Featured
-
-
Article
| Open Access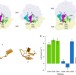 Origin of the omnipotence of eukaryotic release factor 1
Origin of the omnipotence of eukaryotic release factor 1
The eukaryotic release factor eRF1 is able to recognize the three stop codons UAA, UAG and UGA with high accuracy, while discriminating against near-cognate codons. Here the authors use molecular dynamic simulation to provide insight into the molecular basis behind the remarkable codon specificity of eRF1.
- Christoffer Lind
- , Ana Oliveira
- & Johan Åqvist
-
Article
| Open Access The Hsp70 homolog Ssb affects ribosome biogenesis via the TORC1-Sch9 signaling pathway
The Hsp70 homolog Ssb affects ribosome biogenesis via the TORC1-Sch9 signaling pathway
The yeast Hsp70 homolog Ssb is a chaperone that binds translating ribosomes where it is thought to function primarily by promoting nascent peptide folding. Here the authors find that the ribosome biogenesis defect associated with the loss of Ssb is attributable to a specific disruption in TORC1 signaling rather than defects in ribosomal protein folding.
- Kaivalya Mudholkar
- , Edith Fitzke
- & Sabine Rospert
-
Article
| Open Access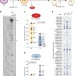 High-throughput RNA structure probing reveals critical folding events during early 60S ribosome assembly in yeast
High-throughput RNA structure probing reveals critical folding events during early 60S ribosome assembly in yeast
Ribosome biogenesis is a dynamic process that involves the ordered assembly of ribosomal proteins and numerous RNA structural rearrangements. Here the authors apply ChemModSeq, a high-throughput RNA structure probing method, to quantitatively measure changes in RNA flexibility during the nucleolar stages of 60S assembly in yeast.
- Elena Burlacu
- , Fredrik Lackmann
- & Sander Granneman
-
Article
| Open Access A general mechanism of ribosome dimerization revealed by single-particle cryo-electron microscopy
A general mechanism of ribosome dimerization revealed by single-particle cryo-electron microscopy
When bacteria enter the stationary growth phase, protein translation is suppressed via the dimerization of 70S ribosomes into inactive complexes. Here the authors provide a structural basis for how the dual domain hibernation promotion factor promotes ribosome dimerization and hibernation in bacteria.
- Linda E. Franken
- , Gert T. Oostergetel
- & Albert Guskov
-
Article
| Open Access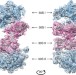 The cryo-EM structure of hibernating 100S ribosome dimer from pathogenic Staphylococcus aureus
The cryo-EM structure of hibernating 100S ribosome dimer from pathogenic Staphylococcus aureus
Under conditions of nutrient limitation, bacterial ribosomes undergo dimerization, forming a 100S complex that is translationally inactive. Here the authors present the structural basis for formation of the 100S complexes in Gram-positive bacteria, shedding light on the mechanism of translation suppression by the ribosome-silencing factors.
- Donna Matzov
- , Shintaro Aibara
- & Ada E. Yonath
-
Article
| Open Access RNA localization is a key determinant of neurite-enriched proteome
RNA localization is a key determinant of neurite-enriched proteome
Subcellular localization of RNAs and proteins is important for polarized cells such as neurons. Here the authors differentiate mouse embryonic stem cells into neurons, and analyze the local transcriptome, proteome, and translated transcriptome in their cell bodies and neurites, providing a unique resource for future studies on neuronal polarity.
- Alessandra Zappulo
- , David van den Bruck
- & Marina Chekulaeva
-
Article
| Open Access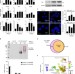 Tia1 dependent regulation of mRNA subcellular location and translation controls p53 expression in B cells
Tia1 dependent regulation of mRNA subcellular location and translation controls p53 expression in B cells
Sequestering mRNA in cytoplasmic stress granules is a mechanism for translational repression. Here the authors find that p53 mRNA, present in stress granules in activated B lymphocytes, is released upon DNA damage and is translated in a CAP-independent manner.
