Featured
-
-
Article
| Open Access CHIKV infection reprograms codon optimality to favor viral RNA translation by altering the tRNA epitranscriptome
CHIKV infection reprograms codon optimality to favor viral RNA translation by altering the tRNA epitranscriptome
Viruses completely depend on the host translational machinery, but their genomes are often enriched in rare codons and therefore should be translated with poor efficiency. Here, Jungfleisch et al. apply Ribo-Seq and RNASeq to provide a global view on the translational changes occurring during Chikungunya virus (CHIKV) infection. CHIKV infection induces a codon-specific reprogramming of the host translation machinery to favor the translation of viral RNA genomes over host mRNAs via tRNA modification.
- Jennifer Jungfleisch
- , René Böttcher
- & Juana Díez
-
Article
| Open Access Ribosome impairment regulates intestinal stem cell identity via ZAKɑ activation
Ribosome impairment regulates intestinal stem cell identity via ZAKɑ activation
Intestinal stem cells are responsible for replenishing cells within the high-turnover intestinal epithelium. Here they show that ribosome dynamics affect intestinal stem cell identity through a mechanism that is triggered by changes in nutrient availability.
- Joana Silva
- , Ferhat Alkan
- & William James Faller
-
Article
| Open Access Modulating co-translational protein folding by rational design and ribosome engineering
Modulating co-translational protein folding by rational design and ribosome engineering
The narrow exit tunnel of the ribosome is important for cotranslational protein folding. Here, authors show that their rationally designed and engineered exit tunnel protein loops modulate the free energy of nascent chain dynamics and folding.
- Minkoo Ahn
- , Tomasz Włodarski
- & John Christodoulou
-
Article
| Open Access Altered tRNA dynamics during translocation on slippery mRNA as determinant of spontaneous ribosome frameshifting
Altered tRNA dynamics during translocation on slippery mRNA as determinant of spontaneous ribosome frameshifting
Slippery sequences in mRNA can cause the ribosome to change its reading frame. Using smFRET, Poulis et al. show how reversible fluctuations of peptidyl-tRNA slow down translocation, alter ribosome dynamics, and favor spontaneous ribosome frameshifting.
- Panagiotis Poulis
- , Anoshi Patel
- & Sarah Adio
-
Article
| Open Access Pervasive translation of circular RNAs driven by short IRES-like elements
Pervasive translation of circular RNAs driven by short IRES-like elements
Unbiased screen of random sequences identified many short IRES-like elements to drive circular RNA translation and hundreds of rolling circle translation events, suggesting a pervasive cap-independent translation in human transcriptome.
- Xiaojuan Fan
- , Yun Yang
- & Zefeng Wang
-
Article
| Open Access Structural remodeling of ribosome associated Hsp40-Hsp70 chaperones during co-translational folding
Structural remodeling of ribosome associated Hsp40-Hsp70 chaperones during co-translational folding
Ribosome associated complex (RAC)- HSP70 (Ssb in yeast) is a eukaryotic chaperone system involved in co-translational folding. Here, authors report structures of RAC-containing ribosomal complexes, which suggest a working model for the dynamic actions of RAC-Ssb during the process.
- Yan Chen
- , Bin Tsai
- & Ning Gao
-
Article
| Open Access Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Compact IF2 allows initiator tRNA accommodation into the P site and gates the ribosome to elongation
Initiation factor 2 (IF2) guides the ribosome to the elongation phase of protein synthesis. Here, Basu et al. provide structural insights into how compact IF2-GDP makes way for initiator tRNA accommodation into the peptidyl (P) site of the ribosome.
- Ritwika S. Basu
- , Michael B. Sherman
- & Matthieu G. Gagnon
-
Article
| Open Access Codon-specific Ramachandran plots show amino acid backbone conformation depends on identity of the translated codon
Codon-specific Ramachandran plots show amino acid backbone conformation depends on identity of the translated codon
Genetic code redundancies are considered inconsequential to protein structure. This study uncovers a dependence between local amino acid conformation in folded proteins and the identity of the codon from which that amino acid was translated.
- Aviv A. Rosenberg
- , Ailie Marx
- & Alex M. Bronstein
-
Article
| Open Access Ribosome inhibition by C9ORF72-ALS/FTD-associated poly-PR and poly-GR proteins revealed by cryo-EM
Ribosome inhibition by C9ORF72-ALS/FTD-associated poly-PR and poly-GR proteins revealed by cryo-EM
The expansion of GGGGCC repeats in the C9ORF72 gene results in the production of disease causing abnormal proteins with polymeric glycine-arginine (poly-GR) and polymeric proline-arginine (poly-PR). Here the authors demonstrate a structural mechanism of how poly-GR and poly-PR inhibit translation and how they might also perturb ribosome assembly.
