Featured
-
-
Article
| Open Access The DEAD-box ATPase Dbp10/DDX54 initiates peptidyl transferase center formation during 60S ribosome biogenesis
The DEAD-box ATPase Dbp10/DDX54 initiates peptidyl transferase center formation during 60S ribosome biogenesis
Cruz et al. describe the role of Dbp10/DDX54 in remodeling rRNA structure within the immature eukaryotic peptidyl transferase center of the ribosome, coupling energy-dependent catalysis to a post-catalytic role in factor exchange during 60S ribosomal subunit assembly.
- Victor E. Cruz
- , Christine S. Weirich
- & Jan P. Erzberger
-
Article
| Open Access Patchy and widespread distribution of bacterial translation arrest peptides associated with the protein localization machinery
Patchy and widespread distribution of bacterial translation arrest peptides associated with the protein localization machinery
Regulatory arrest peptides interact with the bacterial ribosome to halt their own translation. Here, Fujiwara et al. analyse thousands of bacterial genome sequences and identify additional arrest peptides, revealing sequence diversity and patchy, but widespread, distribution across the bacterial domain.
- Keigo Fujiwara
- , Naoko Tsuji
- & Shinobu Chiba
-
Article
| Open Access Diverging co-translational protein complex assembly pathways are governed by interface energy distribution
Diverging co-translational protein complex assembly pathways are governed by interface energy distribution
Protein complex assembly can occur co-translationally. Here, the authors uncover diverging assembly pathways and hotspot disruptions in N-terminal acetyltransferases, enzymes implicated in neurodegenerative diseases. Their model predicts co-translational assembly based on interface energy distribution.
- Johannes Venezian
- , Hagit Bar-Yosef
- & Ayala Shiber
-
Article
| Open Access Extended stop codon context predicts nonsense codon readthrough efficiency in human cells
Extended stop codon context predicts nonsense codon readthrough efficiency in human cells
Stop codon readthrough, the ribosomal bypass of mRNA nonsense codons, has therapeutic potential for diseases caused by nonsense mutations. Here, the authors used machine learning to define readthrough-conducive mRNA sequences and predict specific CFTR alleles likely amenable to readthrough therapy.
- Kotchaphorn Mangkalaphiban
- , Lianwu Fu
- & Allan Jacobson
-
Article
| Open Access The SecM arrest peptide traps a pre-peptide bond formation state of the ribosome
The SecM arrest peptide traps a pre-peptide bond formation state of the ribosome
Stalling of ribosomes by the nascent polypeptide chain is widely used to regulate gene expression. Here, Gersteuer et al determine cryo-EM structures of SecM-stalled ribosomes revealing the mechanism by which the SecM peptide arrests translation.
- Felix Gersteuer
- , Martino Morici
- & Daniel N. Wilson
-
Article
| Open Access RAPP-containing arrest peptides induce translational stalling by short circuiting the ribosomal peptidyltransferase activity
RAPP-containing arrest peptides induce translational stalling by short circuiting the ribosomal peptidyltransferase activity
Translation of RAPP (Arg-AlaPro-Pro) motifs induces ribosome stalling. Here, structures of RAPP-stalled ribosomes reveal that RAPP motifs short circuit the ribosomal peptidyltransferase activity to induce stalling.
- Martino Morici
- , Sara Gabrielli
- & Daniel N. Wilson
-
Article
| Open Access dCas13-mediated translational repression for accurate gene silencing in mammalian cells
dCas13-mediated translational repression for accurate gene silencing in mammalian cells
Current gene silencing tools can have drawbacks. Here the authors report CRISPRδ, an approach for translational silencing, harnessing catalytically inactive Cas13 proteins (dCas13): they also show that fusion of a translational repressor to dCas13 further improved the performance.
- Antonios Apostolopoulos
- , Naohiro Kawamoto
- & Shintaro Iwasaki
-
Article
| Open Access Widespread stable noncanonical peptides identified by integrated analyses of ribosome profiling and ORF features
Widespread stable noncanonical peptides identified by integrated analyses of ribosome profiling and ORF features
By developing computational algorithms, the authors annotated translated open reading frames in five eukaryotes and found many stable peptides are encoded by putative ‘noncoding’ regions of genomes.
