Article
|
Open Access
Featured
-
-
Article
| Open Access Engineered tRNAs suppress nonsense mutations in cells and in vivo
Engineered tRNAs suppress nonsense mutations in cells and in vivo
Suppressor tRNAs adapted to the amino acid that they carry enable readthrough of premature termination codons introduced by nonsense mutations and show potential for the treatment of genetic diseases such as cystic fibrosis.
- Suki Albers
- , Elizabeth C. Allen
- & Zoya Ignatova
-
Article |
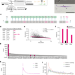 Short tRNA anticodon stem and mutant eRF1 allow stop codon reassignment
Short tRNA anticodon stem and mutant eRF1 allow stop codon reassignment
Analyses of in-frame stop codons in protein-coding genes of Blastocrithidia nonstop with all three stop codons reassigned reveal a mechanism for UGA reassignment in eukaryotes involving shortening of the tRNA anticodon stem and a mutant eRF1 release factor.
- Ambar Kachale
- , Zuzana Pavlíková
- & Julius Lukeš
-
Article |
 Structural basis of regulated m7G tRNA modification by METTL1–WDR4
Structural basis of regulated m7G tRNA modification by METTL1–WDR4
Structures of the human METTL1–WDR4 complex are revealed, providing molecular insights into substrate recognition, modification and catalytic regulation by the N7-methylguanosine methyltransferase complex.
- Jiazhi Li
- , Longfei Wang
- & Richard I. Gregory
-
Article
| Open Access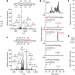 Reversible RNA phosphorylation stabilizes tRNA for cellular thermotolerance
Reversible RNA phosphorylation stabilizes tRNA for cellular thermotolerance
Reversible internal RNA phosphrylation contributes to thermal stability and nuclease resistance of tRNA, and cellular thermotolerance of hyperthermophiles.
- Takayuki Ohira
- , Keiichi Minowa
- & Tsutomu Suzuki
-
Article |
 Valine tRNA levels and availability regulate complex I assembly in leukaemia
Valine tRNA levels and availability regulate complex I assembly in leukaemia
Restriction of dietary valine reduces growth of T cell acute lymphoblastic leukaemia through altered valine tRNA biogenesis and reduced translation of mRNAs that encode subunits of mitochondrial complex I.
- Palaniraja Thandapani
- , Andreas Kloetgen
- & Iannis Aifantis
-
Letter |
 eIF5B gates the transition from translation initiation to elongation
eIF5B gates the transition from translation initiation to elongation
Single-molecule dynamics reveal that the GTPase activity of eukaryotic initiation factor eIF5B serves as a kinetic checkpoint for the transition from translation initiation to elongation.
- Jinfan Wang
- , Alex G. Johnson
- & Joseph D. Puglisi
-
Letter |
 Codon-specific translation reprogramming promotes resistance to targeted therapy
Codon-specific translation reprogramming promotes resistance to targeted therapy
Enzymes that catalyse modifications of wobble uridine 34 tRNA are essential for the survival of melanoma cells that rely on HIF1α-dependent metabolism through codon-dependent regulation of the translation of HIF1A mRNA.
- Francesca Rapino
- , Sylvain Delaunay
- & Pierre Close
-
Article |
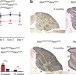 ANKRD16 prevents neuron loss caused by an editing-defective tRNA synthetase
ANKRD16 prevents neuron loss caused by an editing-defective tRNA synthetase
ANKRD16 attenuates neurodegeneration induced by a mutation in the editing domain of alanyl tRNA synthetase by directly accepting mis-activated serine from the synthetase before transfer to the tRNA, establishing a new mechanism by which editing defects are prevented.
- My-Nuong Vo
- , Markus Terrey
- & Susan L. Ackerman
-
Letter |
 Mitochondrial translation requires folate-dependent tRNA methylation
Mitochondrial translation requires folate-dependent tRNA methylation
Mammalian mitochondria use folate-bound one-carbon units generated by the enzyme SHMT2 to methylate tRNA, and this modification is required for mitochondrial translation and thus oxidative phosphorylation.