- Anna B. Loveland
- , Egor Svidritskiy
- & Andrei A. Korostelev
-
Article
| Open Access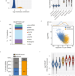 HDLBP binds ER-targeted mRNAs by multivalent interactions to promote protein synthesis of transmembrane and secreted proteins
HDLBP binds ER-targeted mRNAs by multivalent interactions to promote protein synthesis of transmembrane and secreted proteins
RNA binding protein HDLBP (or Vigilin) localizes in the endoplasmic reticulum (ER) membrane. Here the authors show that HDLBP contributes to translation of ER-targeted mRNAs.
- Ulrike Zinnall
- , Miha Milek
- & Markus Landthaler
-
Article
| Open Access Substrate recognition and cryo-EM structure of the ribosome-bound TAC toxin of Mycobacterium tuberculosis
Substrate recognition and cryo-EM structure of the ribosome-bound TAC toxin of Mycobacterium tuberculosis
Toxin-antitoxin systems are widespread in bacteria. Here the authors present structures of M. tuberculosis HigBTAC alone and bound to the ribosome in the presence of native cspA mRNA, shedding light on its mechanism of translation inhibition.
- Moise Mansour
- , Emmanuel Giudice
- & Pierre Genevaux
-
Article
| Open Access Ataluren binds to multiple protein synthesis apparatus sites and competitively inhibits release factor-dependent termination
Ataluren binds to multiple protein synthesis apparatus sites and competitively inhibits release factor-dependent termination
Ataluren is the only nonsense suppressor drug currently approved for clinical use. Here, the authors determine where ataluren binds to the ribosome and how it inhibits termination at nonsense codons.
- Shijie Huang
- , Arpan Bhattacharya
- & Barry S. Cooperman
-
Article
| Open Access Regulation of A-to-I RNA editing and stop codon recoding to control selenoprotein expression during skeletal myogenesis
Regulation of A-to-I RNA editing and stop codon recoding to control selenoprotein expression during skeletal myogenesis
Stop codon recoding is a unique mechanism for selenoprotein biogenesis. Here, the authors report regulatory systems for selenoprotein expression during skeletal muscle differentiation by regulating A-to-I RNA editing and stop codon recoding.
- Yuta Noda
- , Shunpei Okada
- & Tsutomu Suzuki
-
Article
| Open Access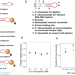 A multiplex platform for small RNA sequencing elucidates multifaceted tRNA stress response and translational regulation
A multiplex platform for small RNA sequencing elucidates multifaceted tRNA stress response and translational regulation
This paper develops a multiplex RNA-seq method that reports tRNA abundance, modification, charging, and fragmentation. Results show stress-induced regulation in translational elongation and association of modification and tRNA fragment biogenesis.
- Christopher P. Watkins
- , Wen Zhang
- & Tao Pan
-
Article
| Open Access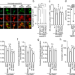 Pantothenate kinase 2 interacts with PINK1 to regulate mitochondrial quality control via acetyl-CoA metabolism
Pantothenate kinase 2 interacts with PINK1 to regulate mitochondrial quality control via acetyl-CoA metabolism
PKAN and PD are two distinct diseases with overlapping pathophysiology. Here, authors show that their pathogenic genes PANK2 and PINK1 interact. PANK2 regulates mitophagy via CoA metabolism, while PINK1 supervises PANK2 translation on mitochondria.
- Yunpeng Huang
- , Zhihui Wan
- & Bing Zhou
-
Article
| Open Access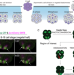 Vasa nucleates asymmetric translation along the mitotic spindle during unequal cell divisions
Vasa nucleates asymmetric translation along the mitotic spindle during unequal cell divisions
Association of mRNA translation with the mitotic spindle is thought to be involved in localized production of cell fate determinants. Here, the authors show Vasa facilitates asymmetric translation, which contributes to differential regulation during sea urchin embryogenesis.
- Ana Fernandez-Nicolas
- , Alicia Uchida
- & Mamiko Yajima
-
Article
| Open Access Structural basis for PoxtA-mediated resistance to phenicol and oxazolidinone antibiotics
Structural basis for PoxtA-mediated resistance to phenicol and oxazolidinone antibiotics
PoxtA confers resistance to ribosome-targeting oxazolidinone (linezolid) and chloramphenicol antibiotics. Here, Crowe-McAuliffe et al. provide structural insights into how binding of PoxtA to the ribosome indirectly promotes drug dissociation.