- Haiwang Yang
- , Qianru Li
- & Zhe Ji
-
Article
| Open Access Transient disome complex formation in native polysomes during ongoing protein synthesis captured by cryo-EM
Transient disome complex formation in native polysomes during ongoing protein synthesis captured by cryo-EM
Direct visualization of short-lived intermediates during active protein synthesis remains challenging. Here, the authors structurally capture transient translation intermediates to uncover temporary disome formation during ribosome collisions.
- Timo Flügel
- , Magdalena Schacherl
- & Christian M. T. Spahn
-
Article
| Open Access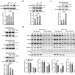 Stalled translation by mitochondrial stress upregulates a CNOT4-ZNF598 ribosomal quality control pathway important for tissue homeostasis
Stalled translation by mitochondrial stress upregulates a CNOT4-ZNF598 ribosomal quality control pathway important for tissue homeostasis
Ribosome associated quality control (RQC) is a new area of biological investigation with emerging connection to a broad range of diseases. Here authors show that mitochondrial stress can upregulate a new RQC pathway important for tissue homeostasis.
- Ji Geng
- , Shuangxi Li
- & Bingwei Lu
-
Article
| Open Access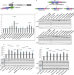 Decryption of sequence, structure, and functional features of SINE repeat elements in SINEUP non-coding RNA-mediated post-transcriptional gene regulation
Decryption of sequence, structure, and functional features of SINE repeat elements in SINEUP non-coding RNA-mediated post-transcriptional gene regulation
Here the authors elucidate structure-function relationships of SINEUPs, antisense long non-coding RNAs that through SINE repeat elements positively regulate protein translation of mRNAs they pair with. SINEUP’s functional domains do not share common ancestors.
- Harshita Sharma
- , Matthew N. Z. Valentine
- & Piero Carninci
-
Article
| Open Access Cryo- EM structure of the mycobacterial 70S ribosome in complex with ribosome hibernation promotion factor RafH
Cryo- EM structure of the mycobacterial 70S ribosome in complex with ribosome hibernation promotion factor RafH
Ribosome hibernation is a key survival strategy bacteria adapt under stress. Here, cryo- EM structure of mycobacterial 70S ribosome with hypoxia stress-induced factor RafH suggests the molecular mechanism of RafH-induced ribosome hibernation.
- Niraj Kumar
- , Shivani Sharma
- & Prem S. Kaushal
-
Article
| Open Access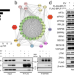 Remodeling of the ribosomal quality control and integrated stress response by viral ubiquitin deconjugases
Remodeling of the ribosomal quality control and integrated stress response by viral ubiquitin deconjugases
Here, the authors show how the vDUB from the large tegument protein from the human herpes virus can reprogram translation in host cells by modulating the activity of the ribosome quality machinery and activating the integrated stress response.
- Jiangnan Liu
- , Noemi Nagy
- & Maria G. Masucci
-
Article
| Open Access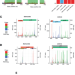 The distinct translational landscapes of gram-negative Salmonella and gram-positive Listeria
The distinct translational landscapes of gram-negative Salmonella and gram-positive Listeria
In this work, Bryant and Lastovka et al. utilise advanced ribosome profiling and transcriptomics techniques, to reveal distinct translation control mechanisms in Salmonella and Listeria, two highly divergent bacterial species.
- Owain J. Bryant
- , Filip Lastovka
- & Betty Y. -W. Chung
-
Article
| Open Access Characterization of nucleolar SUMO isopeptidases unveils a general p53-independent checkpoint of impaired ribosome biogenesis
Characterization of nucleolar SUMO isopeptidases unveils a general p53-independent checkpoint of impaired ribosome biogenesis
Ribosome biogenesis is tightly coordinated with cell-cycle progression. By characterizing the SUMO isopeptidases SENP3/SENP5, Doenig et al. identify a long-sought p53-independent impaired ribosome checkpoint that converges on downregulation of CDK6.
- Judith Dönig
- , Hannah Mende
- & Stefan Müller
-
Article
| Open Access Structural insights into the role of GTPBP10 in the RNA maturation of the mitoribosome
Structural insights into the role of GTPBP10 in the RNA maturation of the mitoribosome
The biogenesis of ribosomes is a highly coordinated process. Here, Nguyen et al. uncover how the mitochondria-specific interplay of the GTPases GTPBP10 and GTPBP7 ensures proper maturation of the catalytic RNA center of the human mitoribosome.