- Raphael J. Morscher
- , Gregory S. Ducker
- & Joshua D. Rabinowitz
-
Article |
 The selective tRNA aminoacylation mechanism based on a single G•U pair
The selective tRNA aminoacylation mechanism based on a single G•U pair
X-ray crystal structures of a tRNA synthetase bound to wild-type and mutant alanine tRNAs reveal the structural basis for selectivity.
- Masahiro Naganuma
- , Shun-ichi Sekine
- & Shigeyuki Yokoyama
-
Letter |
 Analysis of orthologous groups reveals archease and DDX1 as tRNA splicing factors
Analysis of orthologous groups reveals archease and DDX1 as tRNA splicing factors
Using a phylogenetic approach, the protein archease is identified as being a subunit of the human transfer RNA splicing ligase, and found to be necessary for full ligase activity, in cooperation with DDX1.
- Johannes Popow
- , Jennifer Jurkin
- & Javier Martinez
-
Letter |
 Unusual base pairing during the decoding of a stop codon by the ribosome
Unusual base pairing during the decoding of a stop codon by the ribosome
Here, the structure of the 30S ribosomal subunit and the 70S ribosome in complex with a messenger RNA with pseudouridine in the place of uridine reveals unexpected base pairing.
- Israel S. Fernández
- , Chyan Leong Ng
- & V. Ramakrishnan
-
Letter |
 Structure-guided discovery of the metabolite carboxy-SAM that modulates tRNA function
Structure-guided discovery of the metabolite carboxy-SAM that modulates tRNA function
Members of the SAM-dependent methyltransferase superfamily are involved in the modification of wobble uridine to 5-oxacetyl uridine in Gram-negative bacteria; CmoA converts SAM to carboxy-SAM (Cx-SAM; a metabolite that was unknown previously), and CmoB uses Cx-SAM to convert 5-hydroxyuridine to 5-oxyacetyl uridine in tRNA.
- Jungwook Kim
- , Hui Xiao
- & Steven C. Almo
-
Letter |
 Non-optimal codon usage affects expression, structure and function of clock protein FRQ
Non-optimal codon usage affects expression, structure and function of clock protein FRQ
The frq gene, essential for circadian clock function, is shown to differ from most other genes in Neurospora by exhibiting non-optimal codon usage; by contrast, optimization of codon usage is unexpectedly found to affect the structure and function of the coded protein, subsequently impairing circadian feedback loops.
- Mian Zhou
- , Jinhu Guo
- & Yi Liu
-
Letter |
 A new understanding of the decoding principle on the ribosome
A new understanding of the decoding principle on the ribosome
An integrated mechanism for decoding is proposed, based on six X-ray structures of the 70S ribosome determined at 3.1–3.4 Å resolution, modelling cognate or near-cognate states of the decoding centre at the proofreading step.
- Natalia Demeshkina
- , Lasse Jenner
- & Gulnara Yusupova
-
Letter |
 Different substrate-dependent transition states in the active site of the ribosome
Different substrate-dependent transition states in the active site of the ribosome
- Stephan Kuhlenkoetter
- , Wolfgang Wintermeyer
- & Marina V. Rodnina
-
Letter |
 Head swivel on the ribosome facilitates translocation by means of intra-subunit tRNA hybrid sites
Head swivel on the ribosome facilitates translocation by means of intra-subunit tRNA hybrid sites
During translation, tRNAs enter the ribosome and then move sequentially through three sites, known as A, P and E, as they transfer their attached amino acids onto the growing peptide chain. How the ribosome facilitates tRNA translocation between the sites remains largely unknown. Now a study uses multiparticle cryoelectron microscopy of a ribosome bound to the translation elongation factor, EF-G, to get information about tRNA movement. It identifies two new substates and sees that translocation is linked to unratcheting of the 30S ribosomal subunit.
