, Nazor KL, Rodriguez AR, Gifford W, Martin G et al. Adult mice generated from induced pluripotent stem cells. Nature 2009; 461: 91–94.
Kang L, Wang J, Zhang Y, Kou Z, Gao S . iPS cells can support full-term development of tetraploid blastocyst-complemented embryos. Cell Stem Cell 2009; 5: 135–138.
Zhao XY, Li W, Lv Z, Liu L, Tong M, Hai T et al. iPS cells produce viable mice through tetraploid complementation. Nature 2009; 461: 86–90.
Park IH, Zhao R, West JA, Yabuuchi A, Huo H, Ince TA et al. Reprogramming of human somatic cells to pluripotency with defined factors. Nature 2008; 451: 141–146.
Okita K, Ichisaka T, Yamanaka S . Generation of germline-competent induced pluripotent stem cells. Nature 2007; 448: 313–317.
Kaji K, Norrby K, Paca A, Mileikovsky M, Mohseni P, Woltjen K . Virus-free induction of pluripotency and subsequent excision of reprogramming factors. Nature 2009; 458: 771–775.
Woltjen K, Michael IP, Mohseni P, Desai R, Mileikovsky M, Hamalainen R et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 2009; 458: 766–770.
Okita K, Nakagawa M, Hyenjong H, Ichisaka T, Yamanaka S . Generation of mouse induced pluripotent stem cells without viral vectors. Science 2008; 322: 949–953.
Stadtfeld M, Nagaya M, Utikal J, Weir G, Hochedlinger K . Induced pluripotent stem cells generated without viral integration. Science 2008; 322: 945–949.
Yu J, Hu K, Smuga-Otto K, Tian S, Stewart R, Slukvin II et al. Human induced pluripotent stem cells free of vector and transgene sequences. Science 2009; 324: 797–801.
Okita K, Matsumura Y, Sato Y, Okada A, Morizane A, Okamoto S et al. A more efficient method to generate integration-free human iPS cells. Nat Methods 2011; 8: 409–412.
Wang Y, Chen J, Hu JL, Wei XX, Qin D, Gao J et al. Reprogramming of mouse and human somatic cells by high-performance engineered factors. EMBO Rep 2011; 12: 373–378.
Zhao T, Zhang ZN, Rong Z, Xu Y . Immunogenicity of induced pluripotent stem cells. Nature 2011; 474: 212–215.
Anokye-Danso F, Trivedi CM, Juhr D, Gupta M, Cui Z, Tian Y et al. Highly efficient miRNA-mediated reprogramming of mouse and human somatic cells to pluripotency. Cell Stem Cell 2011; 8: 376–388.
Miyoshi N, Ishii H, Nagano H, Haraguchi N, Dewi DL, Kano Y et al2011 Reprogramming of mouse and human cells to pluripotency using mature microRNAs. Cell Stem Cell 8: 633–638.
Kim D, Kim CH, Moon JI, Chung YG, Chang MY, Han BS et al. Generation of human induced pluripotent stem cells by direct delivery of reprogramming proteins. Cell Stem Cell 2009; 4: 472–476.
Zhou H, Wu S, Joo JY, Zhu S, Han DW, Lin T et al. Generation of induced pluripotent stem cells using recombinant proteins. Cell Stem Cell 2009; 4: 381–384.
Cho HJ, Lee CS, Kwon YW, Paek JS, Lee SH, Hur J et al. Induction of pluripotent stem cells from adult somatic cells by protein-based reprogramming without genetic manipulation. Blood 2010; 116: 386–395.
Hou P, Li Y, Zhang X, Liu C, Guan J, Li H et al. Pluripotent stem cells induced from mouse somatic cells by small-molecule compounds. Science 2013; 341: 651–654.
Jullien J, Pasque V, Halley-Stott RP, Miyamoto K, Gurdon JB . Mechanisms of nuclear reprogramming by eggs and oocytes: a deterministic process? Nat Rev Mol Cell Biol 2011; 12: 453–459.
Soufi A, Donahue G, Zaret KS . Facilitators and impediments of the pluripotency reprogramming factors' initial engagement with the genome. Cell 2012; 151: 994–1004.
Utikal J, Polo JM, Stadtfeld M, Maherali N, Kulalert W, Walsh RM et al. Immortalization eliminates a roadblock during cellular reprogramming into iPS cells. Nature 2009; 460: 1145–1148.
Li H, Collado M, Villasante A, Strati K, Ortega S, Canamero M et al. The Ink4/Arf locus is a barrier for iPS cell reprogramming. Nature 2009; 460: 1136–1139.
Kawamura T, Suzuki J, Wang YV, Menendez S, Morera LB, Raya A et al. Linking the p53 tumour suppressor pathway to somatic cell reprogramming. Nature 2009; 460: 1140–1144.
Hong H, Takahashi K, Ichisaka T, Aoi T, Kanagawa O, Nakagawa M et al. Suppression of induced pluripotent stem cell generation by the p53-p21 pathway. Nature 2009; 460: 1132–1135.
