Featured
-
-
Article
| Open Access Regulation by the RNA-binding protein Unkempt at its effector interface
Regulation by the RNA-binding protein Unkempt at its effector interface
How RNA-binding proteins (RBPs) regulate gene expression via effectors of RNA processing is unclear. Here, the authors dissect the effector interface of an essential RBP, Unkempt, and investigate its contribution to translational control in cells.
- Kriti Shah
- , Shiyang He
- & Jernej Murn
-
Article
| Open Access Translation velocity determines the efficacy of engineered suppressor tRNAs on pathogenic nonsense mutations
Translation velocity determines the efficacy of engineered suppressor tRNAs on pathogenic nonsense mutations
An emerging therapeutic strategy is to suppress nonsense mutations with engineered suppressor tRNAs. Here, the authors show that the mRNA translation velocity is a key parameter determining the efficacy of suppressor tRNAs.
- Nikhil Bharti
- , Leonardo Santos
- & Zoya Ignatova
-
Article
| Open Access Patchy and widespread distribution of bacterial translation arrest peptides associated with the protein localization machinery
Patchy and widespread distribution of bacterial translation arrest peptides associated with the protein localization machinery
Regulatory arrest peptides interact with the bacterial ribosome to halt their own translation. Here, Fujiwara et al. analyse thousands of bacterial genome sequences and identify additional arrest peptides, revealing sequence diversity and patchy, but widespread, distribution across the bacterial domain.
- Keigo Fujiwara
- , Naoko Tsuji
- & Shinobu Chiba
-
Article
| Open Access Diverging co-translational protein complex assembly pathways are governed by interface energy distribution
Diverging co-translational protein complex assembly pathways are governed by interface energy distribution
Protein complex assembly can occur co-translationally. Here, the authors uncover diverging assembly pathways and hotspot disruptions in N-terminal acetyltransferases, enzymes implicated in neurodegenerative diseases. Their model predicts co-translational assembly based on interface energy distribution.
- Johannes Venezian
- , Hagit Bar-Yosef
- & Ayala Shiber
-
Article
| Open Access Extended stop codon context predicts nonsense codon readthrough efficiency in human cells
Extended stop codon context predicts nonsense codon readthrough efficiency in human cells
Stop codon readthrough, the ribosomal bypass of mRNA nonsense codons, has therapeutic potential for diseases caused by nonsense mutations. Here, the authors used machine learning to define readthrough-conducive mRNA sequences and predict specific CFTR alleles likely amenable to readthrough therapy.
- Kotchaphorn Mangkalaphiban
- , Lianwu Fu
- & Allan Jacobson
-
Article
| Open Access The SecM arrest peptide traps a pre-peptide bond formation state of the ribosome
The SecM arrest peptide traps a pre-peptide bond formation state of the ribosome
Stalling of ribosomes by the nascent polypeptide chain is widely used to regulate gene expression. Here, Gersteuer et al determine cryo-EM structures of SecM-stalled ribosomes revealing the mechanism by which the SecM peptide arrests translation.
- Felix Gersteuer
- , Martino Morici
- & Daniel N. Wilson
-
Article
| Open Access RAPP-containing arrest peptides induce translational stalling by short circuiting the ribosomal peptidyltransferase activity
RAPP-containing arrest peptides induce translational stalling by short circuiting the ribosomal peptidyltransferase activity
Translation of RAPP (Arg-AlaPro-Pro) motifs induces ribosome stalling. Here, structures of RAPP-stalled ribosomes reveal that RAPP motifs short circuit the ribosomal peptidyltransferase activity to induce stalling.
- Martino Morici
- , Sara Gabrielli
- & Daniel N. Wilson
-
Article
| Open Access Translation efficiency driven by CNOT3 subunit of the CCR4-NOT complex promotes leukemogenesis
Translation efficiency driven by CNOT3 subunit of the CCR4-NOT complex promotes leukemogenesis
Here the authors uncovered CNOT3, a subunit of the CCR4-NOT complex, as an essential modulator of translation in leukemia. The work pointed to the potential of targeting the posttranscriptional circuitry via CNOT3 as a therapeutic vulnerability in acute myeloid leukemia.
- Maryam Ghashghaei
- , Yilin Liu
- & Ly P. Vu
-
Article
| Open Access dCas13-mediated translational repression for accurate gene silencing in mammalian cells
dCas13-mediated translational repression for accurate gene silencing in mammalian cells
Current gene silencing tools can have drawbacks. Here the authors report CRISPRδ, an approach for translational silencing, harnessing catalytically inactive Cas13 proteins (dCas13): they also show that fusion of a translational repressor to dCas13 further improved the performance.