- Manuel D. Díaz-Muñoz
- , Vladimir Yu. Kiselev
- & Martin Turner
-
Article
| Open Access The Gcn4 transcription factor reduces protein synthesis capacity and extends yeast lifespan
The Gcn4 transcription factor reduces protein synthesis capacity and extends yeast lifespan
The transcription factor Gcn4 is known to regulate yeast amino acid synthesis. Here, the authors show that Gcn4 also acts as a repressor of protein biosynthesis in a range of conditions that enhance yeast lifespan, such as ribosomal protein knockout, calorie restriction or mTOR inhibition.
- Nitish Mittal
- , Joao C. Guimaraes
- & Mihaela Zavolan
-
Article
| Open Access A reporter system coupled with high-throughput sequencing unveils key bacterial transcription and translation determinants
A reporter system coupled with high-throughput sequencing unveils key bacterial transcription and translation determinants
Quantitative analysis of how DNA sequence determines transcription and translation regulation is of interest to systems and synthetic biologists. Here the authors present ELM-seq, which uses Dam activity as reporter for high-throughput analysis of promoter and 5’-UTR regions.
- Eva Yus
- , Jae-Seong Yang
- & Luis Serrano
-
Article
| Open Access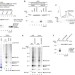 Ubiquitination of stalled ribosome triggers ribosome-associated quality control
Ubiquitination of stalled ribosome triggers ribosome-associated quality control
Several protein quality control mechanisms are in place to trigger the rapid degradation of aberrant polypeptides and mRNAs. Here the authors describe a mechanism of ribosome-mediated quality control that involves the ubiquitination of ribosomal proteins by the E3 ubiquitin ligase Hel2/RQT1.
- Yoshitaka Matsuo
- , Ken Ikeuchi
- & Toshifumi Inada
-
Article
| Open Access Protein-directed ribosomal frameshifting temporally regulates gene expression
Protein-directed ribosomal frameshifting temporally regulates gene expression
Programmed −1 ribosomal frameshifting (−1 PRF) is a mechanism whereby specific signals within mRNAs direct ribosomes to shift into an alternative reading frame. Here the authors describe a mechanism of −1 PRF that is temporally regulated by a viral protein over the course of the virus replicative cycle.
- Sawsan Napthine
- , Roger Ling
- & Andrew E. Firth
-
Article
| Open Access Misfolded polypeptides are selectively recognized and transported toward aggresomes by a CED complex
Misfolded polypeptides are selectively recognized and transported toward aggresomes by a CED complex
Misfolded polypeptide aggregates are actively transported to aggresomes, where they are degraded through aggrephagy. Here the authors show that these aggregates are selectively recognized by the CTIF–eEF1A1–DCTN1 (CED) complex and transported to aggresomes through the interactions of DCTN1 with dynein motors.
- Joori Park
- , Yeonkyoung Park
- & Yoon Ki Kim
-
Article
| Open Access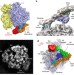 Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome
Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome
Membrane proteins are inserted co-transnationally through the association between ribosome, the signal recognition particle and its receptor, and the membrane-bound translocon. Here the authors present a cryo-EM reconstruction of this quaternary complex in the activated state and propose a model for signal sequence transfer to the translocon.
- Ahmad Jomaa
- , Yu-Hsien Hwang Fu
- & Nenad Ban
-
Article
| Open Access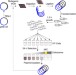 Mistranslation can enhance fitness through purging of deleterious mutations
Mistranslation can enhance fitness through purging of deleterious mutations
Mistranslation results in amino acid changes in proteins known as phenotypic mutations and these occur at a much higher rate than DNA mutations. Here, the authors show that mistranslation can increase the response to directional selection by exacerbating the fitness effects of deleterious DNA mutations.