Zhao Y, Yin X, Qin H, Zhu F, Liu H, Yang W et al. Two supporting factors greatly improve the efficiency of human iPSC generation. Cell Stem Cell 2008; 3: 475–479.
Lin T, Chao C, Saito S, Mazur SJ, Murphy ME, Appella E et al. p53 induces differentiation of mouse embryonic stem cells by suppressing Nanog expression. Nat Cell Biol 2005; 7: 165–171.
Lake BB, Fink J, Klemetsaune L, Fu X, Jeffers JR, Zambetti GP et al. Context-dependent enhancement of induced pluripotent stem cell reprogramming by silencing Puma. Stem Cells 2012; 30: 888–897.
Li R, Liang J, Ni S, Zhou T, Qing X, Li H et al. A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 2010; 7: 51–63.
Samavarchi-Tehrani P, Golipour A, David L, Sung HK, Beyer TA, Datti A et al. Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell 2010; 7: 64–77.
Esteban MA, Wang T, Qin B, Yang J, Qin D, Cai J et al. Vitamin C enhances the generation of mouse and human induced pluripotent stem cells. Cell Stem Cell 2010; 6: 71–79.
Liu X, Sun H, Qi J, Wang L, He S, Liu J et al. Sequential introduction of reprogramming factors reveals a time-sensitive requirement for individual factors and a sequential EMT-MET mechanism for optimal reprogramming. Nat Cell Biol 2013; 15: 829–838.
O'Malley J, Skylaki S, Iwabuchi KA, Chantzoura E, Ruetz T, Johnsson A et al. High-resolution analysis with novel cell-surface markers identifies routes to iPS cells. Nature 2013; 499: 88–91.
Polo JM, Anderssen E, Walsh RM, Schwarz BA, Nefzger CM, Lim SM et al. A molecular roadmap of reprogramming somatic cells into iPS cells. Cell 2012; 151: 1617–1632.
Buganim Y, Faddah DA, Cheng AW, Itskovich E, Markoulaki S, Ganz K et al. Single-cell expression analyses during cellular reprogramming reveal an early stochastic and a late hierarchic phase. Cell 2012; 150: 1209–1222.
Rais Y, Zviran A, Geula S, Gafni O, Chomsky E, Viukov S et al. Deterministic direct reprogramming of somatic cells to pluripotency. Nature 2013; 502: 65–70.
Shu J, Wu C, Wu Y, Li Z, Shao S, Zhao W et al. Induction of pluripotency in mouse somatic cells with lineage specifiers. Cell 2013; 153: 963–975.
Ben-David U, Nissenbaum J, Benvenisty N . New balance in pluripotency: reprogramming with lineage specifiers. Cell 2013; 153: 939–940.
Zhu S, Ma T, Li J, Ding S . Recent advances in chemically induced reprogramming. Cell Cycle 2011; 10: 871–872.
Zhu S, Wei W, Ding S . Chemical strategies for stem cell biology and regenerative medicine. Annu Rev Biomed Eng 2011; 13: 73–90.
Silva J, Barrandon O, Nichols J, Kawaguchi J, Theunissen TW, Smith A . Promotion of reprogramming to ground state pluripotency by signal inhibition. PLoS Biol 2008; 6: e253.
Theunissen TW, van Oosten AL, Castelo-Branco G, Hall J, Smith A, Silva JC . Nanog overcomes reprogramming barriers and induces pluripotency in minimal conditions. Curr Biol 2011; 21: 65–71.
Ying QL, Wray J, Nichols J, Batlle-Morera L, Doble B, Woodgett J et al. The ground state of embryonic stem cell self-renewal. Nature 2008; 453: 519–523.
Ichida JK, Blanchard J, Lam K, Son EY, Chung JE, Egli D et al. A small-molecule inhibitor of tgf-Beta signaling replaces sox2 in reprogramming by inducing nanog. Cell Stem Cell 2009; 5: 491–503.
Tsutsui H, Valamehr B, Hindoyan A, Qiao R, Ding X, Guo S et al. An optimized small molecule inhibitor cocktail supports long-term maintenance of human embryonic stem cells. Nat Commun 2011; 2: 167.
Obokata H, Wakayama T, Sasai Y, Kojima K, Vacanti MP, Niwa H et al. Stimulus-triggered fate conversion of somatic cells into pluripotency. Nature 2014; 505: 641–647.
Cao J, Li X, Lu X, Zhang C, Yu H, Zhao T . Cells derived from iPSC can be immunogenic—yes or no? Protein Cell 2014; 5: 1–3.
Araki R, Uda M, Hoki Y, Sunayama M, Nakamura M, Ando S et al. Negligible immunogenicity of terminally differentiated cells derived from induced pluripotent or embryonic stem cells. Nature 2013; 494: 100–104.
Guha P, Morgan JW, Mostoslavsky G, Rodrigues NP, Boyd AS . Lack of immune response to differentiated cells derived from syngeneic induced pluripotent stem cells. Cell Stem Cell 2013; 12: 407–412.