- Antonios Apostolopoulos
- , Naohiro Kawamoto
- & Shintaro Iwasaki
-
Article
| Open Access N-Acetyltransferase 10 represses Uqcr11 and Uqcrb independently of ac4C modification to promote heart regeneration
N-Acetyltransferase 10 represses Uqcr11 and Uqcrb independently of ac4C modification to promote heart regeneration
Here, Ma et al. investigate the translational profile of cardiac regeneration, pointing to Nat10 as a key regulator of cardiomyocyte proliferative potential, and describing how it regulates cardiac gene expression.
- Wenya Ma
- , Yanan Tian
- & Benzhi Cai
-
Article
| Open Access Widespread stable noncanonical peptides identified by integrated analyses of ribosome profiling and ORF features
Widespread stable noncanonical peptides identified by integrated analyses of ribosome profiling and ORF features
By developing computational algorithms, the authors annotated translated open reading frames in five eukaryotes and found many stable peptides are encoded by putative ‘noncoding’ regions of genomes.
- Haiwang Yang
- , Qianru Li
- & Zhe Ji
-
Article
| Open Access Transient disome complex formation in native polysomes during ongoing protein synthesis captured by cryo-EM
Transient disome complex formation in native polysomes during ongoing protein synthesis captured by cryo-EM
Direct visualization of short-lived intermediates during active protein synthesis remains challenging. Here, the authors structurally capture transient translation intermediates to uncover temporary disome formation during ribosome collisions.
- Timo Flügel
- , Magdalena Schacherl
- & Christian M. T. Spahn
-
Article
| Open Access Stalled translation by mitochondrial stress upregulates a CNOT4-ZNF598 ribosomal quality control pathway important for tissue homeostasis
Stalled translation by mitochondrial stress upregulates a CNOT4-ZNF598 ribosomal quality control pathway important for tissue homeostasis
Ribosome associated quality control (RQC) is a new area of biological investigation with emerging connection to a broad range of diseases. Here authors show that mitochondrial stress can upregulate a new RQC pathway important for tissue homeostasis.
- Ji Geng
- , Shuangxi Li
- & Bingwei Lu
-
Article
| Open Access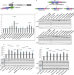 Decryption of sequence, structure, and functional features of SINE repeat elements in SINEUP non-coding RNA-mediated post-transcriptional gene regulation
Decryption of sequence, structure, and functional features of SINE repeat elements in SINEUP non-coding RNA-mediated post-transcriptional gene regulation
Here the authors elucidate structure-function relationships of SINEUPs, antisense long non-coding RNAs that through SINE repeat elements positively regulate protein translation of mRNAs they pair with. SINEUP’s functional domains do not share common ancestors.
- Harshita Sharma
- , Matthew N. Z. Valentine
- & Piero Carninci
-
Article
| Open Access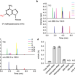 C2-methyladenosine in tRNA promotes protein translation by facilitating the decoding of tandem m2A-tRNA-dependent codons
C2-methyladenosine in tRNA promotes protein translation by facilitating the decoding of tandem m2A-tRNA-dependent codons
Duan et al. demonstrate that the m2A modification is ubiquitous in plants and tRNA m2A37 promotes a relaxed conformation of tRNA, enhancing translation efficiency by facilitating decoding of tandem m2A-tRNA-dependent codons.
- Hong-Chao Duan
- , Chi Zhang
- & Guifang Jia
-
Article
| Open Access Cryo- EM structure of the mycobacterial 70S ribosome in complex with ribosome hibernation promotion factor RafH
Cryo- EM structure of the mycobacterial 70S ribosome in complex with ribosome hibernation promotion factor RafH
Ribosome hibernation is a key survival strategy bacteria adapt under stress. Here, cryo- EM structure of mycobacterial 70S ribosome with hypoxia stress-induced factor RafH suggests the molecular mechanism of RafH-induced ribosome hibernation.
- Niraj Kumar
- , Shivani Sharma
- & Prem S. Kaushal
-
Article
| Open Access Nuclear Hsp104 safeguards the dormant translation machinery during quiescence
Nuclear Hsp104 safeguards the dormant translation machinery during quiescence
During aging, proteins are damaged and can misfold, compromising cellular viability. Here, Kohler et al. uncover how aging cells maintain fitness by redirecting the protein repair factor Hsp104 to the nucleus in response to metabolic cues.
