Featured
-
-
Article
| Open Access RNase III CLASH in MRSA uncovers sRNA regulatory networks coupling metabolism to toxin expression
RNase III CLASH in MRSA uncovers sRNA regulatory networks coupling metabolism to toxin expression
Regulatory small RNA (sRNA) interact with mRNAs to regulate their stability, transcription, and translation via diverse mechanisms. Here, McKellar et al. apply RNase IIICLASH of multi-drug resistant Staphylococcus aureus under different culture conditions to link the network of RNA-RNA interactions to environmental conditions and find that the production of small membrane-permeabilizing toxins is strongly regulated by sRNAs.
- Stuart W. McKellar
- , Ivayla Ivanova
- & Sander Granneman
-
Article
| Open Access A scalable, open-source implementation of a large-scale mechanistic model for single cell proliferation and death signaling
A scalable, open-source implementation of a large-scale mechanistic model for single cell proliferation and death signaling
Mechanistic models of how single cells respond to different perturbations can help integrate disparate big data sets or predict response to varied drug combinations. Here the authors develop a scalable, open-source pipeline for constructing and simulating large-scale, single-cell mechanistic models, an important building block for clinically-predictive mechanistic models and interpretable big data integration.
- Cemal Erdem
- , Arnab Mutsuddy
- & Marc R. Birtwistle
-
Article
| Open Access Global stable-isotope tracing metabolomics reveals system-wide metabolic alternations in aging Drosophila
Global stable-isotope tracing metabolomics reveals system-wide metabolic alternations in aging Drosophila
Stable-isotope tracing allows quantifying metabolic activity by measuring isotopically labeled metabolites, but its metabolome coverage has been limited. Here, the authors develop a global isotope tracing approach with metabolome-wide coverage and use it to characterize metabolic activities in aging Drosophila.
- Ruohong Wang
- , Yandong Yin
- & Zheng-Jiang Zhu
-
Comment
| Open Access Diagonal integration of multimodal single-cell data: potential pitfalls and paths forward
Diagonal integration of multimodal single-cell data: potential pitfalls and paths forward
Diagonal integration of multimodal single-cell data emerges as a trending topic. However, empowering diagonal methods for novel biological discoveries requires bridging huge gaps. Here, we comment on potential risks and future directions of diagonal integration for multimodal single-cell data.
- Yang Xu
- & Rachel Patton McCord
-
Article
| Open Access Enhancing bioreactor arrays for automated measurements and reactive control with ReacSight
Enhancing bioreactor arrays for automated measurements and reactive control with ReacSight
Small-scale bioreactors are increasingly used in quantitative biology. Here, the authors report ReacSight, a software solution to connect reactor arrays with sensitive measurement devices using low-cost pipetting robots and provide applications leveraging optogenetic control in yeast.
- François Bertaux
- , Sebastián Sosa-Carrillo
- & Gregory Batt
-
Article
| Open Access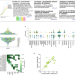 A genome-wide atlas of antibiotic susceptibility targets and pathways to tolerance
A genome-wide atlas of antibiotic susceptibility targets and pathways to tolerance
A lack of understanding in the development and emergence of antimicrobial resistance presents as a problem for accurate infection diagnosis and treatment. Here, authors utilize Streptococcus pneumoniae and build a genome-wide atlas to understand the genes and interactions that contribute to altered drug susceptibility.
- Dmitry Leshchiner
- , Federico Rosconi
- & Tim van Opijnen
-
Article
| Open Access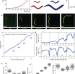 Coupling of growth rate and developmental tempo reduces body size heterogeneity in C. elegans
Coupling of growth rate and developmental tempo reduces body size heterogeneity in C. elegans
Animals must reach the correct size during development, despite stochastic differences in their growth rate. Here, Stojanovski et al. show that a coupling of growth and development by an oscillatory timer buffers fluctuations in the growth of the nematode C. elegans to ensure its correct size.