- Caillan Crowe-McAuliffe
- , Victoriia Murina
- & Vasili Hauryliuk
-
Article
| Open Access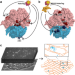 Direct measurements of mRNA translation kinetics in living cells
Direct measurements of mRNA translation kinetics in living cells
Metelev et al. use single-molecule tracking to study kinetics of translation directly in E. coli cells, and how it is affected by translation inhibitors and rRNA mutations. Their results support widespread 70S re-initiation on mRNAs.
- Mikhail Metelev
- , Erik Lundin
- & Magnus Johansson
-
Article
| Open Access eIF6 rebinding dynamically couples ribosome maturation and translation
eIF6 rebinding dynamically couples ribosome maturation and translation
Jaako et al. discover a conserved tier of translational control that dynamically couples ribosome assembly and recycling. This mechanism is corrupted in an inherited bone marrow failure disorder associated with an increased risk of blood cancer.
- Pekka Jaako
- , Alexandre Faille
- & Alan J. Warren
-
Article
| Open Access Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
The authors develop an RNA sequencing-based platform, PERSIST-seq, to simultaneously delineate in-cell mRNA stability, ribosome load, and in-solution stability of a diverse mRNA library to derive design principles for improved mRNA therapeutics.
- Kathrin Leppek
- , Gun Woo Byeon
- & Rhiju Das
-
Article
| Open Access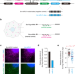 Single-molecule imaging of microRNA-mediated gene silencing in cells
Single-molecule imaging of microRNA-mediated gene silencing in cells
For decades, miRNAs have been studied primarily by ensemble methods, where a bulk collection of molecules is measured outside cells. Here, Kobayashi and Singer report methods to image miRNA function at the single-molecule level inside cells.
- Hotaka Kobayashi
- & Robert H. Singer
-
Article
| Open Access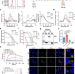 RNA G-quadruplex in TMPRSS2 reduces SARS-CoV-2 infection
RNA G-quadruplex in TMPRSS2 reduces SARS-CoV-2 infection
Understanding the mechanisms of SARS-CoV-2 infection is important to control the pandemic. Here the authors show the biological and pathological role of RNA G-quadruplex structure in both SARS-CoV-2 genome and host factors, particularly TMPRSS2.
- Geng Liu
- , Wenya Du
- & Xianghui Fu
-
Article
| Open Access Co-translational assembly orchestrates competing biogenesis pathways
Co-translational assembly orchestrates competing biogenesis pathways
The biogenesis of nuclear pores imposes a logistic challenge for cells. Here, the authors investigate structural motifs for co-translational interactions in nucleoporins and find that co-translational assembly events differ between paralogous assembly pathways thus contributing to faithful assembly.
- Maximilian Seidel
- , Anja Becker
- & Martin Beck
-
Article
| Open Access ATF-4 and hydrogen sulfide signalling mediate longevity in response to inhibition of translation or mTORC1
ATF-4 and hydrogen sulfide signalling mediate longevity in response to inhibition of translation or mTORC1
The authors showed that, in C. elegans, inhibition of translation or mTORC1 increases ATF-4 expression independently of ISR signalling. ATF-4 promotes longevity by increasing hydrogen sulfide production by the enzyme CTH-2.
- Cyril Statzer
- , Jin Meng
- & Collin Y. Ewald
-
Article
| Open Access A late-stage assembly checkpoint of the human mitochondrial ribosome large subunit
A late-stage assembly checkpoint of the human mitochondrial ribosome large subunit
Rebelo-Guiomar et al. unveil late stage assembly intermediates of the human mitochondrial ribosome by inactivating the methyltransferase MRM2 in cells. Absence of MRM2 impairs organismal homeostasis, while its catalytic activity is dispensable for mitoribosomal biogenesis.
- Pedro Rebelo-Guiomar
- , Simone Pellegrino
- & Michal Minczuk
-
Article
| Open Access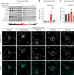 Cyclin B/CDK1 and Cyclin A/CDK2 phosphorylate DENR to promote mitotic protein translation and faithful cell division
Cyclin B/CDK1 and Cyclin A/CDK2 phosphorylate DENR to promote mitotic protein translation and faithful cell division
The cell cycle regulates translation during mitosis by controlling DENR stability. Here, the authors show the non-canonical translation initiation complex DENR·MCTS1 is phosphorylated during mitosis by CDK1 and 2, enabling the translation of genes needed for proper mitotic progression.