- Sinisa Bratulic
- , Macarena Toll-Riera
- & Andreas Wagner
-
Article
| Open Access PTENβ is an alternatively translated isoform of PTEN that regulates rDNA transcription
PTENβ is an alternatively translated isoform of PTEN that regulates rDNA transcription
PTEN is a potent tumour suppressor involved in cell growth, proliferation and survival. Here the authors identify an N-terminal extended isoform of PTEN, termed PTENβ that negatively regulates ribosomal DNA transcription and cell proliferation, expanding the pleiotropic functions of the PTEN family.
- Hui Liang
- , Xi Chen
- & Yuxin Yin
-
Article
| Open Access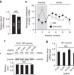 Enhanced expression of ADCY1 underlies aberrant neuronal signalling and behaviour in a syndromic autism model
Enhanced expression of ADCY1 underlies aberrant neuronal signalling and behaviour in a syndromic autism model
Fragile X syndrome (FXS) is a leading cause of autism and neurons lacking FMRP show aberrant mRNA translation and intracellular signalling. Here, the authors show that neurons from Fmr1 knockout mice have increased levels of ADCY1 protein, producing abnormal ERK1/2 signalling, dysregulated protein synthesis and behavioural symptoms associated with FXS.
- Ferzin Sethna
- , Wei Feng
- & Hongbing Wang
-
Article
| Open Access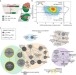 Pervasive translational regulation of the cell signalling circuitry underlies mammalian development
Pervasive translational regulation of the cell signalling circuitry underlies mammalian development
Gene expression is regulated at several levels, including through the modulation of protein translation. Here the authors find that translation control diversifies gene expression between developing tissues and regulates major signalling pathways through a complex landscape of upstream open reading frames (uORFs).
- Kotaro Fujii
- , Zhen Shi
- & Maria Barna
-
Article
| Open Access Recurring RNA structural motifs underlie the mechanics of L1 stalk movement
Recurring RNA structural motifs underlie the mechanics of L1 stalk movement
Translocation of the tRNA on the ribosome is associated with large-scale molecular movements of the ribosomal L1 stalk. Here the authors identify the key determinants that allow these dramatic movements, and suggest they represent general strategies used to enable large-scale motions in functional RNAs.
- Srividya Mohan
- & Harry F Noller
-
Article
| Open Access ATP hydrolysis by UPF1 is required for efficient translation termination at premature stop codons
ATP hydrolysis by UPF1 is required for efficient translation termination at premature stop codons
Nonsense-mediated mRNA decay (NMD) is a quality control pathway that recognizes and degrades transcripts harbouring nonsense mutations. Here the authors show that the ATPase activity of UPF1 mediates functional interactions between the NMD machinery and ribosomes required for efficient ribosome release at premature termination codons.
- Lucas D. Serdar
- , DaJuan L. Whiteside
- & Kristian E. Baker
-
Article
| Open Access Structural insights into ribosomal rescue by Dom34 and Hbs1 at near-atomic resolution
Structural insights into ribosomal rescue by Dom34 and Hbs1 at near-atomic resolution
mRNA surveillance is essential to maintain homeostasis in eukaryotes and is activated by mRNAs lacking a stop codon. Here the authors describe a high resolution cryo-EM structure of a nonstop complex that shows how arrested ribosome recognition is achieved during Dom34-mediated mRNA surveillance.
- Tarek Hilal
- , Hiroshi Yamamoto
- & Christian M.T. Spahn
-
Article
| Open Access Multivalent contacts of the Hsp70 Ssb contribute to its architecture on ribosomes and nascent chain interaction
Multivalent contacts of the Hsp70 Ssb contribute to its architecture on ribosomes and nascent chain interaction
The correct folding of proteins often requires the intervention molecular chaperones, which can occur co-translationally. Here the authors identify elements of yeast Ssb (Hsp70) that mediate ribosomal binding, and suggest a mechanism that directs efficient interaction of Ssb with the nascent chain.
