Featured
-
-
Article
| Open Access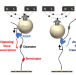 Reciprocating RNA Polymerase batters through roadblocks
Reciprocating RNA Polymerase batters through roadblocks
During transcription, RNA polymerases may encounter protein roadblocks along template DNA. Here, Qian et al. use magnetic tweezers to show that RNA polymerases can backtrack and ram into longer lived roadblocks to transit through them.
- Jin Qian
- , Allison Cartee
- & Laura Finzi
-
Article
| Open Access DNA binding analysis of rare variants in homeodomains reveals homeodomain specificity-determining residues
DNA binding analysis of rare variants in homeodomains reveals homeodomain specificity-determining residues
Analysis of 92 human homeodomain mutants, including disease-associated variants and variants of uncertain significance, reveals variants with altered DNA binding affinity and/or specificity and specificity-determining positions.
- Kian Hong Kock
- , Patrick K. Kimes
- & Martha L. Bulyk
-
Article
| Open Access Concerted transformation of a hyper-paused transcription complex and its reinforcing protein
Concerted transformation of a hyper-paused transcription complex and its reinforcing protein
Here, authors use cryoEM, biochemistry and molecular dynamics simulations to delineate a functional cycle of RfaH, a universally conserved transcription factor that undergoes a fold-switch during recruitment to the transcribing RNA polymerase.
- Philipp K. Zuber
- , Nelly Said
- & Stefan H. Knauer
-
Article
| Open Access Cis-regulatory interfaces reveal the molecular mechanisms underlying the notochord gene regulatory network of Ciona
Cis-regulatory interfaces reveal the molecular mechanisms underlying the notochord gene regulatory network of Ciona
The notochord is an essential hallmark of the chordate phylum. Here, Negrón-Piñeiro et al. study the notochord gene regulatory network in Ciona, and their findings illustrate how notochord transcription factors are coordinated by Brachyury and Foxa2.
- Lenny J. Negrón-Piñeiro
- , Yushi Wu
- & Anna Di Gregorio
-
Article
| Open Access Transcription-driven DNA supercoiling counteracts H-NS-mediated gene silencing in bacterial chromatin
Transcription-driven DNA supercoiling counteracts H-NS-mediated gene silencing in bacterial chromatin
Proteins compacting the bacterial chromosome obstruct transcription and must be transiently displaced to allow gene expression. Here, the authors show that the bacterial nucleoid structuring protein H-NS can be dislodged, from a distance, by the twisting in the DNA generated ahead of approaching RNA polymerase.
- Nara Figueroa-Bossi
- , Rocío Fernández-Fernández
- & Lionello Bossi
-
Article
| Open Access Structure of the human Bre1 complex bound to the nucleosome
Structure of the human Bre1 complex bound to the nucleosome
The structure of the nucleosome-bound human Bre1 complex reveals that its two RING domains bind the acidic patch and nucleosomal DNA, directing the E2 enzyme and ubiquitin for H2BK120-specific ubiquitination. The binding mode suggests a possible regulatory mechanism through nucleosomal DNA flexibility.
- Shuhei Onishi
- , Kotone Uchiyama
- & Toru Sengoku
-
Article
| Open Access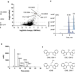 Diindoles produced from commensal microbiota metabolites function as endogenous CAR/Nr1i3 ligands
Diindoles produced from commensal microbiota metabolites function as endogenous CAR/Nr1i3 ligands
Here, combining metabolomic, proteomic and biophysical analyses, the authors identify and characterize a series of diindole molecules produced from commensal bacteria metabolites that act as specific agonists for the orphan constitutive androstane receptor (CAR), having potential to modulate gut and liver inflammation, metabolic diseases and cancer.