- Thu Giang Nguyen
- , Christina Ritter
- & Eva Kummer
-
Article
| Open Access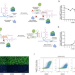 Rational design of microRNA-responsive switch for programmable translational control in mammalian cells
Rational design of microRNA-responsive switch for programmable translational control in mammalian cells
Artificial regulation of translation by intracellular RNAs has many potential applications. Here, authors design a platform capable of miRNA-triggered upregulation or downregulation using a single RNA construct, and demonstrate its use in constructing logic gates and cell-type classifiers.
- Hui Ning
- , Gan Liu
- & Zhen Xie
-
Article
| Open Access The structure of a hibernating ribosome in a Lyme disease pathogen
The structure of a hibernating ribosome in a Lyme disease pathogen
Ribosomes are prime targets for antibiotics in pathogenic bacteria. Here, cryo-electron microscopy reveals features in the Borrelia burgdorferi ribosome that provide insights into ribosome evolution, dormancy, and antibiotic binding.
- Manjuli R. Sharma
- , Swati R. Manjari
- & Nilesh K. Banavali
-
Article
| Open Access RNA-based translation activators for targeted gene upregulation
RNA-based translation activators for targeted gene upregulation
Many diseases are driven by the insufficient expression of critical genes, but few technologies are capable of rescuing these endogenous protein levels. Here, Cao et al. present an RNA-based technology that boosts protein production from endogenous mRNAs by upregulating their translation.
- Yang Cao
- , Huachun Liu
- & Bryan C. Dickinson
-
Article
| Open Access Extensive breaking of genetic code degeneracy with non-canonical amino acids
Extensive breaking of genetic code degeneracy with non-canonical amino acids
Genetic code expansion is limited by the degeneracy of the 61 sense codons which encode for only 20 amino acids. Here, the authors show that by combining hyperaccurate ribosomes and in vitro transcribed tRNAs, dramatic and extensive breaking of sense codon degeneracy can be achieved.
- Clinton A. L. McFeely
- , Bipasana Shakya
- & Matthew C. T. Hartman
-
Article
| Open Access Modulation of translational decoding by m6A modification of mRNA
Modulation of translational decoding by m6A modification of mRNA
m6A is an mRNA modification that slows down translation elongation. Here, Jain et al. show that m6A delays decoding and increases tRNA drop-off from the ribosome by favoring alternative codon conformations that are rejected by the ribosome.
- Sakshi Jain
- , Lukasz Koziej
- & Marina V. Rodnina
-
Article
| Open Access Molecular basis of the pleiotropic effects by the antibiotic amikacin on the ribosome
Molecular basis of the pleiotropic effects by the antibiotic amikacin on the ribosome
Here the authors use fast kinetics, X-ray crystallography, and cryo-EM to uncover the mechanism of ribosome inhibition by amikacin and kanamycin. They find that amikacin binds near the P-site tRNA, offering new strategies to fight antibiotic resistance.
- Savannah M. Seely
- , Narayan P. Parajuli
- & Matthieu G. Gagnon
-
Article
| Open Access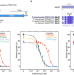 Regulation of the macrolide resistance ABC-F translation factor MsrD
Regulation of the macrolide resistance ABC-F translation factor MsrD
Antibiotic resistance ABC-F factors protect the ribosome from important antibiotics. Here, for one of them, the authors describe its molecular regulation that involves ribosome stalling by antibiotics for which the factor provides resistance.
- Corentin R. Fostier
- , Farès Ousalem
- & Grégory Boël
-
Article
| Open Access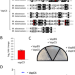 Mycobacterium abscessus VapC5 toxin potentiates evasion of antibiotic killing by ribosome overproduction and activation of multiple resistance pathways
Mycobacterium abscessus VapC5 toxin potentiates evasion of antibiotic killing by ribosome overproduction and activation of multiple resistance pathways
Mycobacterium abscessus (Mab) infections are difficult to clear with antibiotics. Here the authors show that clinical Mab strains can acquire a toxin-antitoxin system that enhances survival upon treatment with current first-line antibiotics through depletion of tRNASerCGA and subsequent ribosome overproduction.