- Verena Kohler
- , Andreas Kohler
- & Sabrina Büttner
-
Article
| Open Access Remodeling of the ribosomal quality control and integrated stress response by viral ubiquitin deconjugases
Remodeling of the ribosomal quality control and integrated stress response by viral ubiquitin deconjugases
Here, the authors show how the vDUB from the large tegument protein from the human herpes virus can reprogram translation in host cells by modulating the activity of the ribosome quality machinery and activating the integrated stress response.
- Jiangnan Liu
- , Noemi Nagy
- & Maria G. Masucci
-
Article
| Open Access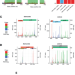 The distinct translational landscapes of gram-negative Salmonella and gram-positive Listeria
The distinct translational landscapes of gram-negative Salmonella and gram-positive Listeria
In this work, Bryant and Lastovka et al. utilise advanced ribosome profiling and transcriptomics techniques, to reveal distinct translation control mechanisms in Salmonella and Listeria, two highly divergent bacterial species.
- Owain J. Bryant
- , Filip Lastovka
- & Betty Y. -W. Chung
-
Article
| Open Access Characterization of nucleolar SUMO isopeptidases unveils a general p53-independent checkpoint of impaired ribosome biogenesis
Characterization of nucleolar SUMO isopeptidases unveils a general p53-independent checkpoint of impaired ribosome biogenesis
Ribosome biogenesis is tightly coordinated with cell-cycle progression. By characterizing the SUMO isopeptidases SENP3/SENP5, Doenig et al. identify a long-sought p53-independent impaired ribosome checkpoint that converges on downregulation of CDK6.
- Judith Dönig
- , Hannah Mende
- & Stefan Müller
-
Article
| Open Access Structural insights into the role of GTPBP10 in the RNA maturation of the mitoribosome
Structural insights into the role of GTPBP10 in the RNA maturation of the mitoribosome
The biogenesis of ribosomes is a highly coordinated process. Here, Nguyen et al. uncover how the mitochondria-specific interplay of the GTPases GTPBP10 and GTPBP7 ensures proper maturation of the catalytic RNA center of the human mitoribosome.
- Thu Giang Nguyen
- , Christina Ritter
- & Eva Kummer
-
Article
| Open Access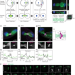 An oocyte meiotic midbody cap is required for developmental competence in mice
An oocyte meiotic midbody cap is required for developmental competence in mice
Midbodies form during cell division and play roles in cell function and fate. Here, the authors show that the meiotic midbody in mouse oocytes has a specialized cap structure required to retain nascent proteins in eggs and for full developmental competence.
- Gyu Ik Jung
- , Daniela Londoño-Vásquez
- & Karen Schindler
-
Article
| Open Access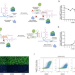 Rational design of microRNA-responsive switch for programmable translational control in mammalian cells
Rational design of microRNA-responsive switch for programmable translational control in mammalian cells
Artificial regulation of translation by intracellular RNAs has many potential applications. Here, authors design a platform capable of miRNA-triggered upregulation or downregulation using a single RNA construct, and demonstrate its use in constructing logic gates and cell-type classifiers.
- Hui Ning
- , Gan Liu
- & Zhen Xie
-
Article
| Open Access The structure of a hibernating ribosome in a Lyme disease pathogen
The structure of a hibernating ribosome in a Lyme disease pathogen
Ribosomes are prime targets for antibiotics in pathogenic bacteria. Here, cryo-electron microscopy reveals features in the Borrelia burgdorferi ribosome that provide insights into ribosome evolution, dormancy, and antibiotic binding.
- Manjuli R. Sharma
- , Swati R. Manjari
- & Nilesh K. Banavali
-
Article
| Open Access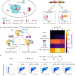 Remodeling the cellular stress response for enhanced genetic code expansion in mammalian cells
Remodeling the cellular stress response for enhanced genetic code expansion in mammalian cells
Genetic code expansion (GCE) is a protein engineering tool that enables programmed and site-specific installation of noncanonical amino acids into proteins. Here, authors show that cellular stress remodelling boosts GCE in mammalian cells including GCE realized by orthogonally translating organelles.