- Jiabao Liu
- , Ainaz Malekoltojari
- & Henry M. Krause
-
Article
| Open Access GAS41 modulates ferroptosis by anchoring NRF2 on chromatin
GAS41 modulates ferroptosis by anchoring NRF2 on chromatin
GAS41 is recognized as a histone reader and oncogene, but the mechanism by which GAS41 contributes to tumorigenesis is not well understood. Here, the authors discover that GAS41 is a ferroptosis repressor that anchors NRF2 to chromatin, promoting tumor growth.
- Zhe Wang
- , Xin Yang
- & Wei Gu
-
Article
| Open Access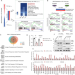 Reciprocal inhibition between TP63 and STAT1 regulates anti-tumor immune response through interferon-γ signaling in squamous cancer
Reciprocal inhibition between TP63 and STAT1 regulates anti-tumor immune response through interferon-γ signaling in squamous cancer
TP63 is a master regulator transcription factor in squamous cell carcinomas (SCCs). Here the authors report that TP63 suppresses IFNγ signaling in SCC tumors and that its inhibition is associated with enhanced anti-tumor immunity and response to anti-PD1.
- Yuan Jiang
- , Yueyuan Zheng
- & Yan-Yi Jiang
-
Article
| Open Access Oncogenic enhancers prime quiescent metastatic cells to escape NK immune surveillance by eliciting transcriptional memory
Oncogenic enhancers prime quiescent metastatic cells to escape NK immune surveillance by eliciting transcriptional memory
Metastasis arises from disseminated tumour cells (DTCs) while the underlying mechanism of DTCs plasticity remains underexplored. Here, the authors show that spatially organized oncogenic enhancers on chromatin sustain the establishment of retinoic acid (RA)-stimulated transcriptional memory through activation of SOX9, supporting the escape of quiescent DTCs from NK-mediated immune surveillance.
- Daniela Michelatti
- , Sven Beyes
- & Alessio Zippo
-
Article
| Open Access Fibroblast-specific PRMT5 deficiency suppresses cardiac fibrosis and left ventricular dysfunction in male mice
Fibroblast-specific PRMT5 deficiency suppresses cardiac fibrosis and left ventricular dysfunction in male mice
Epigenetic mechanisms play a key role in cardiac fibrosis associated with heart failure. Here, the authors show that protein arginine methyltransferase 5 (PRMT5), an epigenetic writer, regulates fibrotic gene transcription through histone methylation in mice.
- Yasufumi Katanasaka
- , Harumi Yabe
- & Tatsuya Morimoto
-
Article
| Open Access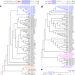 Evolutionary origin of Hoxc13-dependent skin appendages in amphibians
Evolutionary origin of Hoxc13-dependent skin appendages in amphibians
Hair is the main skin appendage of mammals. Here, the authors show that claws of clawed frogs and hair contain homologous keratins and depend on the same transcription factor, Hoxc13, suggesting a common evolutionary origin of these skin appendages.
- Marjolein Carron
- , Attila Placido Sachslehner
- & Leopold Eckhart
-
Article
| Open Access The Deinococcus protease PprI senses DNA damage by directly interacting with single-stranded DNA
The Deinococcus protease PprI senses DNA damage by directly interacting with single-stranded DNA
Lu et al. show that single-stranded DNA produced as a result of DNA damage may directly activate PprI in Deinococcus species, triggering the DNA damage response.
- Huizhi Lu
- , Zijing Chen
- & Yuejin Hua
-
Article
| Open Access Single-molecule RNA sizing enables quantitative analysis of alternative transcription termination
Single-molecule RNA sizing enables quantitative analysis of alternative transcription termination
The development of RNA technologies demands accurate assessment of transcript size and heterogeneity. Here, authors report a nanopore-based approach to study full-length RNA transcripts at the single-molecule level, identify premature transcription termination and study rolling-circle transcription.
- Gerardo Patiño-Guillén
- , Jovan Pešović
- & Ulrich Felix Keyser
-
Article
| Open Access Bacteria employ lysine acetylation of transcriptional regulators to adapt gene expression to cellular metabolism
Bacteria employ lysine acetylation of transcriptional regulators to adapt gene expression to cellular metabolism
The mechanisms underlying adaptation of bacteria to changing environmental conditions remain poorly understood. Here, the authors show bacteria using lysine acetylation of transcriptional regulators to adjust gene expression to changing conditions.