- Katharina Clemm von Hohenberg
- , Sandra Müller
- & Aurelio A. Teleman
-
Article
| Open Access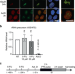 Alteration of ribosome function upon 5-fluorouracil treatment favors cancer cell drug-tolerance
Alteration of ribosome function upon 5-fluorouracil treatment favors cancer cell drug-tolerance
Different mechanisms have been reported to explain resistance to chemotherapy in cancer. Here, the authors show that the chemotherapeutic drug 5-fluorouracil alters the function of ribosomes to promote pro-survival gene translation leading to chemotherapy resistance.
- Gabriel Therizols
- , Zeina Bash-Imam
- & Jean-Jacques Diaz
-
Article
| Open Access Mutations in Hcfc1 and Ronin result in an inborn error of cobalamin metabolism and ribosomopathy
Mutations in Hcfc1 and Ronin result in an inborn error of cobalamin metabolism and ribosomopathy
Combined methylmalonic acidemia (MMA) and hyperhomocysteinemias are inborn errors of vitamin B12 metabolism, and mutations in the transcriptional regulators HCFC1 and RONIN (THAP11) underlie some forms of these disorders. Here the authors generated mouse models of a human syndrome due to mutations in RONIN (THAP11) and HCFC1, and show that this syndrome is both an inborn error of vitamin B12 metabolism and displays some features of ribosomopathy.
- Tiffany Chern
- , Annita Achilleos
- & Ross A. Poché
-
Article
| Open Access Targeted editing and evolution of engineered ribosomes in vivo by filtered editing
Targeted editing and evolution of engineered ribosomes in vivo by filtered editing
Genome editing methods are limited by the inability to selectively edit repetitive sequences. Here the authors demonstrate precise editing of a repetitive genetic element, a ribosome, while avoiding edits to native sites sharing identical sequence.
- Felix Radford
- , Shane D. Elliott
- & Farren J. Isaacs
-
Article
| Open Access Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP
Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP
EF-G drives ribosomal translocation along mRNA. Time-resolved cryo-EM captured translocation with EF-G•GTP—without inhibitors—revealing how EF-G uses ribosome fluctuations to drive translocation and GTP hydrolysis to leave at the right moment.
- Christine E. Carbone
- , Anna B. Loveland
- & Andrei A. Korostelev
-
Article
| Open Access The short isoform of the host antiviral protein ZAP acts as an inhibitor of SARS-CoV-2 programmed ribosomal frameshifting
The short isoform of the host antiviral protein ZAP acts as an inhibitor of SARS-CoV-2 programmed ribosomal frameshifting
Programmed ribosomal frameshifting (PRF) occurs in many viruses including SARS-CoV-2 to allow the translation of multiple proteins from a single transcript. Here, the authors identify the human short isoform of the zinc-finger antiviral protein (ZAP-S) as a direct regulator of PRF in SARS-CoV-2 that severely impairs SARS-CoV-2 frameshifting in cells and directly interacts with the SARS-CoV-2 RNA; interfering with the folding of the frameshift RNA element.
- Matthias M. Zimmer
- , Anuja Kibe
- & Neva Caliskan
-
Article
| Open Access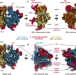 How to build a ribosome from RNA fragments in Chlamydomonas mitochondria
How to build a ribosome from RNA fragments in Chlamydomonas mitochondria
Mitoribosomes are remarkably diverse in their structures and compositions. Here the authors combine biochemistry, genetics, single particle cryo-electron microscopy and in situ cryo-electron tomography to reveal the mitochondrial ribosome of Chlamydomonas reinhardtii as an extreme example of evolution and species-specific adaptation.
- Florent Waltz
- , Thalia Salinas-Giegé
- & Yaser Hashem
-
Article
| Open Access Structural and molecular basis for Cardiovirus 2A protein as a viral gene expression switch
Structural and molecular basis for Cardiovirus 2A protein as a viral gene expression switch
Many RNA viruses employ programmed –1 ribosomal frameshifting (PRF) to expand their coding capacity and optimize production of viral proteins. Here, the authors report structural and biophysical analysis of protein 2A from a cardiovirus, with insights into the mechanism of its PRF-stimulatory function.