- Mikhail E. Sushkin
- , Christine Koehler
- & Edward A. Lemke
-
Article
| Open Access RNA-based translation activators for targeted gene upregulation
RNA-based translation activators for targeted gene upregulation
Many diseases are driven by the insufficient expression of critical genes, but few technologies are capable of rescuing these endogenous protein levels. Here, Cao et al. present an RNA-based technology that boosts protein production from endogenous mRNAs by upregulating their translation.
- Yang Cao
- , Huachun Liu
- & Bryan C. Dickinson
-
Article
| Open Access Secondary structures that regulate mRNA translation provide insights for ASO-mediated modulation of cardiac hypertrophy
Secondary structures that regulate mRNA translation provide insights for ASO-mediated modulation of cardiac hypertrophy
The GAT4A transcription factor mediates cardiac development. Here the authors identify that the 5′ UTR of GATA4 mRNA contains a double stranded structure downstream of an upstream open reading frame (uORF) that promotes uORF-mediated suppression of the main ORF.
- Omar M. Hedaya
- , Kadiam C. Venkata Subbaiah
- & Peng Yao
-
Article
| Open Access Single-molecule imaging reveals distinct elongation and frameshifting dynamics between frames of expanded RNA repeats in C9ORF72-ALS/FTD
Single-molecule imaging reveals distinct elongation and frameshifting dynamics between frames of expanded RNA repeats in C9ORF72-ALS/FTD
Hexanucleotide repeat expansion in the C9ORF72 gene produces toxic dipeptide repeat (DPR) in amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD). Here the authors apply single-molecule methods to study the translation dynamics of C9ORF72 expanded repeat in different frames showing that multiple translation steps contribute to the final toxic dipeptide production.
- Malgorzata J. Latallo
- , Shaopeng Wang
- & Bin Wu
-
Article
| Open Access Antibiotic hyper-resistance in a class I aminoacyl-tRNA synthetase with altered active site signature motif
Antibiotic hyper-resistance in a class I aminoacyl-tRNA synthetase with altered active site signature motif
Aminoacyl-tRNA synthetases translate the genetic code. These enzymes harbor signature catalytic motifs dating from their ancient ancestors. A natural variation of one of the stated motifs was discovered and linked to antibiotic hyper-resistance.
- A. Brkic
- , M. Leibundgut
- & I. Gruic-Sovulj
-
Article
| Open Access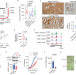 A super-enhancer-regulated RNA-binding protein cascade drives pancreatic cancer
A super-enhancer-regulated RNA-binding protein cascade drives pancreatic cancer
The epigenetic mechanisms underlying pancreatic ductal adenocarcinoma (PDAC) are not fully elucidated. Here, the authors reveal a druggable super-enhancer-mediated RNA-binding protein cascade that supports PDAC growth through enhanced mRNA translation.
- Corina E. Antal
- , Tae Gyu Oh
- & Ronald M. Evans
-
Article
| Open Access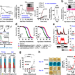 Early-adulthood spike in protein translation drives aging via juvenile hormone/germline signaling
Early-adulthood spike in protein translation drives aging via juvenile hormone/germline signaling
The transient elevation in protein translation during early-adulthood in Drosophila imposes long-lasting negative impacts on future aging trajectories by triggering proteostatic dysfunction at old ages.
- Harper S. Kim
- , Danitra J. Parker
- & Andrew M. Pickering
-
Article
| Open Access Extensive breaking of genetic code degeneracy with non-canonical amino acids
Extensive breaking of genetic code degeneracy with non-canonical amino acids
Genetic code expansion is limited by the degeneracy of the 61 sense codons which encode for only 20 amino acids. Here, the authors show that by combining hyperaccurate ribosomes and in vitro transcribed tRNAs, dramatic and extensive breaking of sense codon degeneracy can be achieved.
- Clinton A. L. McFeely
- , Bipasana Shakya
- & Matthew C. T. Hartman
-
Article
| Open Access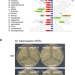 MenT nucleotidyltransferase toxins extend tRNA acceptor stems and can be inhibited by asymmetrical antitoxin binding
MenT nucleotidyltransferase toxins extend tRNA acceptor stems and can be inhibited by asymmetrical antitoxin binding
Bacteria can control their growth using internal toxins regulated by antitoxins. Here, the authors show MenT toxins from Mycobacterium tuberculosis work by modifying tRNAs, and asymmetric antitoxin binding blocks toxin activity.