- Magdalena Kremer
- , Sabrina Schulze
- & Michael Lammers
-
Article
| Open Access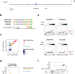 Cbp1 and Cren7 form chromatin-like structures that ensure efficient transcription of long CRISPR arrays
Cbp1 and Cren7 form chromatin-like structures that ensure efficient transcription of long CRISPR arrays
CRISPR arrays form the physical memory of prokaryotic adaptive immune systems by incorporating viral DNA sequences as spacers. Here, Blombach et al. show that transcription factor Cbp1 recruits chromatin protein Cren7 at CRISPR arrays, forming ‘chimeric’ chromatin-like structures that regulate expression of long CRISPR arrays in Sulfolobales archaea.
- Fabian Blombach
- , Michal Sýkora
- & Finn Werner
-
Article
| Open Access Structure of the recombinant RNA polymerase from African Swine Fever Virus
Structure of the recombinant RNA polymerase from African Swine Fever Virus
Pilotto and colleagues produce recombinant and catalytically active RNA polymerase (RNAP) from African Swine Fever Virus. Cryo-EM structures of RNAP with closed and open clamp conformations are presented along with in vitro transcription assays, yielding distinct functional conclusions.
- Simona Pilotto
- , Michal Sýkora
- & Finn Werner
-
Article
| Open Access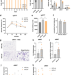 Adipocyte p53 coordinates the response to intermittent fasting by regulating adipose tissue immune cell landscape
Adipocyte p53 coordinates the response to intermittent fasting by regulating adipose tissue immune cell landscape
Adipose tissue (AT) inflammation is strongly associated with obesity and constitutes an obesogenic memory upon weight loss. Here, the authors show that intermittent fasting leads to an adipocyte p53-signaling dependent emergence of lipid-associated macrophages in visceral AT of obese mice which limits the systemic fasting response.
- Isabel Reinisch
- , Helene Michenthaler
- & Andreas Prokesch
-
Article
| Open Access The transcriptional regulatory network modulating human trophoblast stem cells to extravillous trophoblast differentiation
The transcriptional regulatory network modulating human trophoblast stem cells to extravillous trophoblast differentiation
Extravillous trophoblasts are pivotal in placental invasion and artery remodeling. Here the authors report stage-specific transcriptional regulators and their sequential roles in guiding proper trophoblast differentiation.
- Mijeong Kim
- , Yu Jin Jang
- & Jonghwan Kim
-
Article
| Open Access Predicting DNA structure using a deep learning method
Predicting DNA structure using a deep learning method
In this work, the authors report a deep learning method, Deep DNAshape, to predict the influence of flanking regions on three-dimensional DNA structure and in structural readout mechanisms of protein-DNA binding.
- Jinsen Li
- , Tsu-Pei Chiu
- & Remo Rohs
-
Article
| Open Access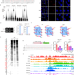 G-quadruplexes promote the motility in MAZ phase-separated condensates to activate CCND1 expression and contribute to hepatocarcinogenesis
G-quadruplexes promote the motility in MAZ phase-separated condensates to activate CCND1 expression and contribute to hepatocarcinogenesis
G-quadruplexes (G4s) can recruit transcription factors to activate genes. Here, the authors revealed that G4s drive molecular motility in phase-separated condensates of MAZ and coactivators, leading to activated CCND1 expression in liver cancer.
- Wenmeng Wang
- , Dangdang Li
- & Guangchao Sui
-
Article
| Open Access The host RNA polymerase II C-terminal domain is the anchor for replication of the influenza virus genome
The host RNA polymerase II C-terminal domain is the anchor for replication of the influenza virus genome
The cellular RNA polymerase II C-terminal domain is known to support transcription of influenza virus mRNAs. Here, Krischuns et al. use cell-based and in vitro approaches to demonstrate that it also plays a role in replication of the viral genome.