- Chris H. Hill
- , Lukas Pekarek
- & Ian Brierley
-
Article
| Open Access eIF2B-capturing viral protein NSs suppresses the integrated stress response
eIF2B-capturing viral protein NSs suppresses the integrated stress response
Here the authors show that a viral protein interferes with the binding of phosphorylated eIF2 to eIF2B, thereby suppressing the host integrated stress response (ISR). This suppression of the ISR abrogates translational changes of the host and ameliorates neurite degradation under stress.
- Kazuhiro Kashiwagi
- , Yuichi Shichino
- & Takuhiro Ito
-
Article
| Open Access Cilia locally synthesize proteins to sustain their ultrastructure and functions
Cilia locally synthesize proteins to sustain their ultrastructure and functions
Cilia are microtubule-based organelles containing proteins transported from the cell body. Here, the authors show that the multicilia of mouse ependymal cells contain ribosomal components, tubulin mRNA,18 S rRNA and nascent tubulin peptides, suggesting local translation in the ciliary compartment.
- Kai Hao
- , Yawen Chen
- & Xueliang Zhu
-
Article
| Open Access A DAP5/eIF3d alternate mRNA translation mechanism promotes differentiation and immune suppression by human regulatory T cells
A DAP5/eIF3d alternate mRNA translation mechanism promotes differentiation and immune suppression by human regulatory T cells
The differentiation of naive T cells to immune suppressing induced regulatory T (iTreg) cells requires TGF-beta-1 and downregulation of mTORC1 activity, which inhibits mRNA translation. Here the authors show that iTreg cell differentiation uses an alternate mRNA translation mechanism involving translation factors DAP5 and eIF3d.
- Viviana Volta
- , Sandra Pérez-Baos
- & Robert J. Schneider
-
Article
| Open Access Functionally distinct roles for eEF2K in the control of ribosome availability and p-body abundance
Functionally distinct roles for eEF2K in the control of ribosome availability and p-body abundance
Processing bodies are phase separated compartments enriched in translationally repressed mRNAs. Here, Smith et al. show that, in sensory neurons, eukaryotic elongation factor 2 kinase (eEF2K) plays key roles in the regulation of processing body abundance and the formation of translationally inactive ribosomes.
- Patrick R. Smith
- , Sarah Loerch
- & Zachary T. Campbell
-
Article
| Open Access Bi-directional ribosome scanning controls the stringency of start codon selection
Bi-directional ribosome scanning controls the stringency of start codon selection
Start codon selection is commonly thought to occur through the unidirectional scanning of the mRNA by the 40 S ribosome. Here the authors provide evidence that the pre-initiation complex can backslide on the mRNA to initiate translation at upstream AUG codons.
- Yifei Gu
- , Yuanhui Mao
- & Shu-Bing Qian
-
Article
| Open Access An integrative proteomics method identifies a regulator of translation during stem cell maintenance and differentiation
An integrative proteomics method identifies a regulator of translation during stem cell maintenance and differentiation
To characterize molecular changes during cell type transitions, the authors develop a method to simultaneously measure protein expression and thermal stability changes. They apply this approach to study differences between human pluripotent stem cells, their progenies, parental and allogeneic cells.
- Pierre Sabatier
- , Christian M. Beusch
- & Roman A. Zubarev
-
Article
| Open Access Nascent chains can form co-translational folding intermediates that promote post-translational folding outcomes in a disease-causing protein
Nascent chains can form co-translational folding intermediates that promote post-translational folding outcomes in a disease-causing protein
Alpha-1-antitrypsin (AAT) deficiency results from misfolding-prone AAT variants. Here the authors show that AAT forms co-translational folding intermediates on the ribosome that persist upon release and determine its folding fate. They show too that the ribosome can also modulate misfolding-prone AAT intermediates during their synthesis.
- Elena Plessa
- , Lien P. Chu
- & Lisa D. Cabrita
-
Article
| Open Access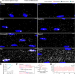 Microtubule-based transport is essential to distribute RNA and nascent protein in skeletal muscle
Microtubule-based transport is essential to distribute RNA and nascent protein in skeletal muscle
It is increasingly recognised that the spatial localisation of RNA is important for proper cellular function. Here, the authors investigate RNA localisation in skeletal muscle and develop methods to show that global active transport of RNA is required to maintain dispersion of gene products in the large muscle syncytium.