- Eduardo A. Troian
- , Heather M. Maldonado
- & Nancy A. Woychik
-
Article
| Open Access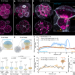 Apicobasal RNA asymmetries regulate cell fate in the early mouse embryo
Apicobasal RNA asymmetries regulate cell fate in the early mouse embryo
How do cells of the preimplantation mouse embryo make decisions? Here the authors discovered that the spatial sorting of mRNAs, tRNA, rRNAs and organelles lead to localized translation, conducive for cell fate allocation and embryonic development.
- Azelle Hawdon
- , Niall D. Geoghegan
- & Jennifer Zenker
-
Article
| Open Access Quality control of protein synthesis in the early elongation stage
Quality control of protein synthesis in the early elongation stage
Peptidyl-tRNAs (pep-tRNAs) frequently dissociate from ribosome, called as pep-tRNA drop-off. But, its function remained unclear. The authors proposed a new quality control mechanism of protein synthesis by active rejection of miscoded pep-tRNAs in the early stage of translation.
- Asuteka Nagao
- , Yui Nakanishi
- & Tsutomu Suzuki
-
Article
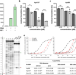
| Open Access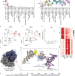 RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function
RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function
RPL3L is a paralog of the ribosomal protein RPL3 and specifically expressed in heart and skeletal muscle. Here, the authors show that RPL3L-containing ribosomes regulate translation elongation dynamics especially for the transcripts related to cardiac muscle contraction.
- Chisa Shiraishi
- , Akinobu Matsumoto
- & Keiichi I. Nakayama
-
Article
| Open Access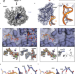 rRNA methylation by Spb1 regulates the GTPase activity of Nog2 during 60S ribosomal subunit assembly
rRNA methylation by Spb1 regulates the GTPase activity of Nog2 during 60S ribosomal subunit assembly
Regulation of 60S biogenesis remains poorly understood. Using cryo-EM, the authors show that failure of Spb1 to methylate the A-loop nucleotide G2922 prematurely activates the GTPase Nog2, suggesting that Spb1 and Nog2 form a kinetic checkpoint during ribosome maturation.
- Kamil Sekulski
- , Victor Emmanuel Cruz
- & Jan P. Erzberger
-
Article
| Open Access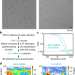 The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services
The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services
Developments in cryo-EM sample preparation and data collection are pivotal for structure determination. Fromm et al. present a 1.55 Å structure of the translating bacterial ribosome that provides new insights on its function and may allow for more precise structure-based drug design.
- Simon A. Fromm
- , Kate M. O’Connor
- & Simone Mattei
-
Article
| Open Access Structural basis for clearing of ribosome collisions by the RQT complex
Structural basis for clearing of ribosome collisions by the RQT complex
Ribosome collisions serve as proxy for aberrant translation to initiate rescue and quality control pathways. Here, authors elucidate the molecular mechanism of collided ribosome clearance by the ribosome quality control trigger complex.
- Katharina Best
- , Ken Ikeuchi
- & Roland Beckmann
-
Article
| Open Access Antibiotic thermorubin tethers ribosomal subunits and impedes A-site interactions to perturb protein synthesis in bacteria
Antibiotic thermorubin tethers ribosomal subunits and impedes A-site interactions to perturb protein synthesis in bacteria
Thermorubin is a ribosome-targeting antibiotic. Here, using fast-kinetics and cryoEM, the authors reveal that thermorubin primarily blocks ribosome-recycling by tethering the ribosomal subunits besides impeding translation elongation and termination steps.
- Narayan Prasad Parajuli
- , Andrew Emmerich
- & Suparna Sanyal
-
Article
| Open Access RSL24D1 sustains steady-state ribosome biogenesis and pluripotency translational programs in embryonic stem cells
RSL24D1 sustains steady-state ribosome biogenesis and pluripotency translational programs in embryonic stem cells
Pluripotency is coordinated at multiple levels of gene expression. Here the authors show that ribosome biogenesis is tightly regulated in embryonic stem cells (ESC) to control the translation of transcription and chromatin factors and dictate ESC fate.
- Sébastien Durand
- , Marion Bruelle
- & Mathieu Gabut
-
Article
| Open Access DAP5 enables main ORF translation on mRNAs with structured and uORF-containing 5′ leaders
DAP5 enables main ORF translation on mRNAs with structured and uORF-containing 5′ leaders
RNA structure and upstream ORFs can modulate the translation of human transcripts. Here, the authors show that structured mRNAs with pervasive uORF translation require DAP5, an eIF4G-like protein, for translation of the main ORF.