- Tim Krischuns
- , Benoît Arragain
- & Nadia Naffakh
-
Article
| Open Access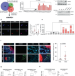 ANKRD1 is a mesenchymal-specific driver of cancer-associated fibroblast activation bridging androgen receptor loss to AP-1 activation
ANKRD1 is a mesenchymal-specific driver of cancer-associated fibroblast activation bridging androgen receptor loss to AP-1 activation
The transcriptional program controlling the activation of cancer-associated fibroblasts (CAFs) remains to be elucidated. Here, the authors identify ANKRD1 as a mesenchymal-specific driver of CAF activation under negative direct control of androgen receptor, triggering AP-1 transcription factor complex activation.
- Luigi Mazzeo
- , Soumitra Ghosh
- & G. Paolo Dotto
-
Article
| Open Access RNA polymerase II pausing is essential during spermatogenesis for appropriate gene expression and completion of meiosis
RNA polymerase II pausing is essential during spermatogenesis for appropriate gene expression and completion of meiosis
Gene expression dynamics are tightly regulated during spermatogenesis, with disruptions resulting in infertility. Here they identify a critical role for RNA PolII pausing in spermatogenesis and show that loss of the RNA PolII pausing factor NELF causes meiotic arrest.
- Emily G. Kaye
- , Kavyashree Basavaraju
- & Prabhakara P. Reddi
-
Article
| Open Access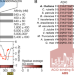 Molecular switching in transcription through splicing and proline-isomerization regulates stress responses in plants
Molecular switching in transcription through splicing and proline-isomerization regulates stress responses in plants
Transcription factor DREB2A interacts with Med25 to regulate stress responses. Here, the authors show that DREB2A uses splicing and proline-isomerization for this regulation and that proline cis-trans switching introduces structural frustration facilitating regulator exchange.
- Frederik Friis Theisen
- , Andreas Prestel
- & Karen Skriver
-
Article
| Open Access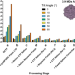 Overcoming resolution attenuation during tilted cryo-EM data collection
Overcoming resolution attenuation during tilted cryo-EM data collection
Here, the authors quantify the effect of cryo-EM data acquisition with stage-tilt on the global resolution of reconstructions and present a tool for predicting an optimal stage-tilt angle to ameliorate the effects of preferred specimen orientation.
- Sriram Aiyer
- , Philip R. Baldwin
- & Dmitry Lyumkis
-
Article
| Open Access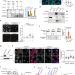 PSIP1/LEDGF reduces R-loops at transcription sites to maintain genome integrity
PSIP1/LEDGF reduces R-loops at transcription sites to maintain genome integrity
R-loop accumulation at transcription sites poses a persistent threat to genome integrity. Here the authors demonstrate a role for PSIP1/LEDGF protein in reducing R-loop levels at the site of transcription and preventing transcription replication conflict to maintain genome integrity.
- Sundarraj Jayakumar
- , Manthan Patel
- & Madapura M. Pradeepa
-
Article
| Open Access Tail-tape-fused virion and non-virion RNA polymerases of a thermophilic virus with an extremely long tail
Tail-tape-fused virion and non-virion RNA polymerases of a thermophilic virus with an extremely long tail
The authors describe the structure and function of two evolutionarily diverged RNA polymerases of a thermophilic phage. One of the polymerases is fused to the phage tape measure protein, a virion component dictating the length of the phage tail.
- Anastasiia Chaban
- , Leonid Minakhin
- & Maria L. Sokolova
-
Article
| Open Access The virulence regulator VirB from Shigella flexneri uses a CTP-dependent switch mechanism to activate gene expression
The virulence regulator VirB from Shigella flexneri uses a CTP-dependent switch mechanism to activate gene expression
Protein VirB regulates the expression of virulence genes in the pathogen Shigella flexneri by binding to DNA sequences far upstream of their promoters. Here, Jakob et al. show that VirB acts as a CTP-dependent molecular switch that uses a loading-and-sliding mechanism to control transcription of its target genes.