- Lance T. Denes
- , Chase P. Kelley
- & Eric T. Wang
-
Article
| Open Access Structural mechanism of GTPase-powered ribosome-tRNA movement
Structural mechanism of GTPase-powered ribosome-tRNA movement
Movement of the ribosome along an mRNA requires the universally-conserved translocase (EF-G in bacteria) that couples GTP hydrolysis to directed movement. Here the authors use time-resolved Cryo-EM to visualize the GTPase-powered step on native translocating ribosomes and capture key translocation intermediates.
- Valentyn Petrychenko
- , Bee-Zen Peng
- & Niels Fischer
-
Article
| Open Access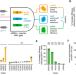 Multiplex suppression of four quadruplet codons via tRNA directed evolution
Multiplex suppression of four quadruplet codons via tRNA directed evolution
Genetic code expansion strategies are limited to specific codons that can be reassigned to new amino acids. Here the authors show that quadruplet-decoding tRNAs (qtRNAs) can be rapidly discovered and evolved to decode new quadruplet codons, enabling four independent decoding events in a single protein in living cells.
- Erika A. DeBenedictis
- , Gavriela D. Carver
- & Ahmed H. Badran
-
Article
| Open Access Pathway of Hsp70 interactions at the ribosome
Pathway of Hsp70 interactions at the ribosome
Here, the authors use in vivo site-specific crosslinking to provide molecular-level insight into how the fungal Hsp70 chaperone system — the Ssb:Ssz1:Zuo1 triad — assists the folding process for the nascent peptide chain emerging from the ribosome tunnel.
- Kanghyun Lee
- , Thomas Ziegelhoffer
- & Elizabeth A. Craig
-
Article
| Open Access Directed evolution of rRNA improves translation kinetics and recombinant protein yield
Directed evolution of rRNA improves translation kinetics and recombinant protein yield
Ribosome kinetics are rate-limiting for protein synthesis. Here the authors evolve diverse 16S rRNAs for enhanced protein synthesis rates and genetic code expansion efficiencies in vivo.
- Fan Liu
- , Siniša Bratulić
- & Ahmed H. Badran
-
Article
| Open Access Structural basis for the tryptophan sensitivity of TnaC-mediated ribosome stalling
Structural basis for the tryptophan sensitivity of TnaC-mediated ribosome stalling
Bacteria adjust the expression of some of their metabolic enzymes through metabolite-sensing ribosome nascent chain complexes. Here the authors present a cryo-EM structure of an E. coli ribosome stalled during translation of the TnaC leader peptide and propose a model for L-Trp dependent ribosome stalling where L-Trp competes with release factor 2 for binding to the TnaC-ribosome complex.
- Anne-Xander van der Stel
- , Emily R. Gordon
- & C. Axel Innis
-
Article
| Open Access A high-resolution temporal atlas of the SARS-CoV-2 translatome and transcriptome
A high-resolution temporal atlas of the SARS-CoV-2 translatome and transcriptome
Here, Kim et al. apply various sequencing techniques (RPF-seq, QTI-seq, mRNA-seq, sRNA-seq) to unravel the high-resolution, longitudinal translatome and transcriptome of SARS-CoV-2. They identify a translation initiation site in the leader sequence of all genomic and subgenomic RNAs and show its relevance for the SARS-CoV-2 translatome.
- Doyeon Kim
- , Sukjun Kim
- & Daehyun Baek
-
Article
| Open Access Humans and other commonly used model organisms are resistant to cycloheximide-mediated biases in ribosome profiling experiments
Humans and other commonly used model organisms are resistant to cycloheximide-mediated biases in ribosome profiling experiments
Ribosome profiling has become the gold standard to analyze mRNA translation dynamics, and the translation inhibitor cycloheximide (CHX) is often used in its application. Here the authors systematically demonstrate that CHX does not bias the outcome of ribosome profiling experiments in most organisms.
- Puneet Sharma
- , Jie Wu
- & Sebastian A. Leidel
-
Article
| Open Access Somatic genetic rescue of a germline ribosome assembly defect
Somatic genetic rescue of a germline ribosome assembly defect
Shwachman-Diamond syndrome (SDS) is a leukemia predisposition disorder that is caused by defective release of eIF6 during ribosome assembly. Here the authors show that acquired somatic EIF6 mutations are frequent in the hematopoietic cells from individuals with SDS and provide a selective advantage over non-modified cells.
- Shengjiang Tan
- , Laëtitia Kermasson
- & Patrick Revy

 Duplicated ribosomal protein paralogs promote alternative translation and drug resistance
Duplicated ribosomal protein paralogs promote alternative translation and drug resistance