- Ramona Weber
- , Leon Kleemann
- & Cátia Igreja
-
Article
| Open Access Structures of the eukaryotic ribosome and its translational states in situ
Structures of the eukaryotic ribosome and its translational states in situ
The translational states of eukaryotic ribosomes have so far been only investigated in vitro. Here, authors obtained the 3.8 Å in situ 80S ribosome structure, the distribution of translational states and unique arrangement of rRNA expansion segments.
- Patrick C. Hoffmann
- , Jan Philipp Kreysing
- & Martin Beck
-
Article
| Open Access Nascent peptide-induced translation discontinuation in eukaryotes impacts biased amino acid usage in proteomes
Nascent peptide-induced translation discontinuation in eukaryotes impacts biased amino acid usage in proteomes
Here the authors find that nascent chains enriched in acidic amino acids destabilize the translating ribosome, eventually leading to stochastic premature termination in eukaryotes as in bacteria. Such risk of premature termination influences the amino acid distribution in the proteomes.
- Yosuke Ito
- , Yuhei Chadani
- & Hideki Taguchi
-
Article
| Open Access Varying strength of selection contributes to the intragenomic diversity of rRNA genes
Varying strength of selection contributes to the intragenomic diversity of rRNA genes
Ribosomal RNA genes are abundant in eukaryotic genomes and code for the universal and essential RNA components of the ribosome. This study uncovers high sequence diversity of the genes within a single species and discusses the contribution of selection in the evolution of ribosomal RNA.
- Daniel Sultanov
- & Andreas Hochwagen
-
Article
| Open Access A nascent peptide code for translational control of mRNA stability in human cells
A nascent peptide code for translational control of mRNA stability in human cells
Here, the authors measure the mRNA levels of thousands of coding sequence motifs in human cells to find that nascent peptides with a combination of β-strand structures and bulky and positively charged sequences reduce mRNA levels by slowing translation.
- Phillip C. Burke
- , Heungwon Park
- & Arvind Rasi Subramaniam
-
Article
| Open Access Structural dynamics of AAA + ATPase Drg1 and mechanism of benzo-diazaborine inhibition
Structural dynamics of AAA + ATPase Drg1 and mechanism of benzo-diazaborine inhibition
Drg1 is a cytoplasmic AAA + ATPase responsible for releasing Rlp24 during ribosome assembly. Here, the authors show that Drg1 structurally and functionally shares many regulatory features with that of Cdc48/p97, suggesting a unfoldase role for Drg1.
- Chengying Ma
- , Damu Wu
- & Ning Gao
-
Article
| Open Access A distinct mammalian disome collision interface harbors K63-linked polyubiquitination of uS10 to trigger hRQT-mediated subunit dissociation
A distinct mammalian disome collision interface harbors K63-linked polyubiquitination of uS10 to trigger hRQT-mediated subunit dissociation
Collided ribosomes are marked by ubiquitination to induce quality control mechanisms. Here, authors show that mammalian disomes form a distinct structural interface, in which uS10 K63-linked polyubiquitination is critical for ribosome dissociation.
- Momoko Narita
- , Timo Denk
- & Toshifumi Inada
-
Article
| Open Access Translational reprogramming in response to accumulating stressors ensures critical threshold levels of Hsp90 for mammalian life
Translational reprogramming in response to accumulating stressors ensures critical threshold levels of Hsp90 for mammalian life
The molecular chaperone Hsp90 decreases with age but whether organisms can physiologically tune expression is unclear. Here, the authors show that mammalian cells can adjust Hsp90 levels to accumulating cellular stress by translational reprogramming.
- Kaushik Bhattacharya
- , Samarpan Maiti
- & Didier Picard
-
Article
| Open Access Structure of a mitochondrial ribosome with fragmented rRNA in complex with membrane-targeting elements
Structure of a mitochondrial ribosome with fragmented rRNA in complex with membrane-targeting elements
Structural and evolutionary analysis of a diverged mitoribosome with rRNA that is split in 13 pieces reveals a non-canonical reduced form of the 5 S, permutation of domain I, and how the fragmentation pattern links to the concept of a primordial ribosome.