- Sara Jakob
- , Wieland Steinchen
- & Martin Thanbichler
-
Article
| Open Access Mutant GGGGCC RNA prevents YY1 from binding to Fuzzy promoter which stimulates Wnt/β-catenin pathway in C9ALS/FTD
Mutant GGGGCC RNA prevents YY1 from binding to Fuzzy promoter which stimulates Wnt/β-catenin pathway in C9ALS/FTD
Intronic GGGGCC repeat expansion in the C9orf72 gene causes ALS/FTD. Here the authors show that mutant GGGGCC RNA triggers YY1-Fuzzy transcriptional dysregulation which subsequently induces Wnt/β-catenin pathway and activates cell death in C9ALS/FTD.
- Zhefan Stephen Chen
- , Mingxi Ou
- & Ho Yin Edwin Chan
-
Article
| Open Access PAX3-FOXO1 uses its activation domain to recruit CBP/P300 and shape RNA Pol2 cluster distribution
PAX3-FOXO1 uses its activation domain to recruit CBP/P300 and shape RNA Pol2 cluster distribution
Different processes are in place to facilitate RNA-Polymerase 2 (Pol2) binding to chromatin. Here the authors reveal that the fusion transcription factor PAX3-FOXO1 shapes RNA Pol2 enhancer loops by recruitment of the histone acetyltransferase p300 via a small alpha-helical hook in its activation domain. Degradation of PAX3-FOXO1 or p300 rapidly collapses Pol2 clusters.
- Yaw Asante
- , Katharina Benischke
- & Marco Wachtel
-
Article
| Open Access MYOD-SKP2 axis boosts tumorigenesis in fusion negative rhabdomyosarcoma by preventing differentiation through p57Kip2 targeting
MYOD-SKP2 axis boosts tumorigenesis in fusion negative rhabdomyosarcoma by preventing differentiation through p57Kip2 targeting
SKP2 is an oncogenic E3-ubiquitin ligase. Here the authors show that SKP2 is epigenetically regulated by the muscle lineage transcription factor MYOD, supports tumorigenesis in the Fusion Negative (FN) subtype of rhabdomyosarcoma (RMS) and impairs differentiation promoting degradation of p57Kip2.
- Silvia Pomella
- , Matteo Cassandri
- & Rossella Rota
-
Article
| Open Access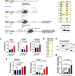 ERK2-topoisomerase II regulatory axis is important for gene activation in immediate early genes
ERK2-topoisomerase II regulatory axis is important for gene activation in immediate early genes
Topoisomerase II (TOP2) has been identified as one of the regulators for the transcriptional activation of immediate early genes IEGs. Here the authors report that ERK1 and ERK2 oppositely regulate transcription and ERK2 regulates TOP2B-DNA interaction and catalysis to favor IEG transcription.
- Heeyoun Bunch
- , Deukyeong Kim
- & Shun-ichi Sekine
-
Article
| Open Access Structure of the complete Saccharomyces cerevisiae Rpd3S-nucleosome complex
Structure of the complete Saccharomyces cerevisiae Rpd3S-nucleosome complex
In this study, the authors present the cryogenic electron microscopy reconstruction of the Rpd3S complex engaged with a nucleosome. The corresponding model describes the interactions that facilitate histone deacetylation within gene bodies by the Rpd3S complex.
- Jonathan W. Markert
- , Seychelle M. Vos
- & Lucas Farnung
-
Article
| Open Access Quorum-sensing synthase mutations re-calibrate autoinducer concentrations in clinical isolates of Pseudomonas aeruginosa to enhance pathogenesis
Quorum-sensing synthase mutations re-calibrate autoinducer concentrations in clinical isolates of Pseudomonas aeruginosa to enhance pathogenesis
Simanek et al. discovered variants that arise in the protein responsible for synthesizing a molecule required for bacterial communication, which mediates the progression of virulence in the pathogen Pseudomonas aeruginosa.