- Andreas H. Ratje
- , Justus Loerke
- & Christian M. T. Spahn
-
Article |
 Structure of a bacterial ribonuclease P holoenzyme in complex with tRNA
Structure of a bacterial ribonuclease P holoenzyme in complex with tRNA
tRNAs are synthesized in a premature form that requires trimming of the 5′ and 3′ ends and modification of specific nucleotides. RNase P, a complex containing a long catalytic RNA and a protein cofactor, catalyses the cleavage that generates the mature 5′ end. Here, the structure of RNase P bound to mature tRNAPhe is solved. Recognition of the leader sequence and its mechanism of cleavage is determined by soaking an oligonucleotide corresponding to the premature 5′ end into the crystal.
- Nicholas J. Reiter
- , Amy Osterman
- & Alfonso Mondragón
-
Letter |
 Two enzymes bound to one transfer RNA assume alternative conformations for consecutive reactions
Two enzymes bound to one transfer RNA assume alternative conformations for consecutive reactions
In most bacteria and all archaea, glutamyl-tRNA synthetase (GluRS) glutamylates both tRNAGlu and tRNAGln; Glu-tRNAGln is then converted to Gln-tRNAGln by an amidotransferase. Here the structure is reported of a bacterial complex containing tRNAGln, GluRS and the amidotransferase GatCAB. The structure provides an explanation for how the enzymes work consecutively: only one can assume a productive state at any time. There also seems to be an intermediary state in which neither enzyme is productive.
- Takuhiro Ito
- & Shigeyuki Yokoyama
-
Letter |
 A ribosome-associating factor chaperones tail-anchored membrane proteins
A ribosome-associating factor chaperones tail-anchored membrane proteins
Tail-anchored proteins have a single transmembrane domain at their carboxy termini and are post-translationally targeted to the endoplasmic reticulum via the cytosolic ATPase TRC40. These authors identify a conserved protein complex called Bat3 complex that is recruited to ribosomes, interacts with the transmembrane domain of newly released tail-anchored proteins and transfers them to TRC40 for subsequent targeting to the endoplasmic reticulum.
- Malaiyalam Mariappan
- , Xingzhe Li
- & Ramanujan S. Hegde
-
Article |
 Real-time tRNA transit on single translating ribosomes at codon resolution
Real-time tRNA transit on single translating ribosomes at codon resolution
Single-molecule studies allow biological processes to be examined one molecule at a time, as they occur. Here, zero-mode waveguides have been used to concentrate reactions in zeptolitre-sized volumes, making it possible to study real-time translocation by the ribosome. The binding of transfer RNAs (tRNAs) to the ribosome could be followed; the results show that tRNA release from the exit site is uncoupled from tRNA binding to the aminoacyl-tRNA site.
- Sotaro Uemura
- , Colin Echeverría Aitken
- & Joseph D. Puglisi
-
Letter |
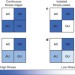 Compensatory evolution in mitochondrial tRNAs navigates valleys of low fitness
Compensatory evolution in mitochondrial tRNAs navigates valleys of low fitness
Evolution from one fitness peak to another must involve either transitions through intermediates of low fitness or skirting round the fitness valley through compensatory mutations elsewhere. Here, the base pairs in mitochondrial tRNA stems is used as a model to show that deep fitness valleys can be traversed. Transitions between AU and GC pairs have occurred during mammalian evolution without help from genetic drift or mutations elsewhere.
- Margarita V. Meer
- , Alexey S. Kondrashov
- & Fyodor A. Kondrashov
-
Letter |
 Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome
Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome
Although new amino acids with desirable properties can be devised, only a few have been successfully introduced into proteins by the cellular machinery. Even then, only one type of unnatural amino acid can be added to a given protein. Here, a new system has been designed that could allow the incorporation of up to 200 novel amino acids. The system involves an orthogonal ribosome that uses quadruplet — rather than triplet — codons, as well as orthogonal tRNA synthetase–tRNA pairs.
- Heinz Neumann
- , Kaihang Wang
- & Jason W. Chin

 Adding α,α-disubstituted and β-linked monomers to the genetic code of an organism
Adding α,α-disubstituted and β-linked monomers to the genetic code of an organism