- Marie A. Hanebuth
- , Roman Kityk
- & Elke Deuerling
-
Article
| Open Access Interaction of the cotranslational Hsp70 Ssb with ribosomal proteins and rRNA depends on its lid domain
Interaction of the cotranslational Hsp70 Ssb with ribosomal proteins and rRNA depends on its lid domain
In yeast, the heterodimeric ribosome-associated complex (RAC) acts in concert with the Hsp70 protein Ssb, forming a unique chaperone triad. Here the authors use structural and biochemical approaches to shed light on how translation and folding are coupled in eukaryotes.
- Andrea Gumiero
- , Charlotte Conz
- & Irmgard Sinning
-
Article
| Open Access CELF1 is a central node in post-transcriptional regulatory programmes underlying EMT
CELF1 is a central node in post-transcriptional regulatory programmes underlying EMT
Epithelial-to-mesenchymal transition is a key process in tumorigenesis but little is known about the molecular mechanism regulating such process at the translational level. Here, the authors identify a subset of mRNAs important for this process that are specifically modulated by the RNA-binding protein CELF1.
- Arindam Chaudhury
- , Shebna Cheema
- & Joel R. Neilson
-
Article
| Open Access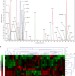 tRNA-mediated codon-biased translation in mycobacterial hypoxic persistence
tRNA-mediated codon-biased translation in mycobacterial hypoxic persistence
Mycobacteria can adapt to the stress of human infection by entering a dormant state. Here the authors show that hypoxia-induced dormancy in M. bovisBCG involves the reprogramming of tRNA wobble modifications and copy numbers, coupled with biased use of synonymous codons in survival genes.
- Yok Hian Chionh
- , Megan McBee
- & Peter C. Dedon
-
Article
| Open Access Structure of the ribosome post-recycling complex probed by chemical cross-linking and mass spectrometry
Structure of the ribosome post-recycling complex probed by chemical cross-linking and mass spectrometry
Ribosome recycling orchestrated by ABCE1 connects translation termination and mRNA surveillance mechanisms with re-initiation. Using a cross-linking and mass spectrometry approach, Kiosze-Becker et al. provide new information on the large conformational rearrangements that occur during ribosome recycling.
- Kristin Kiosze-Becker
- , Alessandro Ori
- & Robert Tampé
-
Article
| Open Access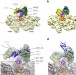 Cryo-EM study of start codon selection during archaeal translation initiation
Cryo-EM study of start codon selection during archaeal translation initiation
Initiation factor eIF2, common to eukaryotes and archaea, is a central actor in translation initiation. Here the authors describe two cryo-EM structures of archaeal 30S initiation complexes that provide a novel view of the central role that e/aIF2 plays in start codon selection.
- Pierre-Damien Coureux
- , Christine Lazennec-Schurdevin
- & Yves Mechulam
-
Article
| Open Access How EF-Tu can contribute to efficient proofreading of aa-tRNA by the ribosome
How EF-Tu can contribute to efficient proofreading of aa-tRNA by the ribosome
The translation of mRNA by the ribosome is governed by a series of large-scale conformational transitions. Here the authors use MD simulations to demonstrate how the rate of dissociation of elongation factor Tu affects the dynamics of tRNA accommodation and proofreading.
- Jeffrey K. Noel
- & Paul C. Whitford
-
Article
| Open Access Structures and stabilization of kinetoplastid-specific split rRNAs revealed by comparing leishmanial and human ribosomes
Structures and stabilization of kinetoplastid-specific split rRNAs revealed by comparing leishmanial and human ribosomes
Leishmania donovani is a protozoan parasite that can cause fatal infections in humans. Here the authors present a high resolution cryoEM structure of the L. donovani80S ribosome and compare it to its human counterpart to provide insight into the basis for drug selectivity towards this eukaryotic parasite.