- Kayla A. Simanek
- , Megan L. Schumacher
- & Jon E. Paczkowski
-
Article
| Open Access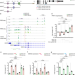 The SPOC proteins DIDO3 and PHF3 co-regulate gene expression and neuronal differentiation
The SPOC proteins DIDO3 and PHF3 co-regulate gene expression and neuronal differentiation
Death-inducer obliterator 3 (DIDO3) and PHD finger protein 3 (PHF3) are paralogue proteins that regulate transcription elongation by docking onto phosphorylated serine-2 in the C-terminal domain (CTD) of Pol II through their SPOC domains. Here the authors characterize the interplay of these proteins and show that they coregulate neuronal target genes.
- Johannes Benedum
- , Vedran Franke
- & Dea Slade
-
Article
| Open Access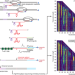 Observation of coordinated RNA folding events by systematic cotranscriptional RNA structure probing
Observation of coordinated RNA folding events by systematic cotranscriptional RNA structure probing
RNA begins to fold as the nascent transcript emerges from a transcribing RNA polymerase. Here, the authors develop a concise method for mapping RNA folding pathways that couples in vitro transcription with high-throughput RNA chemical probing.
- Courtney E. Szyjka
- & Eric J. Strobel
-
Article
| Open Access ID1 expressing macrophages support cancer cell stemness and limit CD8+ T cell infiltration in colorectal cancer
ID1 expressing macrophages support cancer cell stemness and limit CD8+ T cell infiltration in colorectal cancer
Inhibitor of differentiation 1 (ID1) has been described as a cancer-promoting factor and also involved in the formation of an immunosuppressive tumor microenvironment. Here the authors report that ID1-expressing tumor associated macrophages favor colorectal cancer progression by promoting cancer cell stemness and CD8+ T cell exclusion.
- Shuang Shang
- , Chen Yang
- & Fang Hua
-
Article
| Open Access c-di-GMP inhibits the DNA binding activity of H-NS in Salmonella
c-di-GMP inhibits the DNA binding activity of H-NS in Salmonella
H-NS is a global regulatory protein that represses expression of many genes in bacteria. Here, Li et al. show that a second messenger, cyclic di-GMP, binds to H-NS and inhibits its binding to DNA, thus relieving H-NS-mediated transcriptional silencing.
- Shuyu Li
- , Qinmeng Liu
- & Lei Zhang
-
Article
| Open Access Synthetic genetic oscillators demonstrate the functional importance of phenotypic variation in pneumococcal-host interactions
Synthetic genetic oscillators demonstrate the functional importance of phenotypic variation in pneumococcal-host interactions
Here, Rueff et al engineered a CRISPRi-based oscillator to rewire capsule production in Streptococcus pneumoniae from its native control. They show that heterogeneity in capsule production is beneficial for fitness in several virulence associated traits.
- Anne-Stéphanie Rueff
- , Renske van Raaphorst
- & Jan-Willem Veening
-
Article
| Open Access The environmentally-regulated interplay between local three-dimensional chromatin organisation and transcription of proVWX in E. coli
The environmentally-regulated interplay between local three-dimensional chromatin organisation and transcription of proVWX in E. coli
Here, the authors use the proVWXoperon of Escherichia coli as a model system to show how the nucleoid associated protein H-NS regulates gene expression in vivo by local chromatin remodelling.
- Fatema-Zahra M. Rashid
- , Frédéric G. E. Crémazy
- & Remus T. Dame
-
Article
| Open Access Transposable element-initiated enhancer-like elements generate the subgenome-biased spike specificity of polyploid wheat
Transposable element-initiated enhancer-like elements generate the subgenome-biased spike specificity of polyploid wheat
The direct impacts of transposable element dynamics on polyploid regulation and developmental specificity remain unclear. Here, the authors show that a large proportion of enhancer-like elements (ELEs) are mainly originated from RLG_famc7.3 specifically expanded in subgenome A, producing active nascent transcripts and influencing wheat spike development.