- Klement Stojanovski
- , Helge Großhans
- & Benjamin D. Towbin
-
Article
| Open Access High resolution microfluidic assay and probabilistic modeling reveal cooperation between T cells in tumor killing
High resolution microfluidic assay and probabilistic modeling reveal cooperation between T cells in tumor killing
Anti-cancer cytotoxic T cell responses largely vary among individuals. Here authors show, by stochastic modeling on high throughput T cell behavior and matched tumor spheroid fate data generated by a microfluidics system, that tumor killing is dependent on T cell cooperativity, which might contribute to the heterogeneity of T cell responses.
- Gustave Ronteix
- , Shreyansh Jain
- & Charles N. Baroud
-
Article
| Open Access Artificial neural networks enable genome-scale simulations of intracellular signaling
Artificial neural networks enable genome-scale simulations of intracellular signaling
Many diseases are caused by disruptions to the network of biochemical reactions that allow cells to respond to external signals. Here Nilsson et al develop a method to simulate cellular signaling using artificial neural networks to predict cellular responses and activities of signaling molecules.
- Avlant Nilsson
- , Joshua M. Peters
- & Douglas A. Lauffenburger
-
Article
| Open Access Machine learning aided construction of the quorum sensing communication network for human gut microbiota
Machine learning aided construction of the quorum sensing communication network for human gut microbiota
Microbes communicate with each other by Quorum sensing (QS) languages. Here the authors construct a QS database and the QS communication network to decipher intricate QSbased communications and form one of the key knowledge maps for human gut microbiota.
- Shengbo Wu
- , Jie Feng
- & Jianjun Qiao
-
Article
| Open Access De novo biosynthesis of rubusoside and rebaudiosides in engineered yeasts
De novo biosynthesis of rubusoside and rebaudiosides in engineered yeasts
Rubusoside and rebaudiosides are considered the next generation of sugar substitutes. In this article, the authors report the engineering of Saccharomyces cerevisiae, remodelling the complex metabolic networks by a modular engineering approach, obtaining rubusoside and rebaudiosides at titers of around 1.4 g/L and 100 mg/L, respectively.
- Yameng Xu
- , Xinglong Wang
- & Long Liu
-
Article
| Open Access Characterisation of a nucleo-adhesome
Characterisation of a nucleo-adhesome
Cell adhesion proteins have been described at sites away from the cell surface, including in the nucleus. Here, the authors report the scale of nuclear localisation of adhesion proteins, establishing a nucleo-adhesome and showing that nuclear adhesion proteins can cooperate to control transcription.
- Adam Byron
- , Billie G. C. Griffith
- & Margaret C. Frame
-
Article
| Open Access Modular (de)construction of complex bacterial phenotypes by CRISPR/nCas9-assisted, multiplex cytidine base-editing
Modular (de)construction of complex bacterial phenotypes by CRISPR/nCas9-assisted, multiplex cytidine base-editing
Rapid engineering of bacterial genomes is a requisite for both fundamental and applied studies. Here the authors develop an enhanced, broad-host-range cytidine base editor that enables multiplexed and efficient genome editing of Gram-negative bacteria.
- Daniel C. Volke
- , Román A. Martino
- & Pablo I. Nikel
-
Article
| Open Access Improving recombinant protein production by yeast through genome-scale modeling using proteome constraints
Improving recombinant protein production by yeast through genome-scale modeling using proteome constraints
Due to the complexity of the protein secretory pathway, strategy suitable for the production of a certain recombination protein cannot be generalized. Here, the authors construct a proteome-constrained genome-scale protein secretory model for yeast and show its application in the production of different misfolded or recombinant proteins.
- Feiran Li
- , Yu Chen
- & Jens Nielsen
-
Article
| Open Access Rewritable two-dimensional DNA-based data storage with machine learning reconstruction
Rewritable two-dimensional DNA-based data storage with machine learning reconstruction
Current DNA-based data storage platforms encode information only in the nucleotide sequence. Here, the authors report a 2DDNA platform that can store data in both sequence context and backbone structure, and has improved image inpainting and enhancement via automatic discoloration detection and deep learning.