- Victor Tobiasson
- , Ieva Berzina
- & Alexey Amunts
-
Article
| Open Access A stem cell roadmap of ribosome heterogeneity reveals a function for RPL10A in mesoderm production
A stem cell roadmap of ribosome heterogeneity reveals a function for RPL10A in mesoderm production
How ribosomes differ in composition and function to regulate gene expression is poorly understood. Here, the authors show that ribosome composition changes during stem cell differentiation and identify a ribosomal protein that regulates production of the mesoderm lineage.
- Naomi R. Genuth
- , Zhen Shi
- & Maria Barna
-
Article
| Open Access Single-cell transcriptome and translatome dual-omics reveals potential mechanisms of human oocyte maturation
Single-cell transcriptome and translatome dual-omics reveals potential mechanisms of human oocyte maturation
Development of methods for simultaneous single cell analysis of transcription and translation is still underway. Here, Hu et al. develop single-cell transcriptome and translatome dual-omics on human oocytes, which enables them to identify OOSP2 as an induction factor during human oocyte maturation.
- Wenqi Hu
- , Haitao Zeng
- & Kehkooi Kee
-
Article
| Open Access Duplicated ribosomal protein paralogs promote alternative translation and drug resistance
Duplicated ribosomal protein paralogs promote alternative translation and drug resistance
Most yeast ribosomal protein genes are duplicated but the functional significance of this duplication remains unclear. This study identifies a natural program where changing the ratio of proteins produced from duplicated genes modifies translation in response to drugs regardless of ribosome number.
- Mustafa Malik Ghulam
- , Mathieu Catala
- & Sherif Abou Elela
-
Article
| Open Access Ribosome impairment regulates intestinal stem cell identity via ZAKɑ activation
Ribosome impairment regulates intestinal stem cell identity via ZAKɑ activation
Intestinal stem cells are responsible for replenishing cells within the high-turnover intestinal epithelium. Here they show that ribosome dynamics affect intestinal stem cell identity through a mechanism that is triggered by changes in nutrient availability.
- Joana Silva
- , Ferhat Alkan
- & William James Faller
-
Article
| Open Access Modulating co-translational protein folding by rational design and ribosome engineering
Modulating co-translational protein folding by rational design and ribosome engineering
The narrow exit tunnel of the ribosome is important for cotranslational protein folding. Here, authors show that their rationally designed and engineered exit tunnel protein loops modulate the free energy of nascent chain dynamics and folding.
- Minkoo Ahn
- , Tomasz Włodarski
- & John Christodoulou
-
Article
| Open Access Altered tRNA dynamics during translocation on slippery mRNA as determinant of spontaneous ribosome frameshifting
Altered tRNA dynamics during translocation on slippery mRNA as determinant of spontaneous ribosome frameshifting
Slippery sequences in mRNA can cause the ribosome to change its reading frame. Using smFRET, Poulis et al. show how reversible fluctuations of peptidyl-tRNA slow down translocation, alter ribosome dynamics, and favor spontaneous ribosome frameshifting.
- Panagiotis Poulis
- , Anoshi Patel
- & Sarah Adio
-
Article
| Open Access Structural remodeling of ribosome associated Hsp40-Hsp70 chaperones during co-translational folding
Structural remodeling of ribosome associated Hsp40-Hsp70 chaperones during co-translational folding
Ribosome associated complex (RAC)- HSP70 (Ssb in yeast) is a eukaryotic chaperone system involved in co-translational folding. Here, authors report structures of RAC-containing ribosomal complexes, which suggest a working model for the dynamic actions of RAC-Ssb during the process.
- Yan Chen
- , Bin Tsai
- & Ning Gao
-
Article
| Open Access Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Initiation factor 2 (IF2) guides the ribosome to the elongation phase of protein synthesis. Here, Basu et al. provide structural insights into how compact IF2-GDP makes way for initiator tRNA accommodation into the peptidyl (P) site of the ribosome.
- Ritwika S. Basu
- , Michael B. Sherman
- & Matthieu G. Gagnon

 Multimodal binding and inhibition of bacterial ribosomes by the antimicrobial peptides Api137 and Api88
Multimodal binding and inhibition of bacterial ribosomes by the antimicrobial peptides Api137 and Api88