- Xibing Xu
- , Ben Usher
- & Pierre Genevaux
-
Article
| Open Access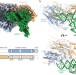 Structural basis for a degenerate tRNA identity code and the evolution of bimodal specificity in human mitochondrial tRNA recognition
Structural basis for a degenerate tRNA identity code and the evolution of bimodal specificity in human mitochondrial tRNA recognition
Aminoacyl-tRNA synthetases catalyze the ligation of amino acids to their cognate tRNAs. Here the authors report the cryo-EM structure of a human mitochondrial seryl-tRNA synthetase•mtRNASer complex showing how strong mutation pressure on mtRNA genes drove a rewiring of intermolecular recognition rules.
- Bernhard Kuhle
- , Marscha Hirschi
- & Paul Schimmel
-
Article
| Open Access Modulation of translational decoding by m6A modification of mRNA
Modulation of translational decoding by m6A modification of mRNA
m6A is an mRNA modification that slows down translation elongation. Here, Jain et al. show that m6A delays decoding and increases tRNA drop-off from the ribosome by favoring alternative codon conformations that are rejected by the ribosome.
- Sakshi Jain
- , Lukasz Koziej
- & Marina V. Rodnina
-
Article
| Open Access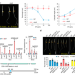 A translational regulator MHZ9 modulates ethylene signaling in rice
A translational regulator MHZ9 modulates ethylene signaling in rice
The authors identify a GYF domain-containing protein MHZ9 in rice, which regulates ethylene signaling by directly binding to OsEBF1/2 and other mRNAs thus regulating their translation efficiency in P-body via interacting with OsEIN2.
- Yi-Hua Huang
- , Jia-Qi Han
- & Jin-Song Zhang
-
Article
| Open Access Molecular basis of the pleiotropic effects by the antibiotic amikacin on the ribosome
Molecular basis of the pleiotropic effects by the antibiotic amikacin on the ribosome
Here the authors use fast kinetics, X-ray crystallography, and cryo-EM to uncover the mechanism of ribosome inhibition by amikacin and kanamycin. They find that amikacin binds near the P-site tRNA, offering new strategies to fight antibiotic resistance.
- Savannah M. Seely
- , Narayan P. Parajuli
- & Matthieu G. Gagnon
-
Article
| Open Access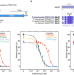 Regulation of the macrolide resistance ABC-F translation factor MsrD
Regulation of the macrolide resistance ABC-F translation factor MsrD
Antibiotic resistance ABC-F factors protect the ribosome from important antibiotics. Here, for one of them, the authors describe its molecular regulation that involves ribosome stalling by antibiotics for which the factor provides resistance.
- Corentin R. Fostier
- , Farès Ousalem
- & Grégory Boël
-
Article
| Open Access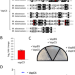 Mycobacterium abscessus VapC5 toxin potentiates evasion of antibiotic killing by ribosome overproduction and activation of multiple resistance pathways
Mycobacterium abscessus VapC5 toxin potentiates evasion of antibiotic killing by ribosome overproduction and activation of multiple resistance pathways
Mycobacterium abscessus (Mab) infections are difficult to clear with antibiotics. Here the authors show that clinical Mab strains can acquire a toxin-antitoxin system that enhances survival upon treatment with current first-line antibiotics through depletion of tRNASerCGA and subsequent ribosome overproduction.
- Eduardo A. Troian
- , Heather M. Maldonado
- & Nancy A. Woychik
-
Article
| Open Access A viral pan-end RNA element and host complex define a SARS-CoV-2 regulon
A viral pan-end RNA element and host complex define a SARS-CoV-2 regulon
Here, Khan et al. identify a pan-genome-end sarbecoviral RNA element in SARS-CoV-2 that recruits an unconventional host multiprotein-complex to enhance viral translation and assigns a new function to ORF10. Antisense targeting of the element points to a potential novel therapeutic modality.
- Debjit Khan
- , Fulvia Terenzi
- & Paul L. Fox
-
Article
| Open Access THRONCAT: metabolic labeling of newly synthesized proteins using a bioorthogonal threonine analog
THRONCAT: metabolic labeling of newly synthesized proteins using a bioorthogonal threonine analog
Incorporating bioorthogonally-modified amino acids into newly synthesized proteins allows studying the nascent proteome, but current methods often require special conditions or are toxic to cells. Here, the authors develop a method that uses β-ethynylserine to label nascent proteins under standard conditions without harming cells.