- Chao Pan
- , S. Kasra Tabatabaei
- & Olgica Milenkovic
-
Article
| Open Access Complex dynamics in a synchronized cell-free genetic clock
Complex dynamics in a synchronized cell-free genetic clock
In theory, driven biological oscillators can display complex dynamic behaviors, but these are experimentally difficult to observe. Here the authors, using microfluidics, show that a synthetic cell-free gene oscillator displays period doubling and even quadrupling.
- Lukas Aufinger
- , Johann Brenner
- & Friedrich C. Simmel
-
Article
| Open Access Proteome allocations change linearly with the specific growth rate of Saccharomyces cerevisiae under glucose limitation
Proteome allocations change linearly with the specific growth rate of Saccharomyces cerevisiae under glucose limitation
Understanding how yeast organizes its functional proteome is a fundamental task in systems biology. Here, the authors conduct a multiomics analysis on yeast cells cultured with different growth rates, identifying a linear dependence of the functional proteome on the growth rate.
- Jianye Xia
- , Benjamin J. Sánchez
- & Jens Nielsen
-
Article
| Open Access Frequency modulation of a bacterial quorum sensing response
Frequency modulation of a bacterial quorum sensing response
Quorum-sensing bacteria produce and secrete autoinducers that trigger a behavioral change in the population when reaching a certain threshold. Here, Bettenworth et al. show that autoinducer synthase gene expression in Sinorhizobium meliloti occurs in asynchronous stochastic pulses, and that physiological cues modulate pulse frequency and, consequently, response behavior dynamics. Frequency-modulated pulsing in autoinducer synthase gene expression thus represents a time-based mechanism for information integration and collective decision-making.
- Vera Bettenworth
- , Simon van Vliet
- & Anke Becker
-
Article
| Open Access Broad-spectrum CRISPR-mediated inhibition of SARS-CoV-2 variants and endemic coronaviruses in vitro
Broad-spectrum CRISPR-mediated inhibition of SARS-CoV-2 variants and endemic coronaviruses in vitro
A major challenge in coronavirus vaccination and treatment is to counteract rapid viral evolution and mutations. Here the authors show that CRISPR-Cas13d can be used as a broad-spectrum antiviral to inhibit human coronaviruses, including new SARS-CoV-2 variants, combined with small molecule drugs for an enhanced antiviral effect in human primary cells.
- Leiping Zeng
- , Yanxia Liu
- & Lei S. Qi
-
Article
| Open Access Global diversity dynamics in the fossil record are regionally heterogeneous
Global diversity dynamics in the fossil record are regionally heterogeneous
Global diversity trends in the fossil record vary regionally through time and space, affecting our ability to interpret macroevolutionary history. Here, the authors propose a method to eliminate spatial sampling bias, estimate origination and extinction rates, and generate diversity estimates, applying this method to the Late Permian to Early Jurassic marine fossil record.
- Joseph T. Flannery-Sutherland
- , Daniele Silvestro
- & Michael J. Benton
-
Article
| Open Access Visual barcodes for clonal-multiplexing of live microscopy-based assays
Visual barcodes for clonal-multiplexing of live microscopy-based assays
Multiplex analyses of samples allow understanding complex processes in cancer initiation, progression and therapy response. Here, the authors present a fluorescence imaging-based visual barcode for livecell clonal-multiplexing which allows identifying signalling pathways clusters in response to different chemotherapy compounds.
- Tom Kaufman
- , Erez Nitzan
- & Ravid Straussman
-
Article
| Open Access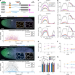 Differential regulation of alternative promoters emerges from unified kinetics of enhancer-promoter interaction
Differential regulation of alternative promoters emerges from unified kinetics of enhancer-promoter interaction
Alternative promoters differ in their expression patterns, whose mechanisms are not well understood. Here the authors show that alternative promoters of a Drosophila embryonic gene hunchback are regulated by different action modes of two enhancers.