Stadtfeld M, Apostolou E, Akutsu H, Fukuda A, Follett P, Natesan S et al. Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. Nature 2010; 465: 175–181.
Barrilleaux B, Knoepfler PS . Inducing iPSCs to escape the dish. Cell Stem Cell 2011; 9: 103–111.
Gore A, Li Z, Fung HL, Young JE, Agarwal S, Antosiewicz-Bourget J et al. Somatic coding mutations in human induced pluripotent stem cells. Nature 2011; 471: 63–67.
Lister R, Pelizzola M, Kida YS, Hawkins RD, Nery JR, Hon G et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 2011; 471: 68–73.
Pasi CE, Dereli-Oz A, Negrini S, Friedli M, Fragola G, Lombardo A et al. Genomic instability in induced stem cells. Cell Death Differ 2011; 18: 745–753.
Lu X, Zhao T . Clinical therapy using iPSCs: hopes and challenges. Genomics Proteomics Bioinformatics 2013; 11: 294–298.
Polychronopoulos P, Magiatis P, Skaltsounis AL, Myrianthopoulos V, Mikros E, Tarricone A et al. Structural basis for the synthesis of indirubins as potent and selective inhibitors of glycogen synthase kinase-3 and cyclin-dependent kinases. J Med Chem 2004; 47: 935–946.
Sato N, Meijer L, Skaltsounis L, Greengard P, Brivanlou AH . Maintenance of pluripotency in human and mouse embryonic stem cells through activation of Wnt signaling by a pharmacological GSK-3-specific inhibitor. Nat Med 2004; 10: 55–63.
Li W, Zhou H, Abujarour R, Zhu S, Young Joo J, Lin T et al. Generation of human-induced pluripotent stem cells in the absence of exogenous Sox2. Stem Cells 2009; 27: 2992–3000.
Hammerschmidt M, Bitgood MJ, McMahon AP . Protein kinase A is a common negative regulator of Hedgehog signaling in the vertebrate embryo. Genes Dev 1996; 10: 647–658.
Sands WA, Palmer TM . Regulating gene transcription in response to cyclic AMP elevation. Cell Signal 2008; 20: 460–466.
Fiskus W, Wang Y, Sreekumar A, Buckley KM, Shi H, Jillella A et al. Combined epigenetic therapy with the histone methyltransferase EZH2 inhibitor 3-deazaneplanocin A and the histone deacetylase inhibitor panobinostat against human AML cells. Blood 2009; 114: 2733–2743.
Bissinger EM, Heinke R, Sippl W, Jung M . Targeting epigenetic modifiers: inhibitors of histone methyltransferases. Med Chem Commun 2010; 1: 114–124.
Huangfu D, Maehr R, Guo W, Eijkelenboom A, Snitow M, Chen AE et al. Induction of pluripotent stem cells by defined factors is greatly improved by small-molecule compounds. Nat Biotechnol 2008; 26: 795–797.
Jung GA, Yoon JY, Moon BS, Yang DH, Kim HY, Lee SH et al. Valproic acid induces differentiation and inhibition of proliferation in neural progenitor cells via the beta-catenin-Ras-ERK-p21Cip/WAF1 pathway. BMC Cell Biol 2008; 9: 66.
Whyte WA, Bilodeau S, Orlando DA, Hoke HA, Frampton GM, Foster CT et al. Enhancer decommissioning by LSD1 during embryonic stem cell differentiation. Nature 2012; 482: 221–225.
Kurokawa R, DiRenzo J, Boehm M, Sugarman J, Gloss B, Rosenfeld MG et al. Regulation of retinoid signalling by receptor polarity and allosteric control of ligand binding. Nature 1994; 371: 528–531.
Acknowledgements
This work was supported by grants from the China National Basic Research Program 2012CB966901, the Strategic Priority Research Program of the Chinese Academy of Sciences XDA01040108, the National Natural Science Foundation of China Program 31271592 and the National Thousand Young Talents Program to TZ.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by M Agostini
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Liu, K., Song, Y., Yu, H. et al. Understanding the roadmaps to induced pluripotency. Cell Death Dis 5, e1232 (2014). https://doi.org/10.1038/cddis.2014.205
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2014.205
Keywords
This article is cited by
-
A rapid and stable spontaneous reprogramming system of Spermatogonial stem cells to Pluripotent State
Cell & Bioscience (2023)
-
BNIP3-dependent mitophagy safeguards ESC genomic integrity via preventing oxidative stress-induced DNA damage and protecting homologous recombination
Cell Death & Disease (2022)
-
Phosphorylation of ULK1 by AMPK is essential for mouse embryonic stem cell self-renewal and pluripotency
Cell Death & Disease (2018)
-
High autophagic flux guards ESC identity through coordinating autophagy machinery gene program by FOXO1
Cell Death & Differentiation (2017)
-
p18 inhibits reprogramming through inactivation of Cdk4/6
Scientific Reports (2016)