- Xing Zhang
- , Mason Lai
- & Z. Hong Zhou
-
Article
| Open Access Comparative survey of the relative impact of mRNA features on local ribosome profiling read density
Comparative survey of the relative impact of mRNA features on local ribosome profiling read density
Ribosome profiling data can suffer from uneven coverage which hampers estimation of elongation rates. Connor et al.present an enhanced data smoothing method for Ribo-seq data and highlight significant variability in sequence determinants of ribosome density in publicly available data sets.
- Patrick B. F. O’Connor
- , Dmitry E. Andreev
- & Pavel V. Baranov
-
Article
| Open Access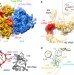 Ribosomal 18S rRNA base pairs with mRNA during eukaryotic translation initiation
Ribosomal 18S rRNA base pairs with mRNA during eukaryotic translation initiation
Prokaryotic translation initiation involves mRNA-ribosomal RNA base pairing interactions. Here, the authors provide evidence for a similar base pairing interactions occurring between the human h4 mRNA and helix 16 of the small subunit rRNA to position the correct AUG codon in the decoding site.
- Franck Martin
- , Jean-François Ménétret
- & Gilbert Eriani
-
Article
| Open Access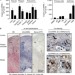 Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans
Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans
Circular RNAs are widely expressed in eukaryotic cells but their functions and mechanisms of action are still being elucidated. Here the authors show that circANRILmodulates rRNA maturation and confers protection again atherosclerosis.
- Lesca M. Holdt
- , Anika Stahringer
- & Daniel Teupser
-
Article
| Open Access UtpA and UtpB chaperone nascent pre-ribosomal RNA and U3 snoRNA to initiate eukaryotic ribosome assembly
UtpA and UtpB chaperone nascent pre-ribosomal RNA and U3 snoRNA to initiate eukaryotic ribosome assembly
Eukaryotic ribosome biogenesis involves a large number of maturations factors which are responsible for the stepwise assembly of the ribosomal subunits. Here the authors use an array of biochemical and structural biology methods to investigate the function of the UtpA and UtpB complexes as part of the small subunit processome.
- Mirjam Hunziker
- , Jonas Barandun
- & Sebastian Klinge
-
Article
| Open Access Structural basis of suppression of host translation termination by Moloney Murine Leukemia Virus
Structural basis of suppression of host translation termination by Moloney Murine Leukemia Virus
Retroviral reverse transcriptase from Moloney Murine Leukemia Virus (MoMLV) requires interaction with peptidyl release factor 1. Here, the authors report the crystal structure of this complex, and provide insights into how MoMLV uses the host translation machinery to synthesize its own proteins.
- Xuhua Tang
- , Yiping Zhu
- & Haiwei Song
-
Article
| Open Access Loss of the RNA-binding protein TACO1 causes late-onset mitochondrial dysfunction in mice
Loss of the RNA-binding protein TACO1 causes late-onset mitochondrial dysfunction in mice
Mutations in the translational activator of cytochrome c oxidase subunit I (TACO1) causes cytochrome c oxidase deficiency and Leigh Syndrome in patients. Here, the authors characterize mice with a mutation that causes lack of TACO1 expression, identifying a mouse model that could be useful for preclinical trials.
- Tara R. Richman
- , Henrik Spåhr
- & Aleksandra Filipovska
-
Article
| Open Access Pre-40S ribosome biogenesis factor Tsr1 is an inactive structural mimic of translational GTPases
Pre-40S ribosome biogenesis factor Tsr1 is an inactive structural mimic of translational GTPases
Tsr1 is an essential ribosome biogenesis factor that has known similarity to GTPases. Here, the authors report the Tsr1 crystal structure and show that it is similar to GTPases but that active site residues are not conserved; modelling of the structure into the pre-40S maps allows inferences on ribosomal maturation to be drawn.