- Bob J. Ignacio
- , Jelmer Dijkstra
- & Kimberly M. Bonger
-
Article
| Open Access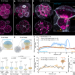 Apicobasal RNA asymmetries regulate cell fate in the early mouse embryo
Apicobasal RNA asymmetries regulate cell fate in the early mouse embryo
How do cells of the preimplantation mouse embryo make decisions? Here the authors discovered that the spatial sorting of mRNAs, tRNA, rRNAs and organelles lead to localized translation, conducive for cell fate allocation and embryonic development.
- Azelle Hawdon
- , Niall D. Geoghegan
- & Jennifer Zenker
-
Article
| Open Access Quality control of protein synthesis in the early elongation stage
Quality control of protein synthesis in the early elongation stage
Peptidyl-tRNAs (pep-tRNAs) frequently dissociate from ribosome, called as pep-tRNA drop-off. But, its function remained unclear. The authors proposed a new quality control mechanism of protein synthesis by active rejection of miscoded pep-tRNAs in the early stage of translation.
- Asuteka Nagao
- , Yui Nakanishi
- & Tsutomu Suzuki
-
Article
| Open Access Translational reprogramming as a driver of antimony-drug resistance in Leishmania
Translational reprogramming as a driver of antimony-drug resistance in Leishmania
Leishmania is a unicellular protozoan that has limited transcriptional control. Here, the authors show that translational control is a major mechanism of antimony drug resistance in Leishmania. They observe a dramatic translatome reprogramming during development of resistance to the drug and report translational control as a major driver of antimony-resistant phenotypes.
- Sneider Alexander Gutierrez Guarnizo
- , Elena B. Tikhonova
- & Zemfira N. Karamysheva
-
Article
| Open Access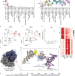 RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function
RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function
RPL3L is a paralog of the ribosomal protein RPL3 and specifically expressed in heart and skeletal muscle. Here, the authors show that RPL3L-containing ribosomes regulate translation elongation dynamics especially for the transcripts related to cardiac muscle contraction.
- Chisa Shiraishi
- , Akinobu Matsumoto
- & Keiichi I. Nakayama
-
Article
| Open Access Copy number variation in tRNA isodecoder genes impairs mammalian development and balanced translation
Copy number variation in tRNA isodecoder genes impairs mammalian development and balanced translation
Enigmatically tRNA genes exist in several hundred copies in mammalian genomes. Here the authors find a precipitous failure of development and increased amino acid misincorporation upon systematic elimination of tRNA-Phe genes in mice using CRISPR.
- Laetitia A. Hughes
- , Danielle L. Rudler
- & Aleksandra Filipovska
-
Article
| Open Access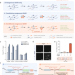 Autocatalytic base editing for RNA-responsive translational control
Autocatalytic base editing for RNA-responsive translational control
Genetic circuits that control transgene expression in response to pre-defined transcriptional cues would enable the development of smart therapeutics. Here the authors engineer programmable RNA sensors, DART VADARs, in which ADARs autocatalytically convert target hybridization into a translational output, thus amplifying editing by endogenous ADAR via positive feedback and conferring high dynamic range and a small genetic footprint.
- Raphaël V. Gayet
- , Katherine Ilia
- & James J. Collins
-
Article
| Open Access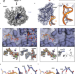 rRNA methylation by Spb1 regulates the GTPase activity of Nog2 during 60S ribosomal subunit assembly
rRNA methylation by Spb1 regulates the GTPase activity of Nog2 during 60S ribosomal subunit assembly
Regulation of 60S biogenesis remains poorly understood. Using cryo-EM, the authors show that failure of Spb1 to methylate the A-loop nucleotide G2922 prematurely activates the GTPase Nog2, suggesting that Spb1 and Nog2 form a kinetic checkpoint during ribosome maturation.
- Kamil Sekulski
- , Victor Emmanuel Cruz
- & Jan P. Erzberger
-
Article
| Open Access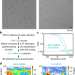 The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services
The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services
Developments in cryo-EM sample preparation and data collection are pivotal for structure determination. Fromm et al. present a 1.55 Å structure of the translating bacterial ribosome that provides new insights on its function and may allow for more precise structure-based drug design.
- Simon A. Fromm
- , Kate M. O’Connor
- & Simone Mattei

 The DEAD-box ATPase Dbp10/DDX54 initiates peptidyl transferase center formation during 60S ribosome biogenesis
The DEAD-box ATPase Dbp10/DDX54 initiates peptidyl transferase center formation during 60S ribosome biogenesis