- Jingyao Wang
- , Shihe Zhang
- & Heng Xu
-
Article
| Open Access An Artificial Intelligence-guided signature reveals the shared host immune response in MIS-C and Kawasaki disease
An Artificial Intelligence-guided signature reveals the shared host immune response in MIS-C and Kawasaki disease
Multisystem inflammatory syndrome may occur in children following COVID-19 infection. Here, the authors analyze gene signatures to show that MIS-C shares the same host immune response as the pre-pandemic inflammatory syndrome of Kawasaki disease but is further along in the spectrum in disease severity
- Pradipta Ghosh
- , Gajanan D. Katkar
- & Debashis Sahoo
-
Article
| Open Access PlasmidMaker is a versatile, automated, and high throughput end-to-end platform for plasmid construction
PlasmidMaker is a versatile, automated, and high throughput end-to-end platform for plasmid construction
Despite their broad utility, design and construction of plasmids remains laborious and time-consuming. Here the authors report a robust, versatile, and automated end-to-end platform that enables scarless construction of virtually any plasmid.
- Behnam Enghiad
- , Pu Xue
- & Huimin Zhao
-
Article
| Open Access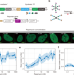 Modulating gene regulation function by chemically controlled transcription factor clustering
Modulating gene regulation function by chemically controlled transcription factor clustering
Transcription factor (TF) condensates appear to be pervasive, yet their roles remain debated. Here, the authors use a synthetic biology approach to show that TF clusters causally amplify transcription and can confer bimodality and “memory”.
- Jiegen Wu
- , Baoqiang Chen
- & Yihan Lin
-
Article
| Open Access In vitro reconstitution of Escherichia coli divisome activation
In vitro reconstitution of Escherichia coli divisome activation
In E. coli, FtsA and FtsZ control the place and time of cell division. Here, the authors use in vitro experiments to show how FtsA can follow FtsZ treadmilling and that downstream proteins form dynamic copolymers with FtsA to initiate division.
- Philipp Radler
- , Natalia Baranova
- & Martin Loose
-
Article
| Open Access Signal transduction in light-oxygen-voltage receptors lacking the active-site glutamine
Signal transduction in light-oxygen-voltage receptors lacking the active-site glutamine
Light-oxygen-voltage (LOV) photoreceptors perceive blue light to elicit spatio-temporally defined cellular responses, and their signalling process has been extensively characterized. Here the authors report that the light signal is still transduced in the absence of a conserved Gln residue, thought to be key.
- Julia Dietler
- , Renate Gelfert
- & Andreas Möglich
-
Article
| Open Access Context-specific effects of sequence elements on subcellular localization of linear and circular RNAs
Context-specific effects of sequence elements on subcellular localization of linear and circular RNAs
Ron and Ulitsky found using massively parallel assays that the effects of short RNA sequences on the subcellular localization of their host RNAs are strongly dependent on the host RNA form, linear or circular, and spliced or unspliced.
- Maya Ron
- & Igor Ulitsky
-
Article
| Open Access Enhancer RNAs stimulate Pol II pause release by harnessing multivalent interactions to NELF
Enhancer RNAs stimulate Pol II pause release by harnessing multivalent interactions to NELF
Enhancer RNAs (eRNAs) can stimulate gene transcription through various mechanisms. Here, the authors identify the molecular features within eRNAs that are critical for their action in facilitating RNA Polymerase II release from the paused state.
- Vladyslava Gorbovytska
- , Seung-Kyoon Kim
- & Claus-D. Kuhn
-
Article
| Open Access Elucidating Human Milk Oligosaccharide biosynthetic genes through network-based multi-omics integration
Elucidating Human Milk Oligosaccharide biosynthetic genes through network-based multi-omics integration
Human milk oligosaccharides are fundamental to infant health. Here the authors deploy a multi-omics systems biology approach to elucidate their biosynthetic network, including the associated enzymes and likely structures of ambiguous oligosaccharides.
- Benjamin P. Kellman
- , Anne Richelle
- & Nathan E. Lewis
-
Article
| Open Access A systems genomics approach to uncover patient-specific pathogenic pathways and proteins in ulcerative colitis
A systems genomics approach to uncover patient-specific pathogenic pathways and proteins in ulcerative colitis
Single Nucleotide Polymorphisms (SNPs) affect cellular regulatory networks, and SNP co-occurrences contribute to disease pathogenesis in ulcerative colitis (UC). Here the authors introduce iSNP, a precision medicine pipeline that combines genomics and network biology approaches to uncover patient specific pathways affected in complex diseases.