- Yilin Xie
- , Songbei Ying
- & Yijing Zhang
-
Article
| Open Access Transcriptional responses of cancer cells to heat shock-inducing stimuli involve amplification of robust HSF1 binding
Transcriptional responses of cancer cells to heat shock-inducing stimuli involve amplification of robust HSF1 binding
The authors compare the heat shock response between different cell lines and stimuli and reveal the genome-wide binding of its master transcription factor HSF1 as a platform for context-specific transcription activation.
- Sayantani Ghosh Dastidar
- , Bony De Kumar
- & Sergei Nechaev
-
Article
| Open Access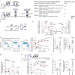 Themis2 regulates natural killer cell memory function and formation
Themis2 regulates natural killer cell memory function and formation
Innate immunity represents the first line of defence against pathogens, but certain innate cells are capable of memory formation, albeit with different and lesser-known mechanisms than adoptive immune cells. Here authors show that Themis2 regulates both memory NK cell development and function, via distinct downstream pathways.
- Tsukasa Nabekura
- , Elfira Amalia Deborah
- & Akira Shibuya
-
Article
| Open Access Transcriptional reprogramming by mutated IRF4 in lymphoma
Transcriptional reprogramming by mutated IRF4 in lymphoma
Cancer is often associated with mutant transcription factors (TFs) but their functional characterization is challenging. Here, the authors describe a recurrent mutation within TF IRF4 in human lymphomas and they show how it causes a complex switch in TF specificity and functionality.
- Nikolai Schleussner
- , Pierre Cauchy
- & Stephan Mathas
-
Article
| Open Access Paired yeast one-hybrid assays to detect DNA-binding cooperativity and antagonism across transcription factors
Paired yeast one-hybrid assays to detect DNA-binding cooperativity and antagonism across transcription factors
Combinations of transcription factors (TFs) bind DNA to fine-tune gene expression. Here, the authors map cooperative and antagonistic DNA binding across hundreds of TF-pairs. TF-TF relationships vary depending on DNA targets and TF isoforms.
- Anna Berenson
- , Ryan Lane
- & Juan I. Fuxman Bass
-
Article
| Open Access Structure of the transcription open complex of distinct σI factors
Structure of the transcription open complex of distinct σI factors
Here the authors show that σI factors encompass a unique, hitherto-unknown recognition mode of bacterial transcriptional promoters and represent a new distinctive class of σ70-family σ factors for bacterial transcription.
- Jie Li
- , Haonan Zhang
- & Ping Zhu
-
Article
| Open Access Transcriptional repression by a secondary DNA binding surface of DNA topoisomerase I safeguards against hypertranscription
Transcriptional repression by a secondary DNA binding surface of DNA topoisomerase I safeguards against hypertranscription
Aberrant hypertranscription in cells can affect development and disease. Here, the authors show that DNA topoisomerase I prevents hypertranscription; not through its catalytic function, but through a DNA binding mechanism.
- Mei Sheng Lau
- , Zhenhua Hu
- & Wee-Wei Tee
-
Article
| Open Access Acetylation reprograms MITF target selectivity and residence time
Acetylation reprograms MITF target selectivity and residence time
The microphthalmia-associated transcription factor MITF is a lineage-survival oncogene that plays a crucial role in melanocyte development and melanoma. Here, the authors reveal that MITF has a very long chromatin-bound half-life, and that MITF target selectivity is regulated by K206 acetylation, a residue linked to Waardenburg syndrome.
- Pakavarin Louphrasitthiphol
- , Alessia Loffreda
- & Colin R. Goding

 Sm-like protein Rof inhibits transcription termination factor ρ by binding site obstruction and conformational insulation
Sm-like protein Rof inhibits transcription termination factor ρ by binding site obstruction and conformational insulation