- Urszula M. McCaughan
- , Uma Jayachandran
- & Atlanta G. Cook
-
Article
| Open Access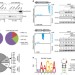 The CsrA-FliW network controls polar localization of the dual-function flagellin mRNA in Campylobacter jejuni
The CsrA-FliW network controls polar localization of the dual-function flagellin mRNA in Campylobacter jejuni
The CsrA protein binds to and represses translation of certain bacterial mRNAs. Here, Dugar et al. show for the human pathogen Campylobacter jejunithat the major flagellin mRNA acts as both a target and a regulatory 'sponge' for CsrA, and is localized at the cell poles in a translation-dependent manner.
- Gaurav Dugar
- , Sarah L. Svensson
- & Cynthia M. Sharma
-
Article
| Open Access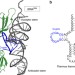 Essential structural elements in tRNAPro for EF-P-mediated alleviation of translation stalling
Essential structural elements in tRNAPro for EF-P-mediated alleviation of translation stalling
Ribosomes tend to stall during the translation of consecutive proline residues, which can be rescued by the co-translational factor EF-P. Here the authors identify a structural element of tRNAProresponsible for specific recognition by EF-P and stimulation of Pro-Pro peptide bond formation.
- Takayuki Katoh
- , Ingo Wohlgemuth
- & Hiroaki Suga
-
Article
| Open Access Conservation of uORF repressiveness and sequence features in mouse, human and zebrafish
Conservation of uORF repressiveness and sequence features in mouse, human and zebrafish
Upstream open reading frames (uORFs) can repress gene expression. Here, Guo-Liang Chew and colleagues use bioinformatics approaches to show that conservation of uORF-mediated translational repression is mediated by sequence features in human, mouse and zebrafish genomes.
- Guo-Liang Chew
- , Andrea Pauli
- & Alexander F. Schier
-
Article
| Open Access Mincle-mediated translational regulation is required for strong nitric oxide production and inflammation resolution
Mincle-mediated translational regulation is required for strong nitric oxide production and inflammation resolution
Resolution of granulomas in mycobacterial infection requires macrophages to switch from cytokine to nitric oxide (NO) production. Here the authors show that Mincle stimulates translation of the key NO synthesis genes by a mechanism dependent on p38-mediated hypusination of eiF5A.
- Wook-Bin Lee
- , Ji-Seon Kang
- & Young-Joon Kim
-
Article
| Open Access Genome-wide assessment of differential translations with ribosome profiling data
Genome-wide assessment of differential translations with ribosome profiling data
The global measurement of ribosome occupancy on mRNAs is commonly used as a proxy in estimating rates of protein synthesis. Here the authors describe Xtail, a computational approach that facilitates the extraction of accurate quantitative insight from ribosome profiling data (Ribo-Seq).
- Zhengtao Xiao
- , Qin Zou
- & Xuerui Yang
-
Article
| Open Access Steric interactions lead to collective tilting motion in the ribosome during mRNA–tRNA translocation
Steric interactions lead to collective tilting motion in the ribosome during mRNA–tRNA translocation
During protein elongation, the translocation of mRNA and tRNA molecules across the 30S ribosomal subunit is associated with large-scale motions of the 30S head domain. Here the authors carry out MD simulations to probe the associated steric interactions and identify novel tilting motions during the late stages of translocation.
- Kien Nguyen
- & Paul C. Whitford
-
Article
| Open Access Structures of the E. coli translating ribosome with SRP and its receptor and with the translocon
Structures of the E. coli translating ribosome with SRP and its receptor and with the translocon
The co-translational insertion of proteins into membranes requires interaction between a ribosome-bound signal recognition particle (SRP) and a membrane-bound translocon. Here the authors use cryo-EM and single particle reconstructions to obtain a comprehensive view of the co-translational protein targeting process.
- Ahmad Jomaa
- , Daniel Boehringer
- & Nenad Ban
-
Article
| Open Access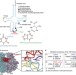 Novel base-pairing interactions at the tRNA wobble position crucial for accurate reading of the genetic code
Novel base-pairing interactions at the tRNA wobble position crucial for accurate reading of the genetic code
The anticodon loops of almost all tRNAs contain modifications known to be important for their function. Here the authors use crystallography to provide new mechanistic insights into how the modification at the wobble position of the E. coli tRNALysUUUassists in discrimination between cognate and near-cognate codons.
- Alexey Rozov
- , Natalia Demeshkina
- & Gulnara Yusupova
-
Article
| Open Access The Shine-Dalgarno sequence of riboswitch-regulated single mRNAs shows ligand-dependent accessibility bursts
The Shine-Dalgarno sequence of riboswitch-regulated single mRNAs shows ligand-dependent accessibility bursts
In response to intracellular signals, bacterial translational riboswitches embedded in mRNAs can regulate gene expression through inhibition of translation initiation. Here, the authors describe SiM-KARTS, a novel approach for detecting changes in the structure of single RNA molecules in response to a ligand.
- Arlie J. Rinaldi
- , Paul E. Lund
- & Nils G. Walter
-
Article
| Open Access A DNA-based system for selecting and displaying the combined result of two input variables
A DNA-based system for selecting and displaying the combined result of two input variables
DNA based sensors provide substantial amounts of information that requires integration and processing. Here the authors demonstrate a DNA-based calculator that takes two inputs, identifies the solution in a library of answers and displays the result.
- Huajie Liu
- , Jianbang Wang
- & Kurt V. Gothelf
-
Article
| Open Access Mammalian SRP receptor switches the Sec61 translocase from Sec62 to SRP-dependent translocation
Mammalian SRP receptor switches the Sec61 translocase from Sec62 to SRP-dependent translocation
Sec62 is a membrane-bound protein that is involved in the translocation of proteins via the signal recognition particle-independent pathway. Here, the authors show that the receptor SRα displaces Sec62 from the translocon and isolate the domain on SRα that is responsible for this.
- Bhalchandra Jadhav
- , Michael McKenna
- & Martin R. Pool
-
Article
| Open Access eIF6 coordinates insulin sensitivity and lipid metabolism by coupling translation to transcription
eIF6 coordinates insulin sensitivity and lipid metabolism by coupling translation to transcription
Insulin enhances mRNA translation via the translation initiation factor eIF6. Here, Brina et al. show that insulin-mediated activation of eIF6 is associated with the selective translation of genes involved in glycolysis and lipid synthesis with characteristic G/C-rich and uORF sequences in their mRNA.
- Daniela Brina
- , Annarita Miluzio
- & Stefano Biffo
-
Article |
 Growth-regulating Mycobacterium tuberculosis VapC-mt4 toxin is an isoacceptor-specific tRNase
Growth-regulating Mycobacterium tuberculosis VapC-mt4 toxin is an isoacceptor-specific tRNase
Toxin–antitoxin systems of the Vap class regulate the growth of several bacterial pathogens including Mycobacterium tuberculosis. Here, the authors show that toxin VapC-mt4 arrests M. tuberculosis growth by specifically cleaving three tRNAs at a single site in their anticodon stem loop, leading to translation inhibition.
- Jonathan W. Cruz
- , Jared D. Sharp
- & Nancy A. Woychik
-
Article
| Open Access Cryo-EM structure of Hepatitis C virus IRES bound to the human ribosome at 3.9-Å resolution
Cryo-EM structure of Hepatitis C virus IRES bound to the human ribosome at 3.9-Å resolution
The Hepatitis C virus (HCV) relies on an internal ribosome entry site (IRES) for translation of all the proteins encoded by its single-stranded RNA genome. Here the authors present a near-atomic cryo-EM structure of the HCV IRES bound to the human ribosome, shedding light on the initiation mechanism of HCV's and related IRESs.
- Nick Quade
- , Daniel Boehringer
- & Nenad Ban

 A conformational switch in initiation factor 2 controls the fidelity of translation initiation in bacteria
A conformational switch in initiation factor 2 controls the fidelity of translation initiation in bacteria