- Johanne Brooks-Warburton
- , Dezso Modos
- & Tamas Korcsmaros
-
Article
| Open Access Engineering artificial photosynthetic life-forms through endosymbiosis
Engineering artificial photosynthetic life-forms through endosymbiosis
The endosymbiotic theory posits that chloroplasts in eukaryotes arise from bacterial endosymbionts. Here, the authors engineer the yeast/cyanobacteria chimeras and show that the engineered cyanobacteria perform chloroplast-like functions to support the growth of yeast cells under photosynthetic conditions.
- Jason E. Cournoyer
- , Sarah D. Altman
- & Angad P. Mehta
-
Article
| Open Access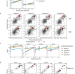 Machine learning-coupled combinatorial mutagenesis enables resource-efficient engineering of CRISPR-Cas9 genome editor activities
Machine learning-coupled combinatorial mutagenesis enables resource-efficient engineering of CRISPR-Cas9 genome editor activities
Screening combinatorial mutants is too massive for wet-lab experiment alone. Here the authors present a machine learning-coupled combinatorial mutagenesis approach to vastly reduce experimental burden for engineering Cas9 genome editing enzymes.
- Dawn G. L. Thean
- , Hoi Yee Chu
- & Alan S. L. Wong
-
Article
| Open Access Enabling reactive microscopy with MicroMator
Enabling reactive microscopy with MicroMator
In microscopy, applications in which reactiveness is needed are multifarious. Here the authors report MicroMator, a Python software package for reactive experiments, which they use for applications requiring real-time tracking and light-targeting at the single-cell level.
- Zachary R. Fox
- , Steven Fletcher
- & Gregory Batt
-
Article
| Open Access Bipartite network models to design combination therapies in acute myeloid leukaemia
Bipartite network models to design combination therapies in acute myeloid leukaemia
Identifying effective drug combinations to treat cancer is a challenging task, either experimentally or computationally. Here, the authors develop a bipartite network modelling approach to propose drug combination strategies in acute myeloid leukaemia using patient and cell line drug screening data.
- Mohieddin Jafari
- , Mehdi Mirzaie
- & Jing Tang
-
Article
| Open Access Data-driven learning how oncogenic gene expression locally alters heterocellular networks
Data-driven learning how oncogenic gene expression locally alters heterocellular networks
While mechanistic models play increasing roles in immuno-oncology, hand network curation is current practice. Here the authors use a Bayesian data-driven approach to infer how expression of a secreted oncogene alters the cellular landscape within the tumor.
- David J. Klinke II
- , Audry Fernandez
- & Anika C. Pirkey
-
Article
| Open Access Using high-resolution contact networks to evaluate SARS-CoV-2 transmission and control in large-scale multi-day events
Using high-resolution contact networks to evaluate SARS-CoV-2 transmission and control in large-scale multi-day events
Here, the authors simulate COVID-19 outbreaks on an empirical contact network derived from digital contact data collected on cruise ships. They model impacts of different control measures and find that combinations of measures, particularly vaccination and rapid antigen testing, are important for mitigating outbreaks.
- Rachael Pung
- , Josh A. Firth
- & Adam J. Kucharski
-
Article
| Open Access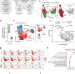 Single-cell characterization of leukemic and non-leukemic immune repertoires in CD8+ T-cell large granular lymphocytic leukemia
Single-cell characterization of leukemic and non-leukemic immune repertoires in CD8+ T-cell large granular lymphocytic leukemia
T cell large granular lymphocytic leukemia (T-LGLL) is a lymphoproliferative disorder involving clonally expanded T cell clones and is not fully understood. Here the authors show that the rest of the immune repertoire is interconnected with the T-LGLL clonotype(s) and is more mature, cytotoxic and clonally restricted than in other cancers and autoimmune disorders.
- Jani Huuhtanen
- , Dipabarna Bhattacharya
- & Satu Mustjoki
-
Article
| Open Access A genetic toolkit and gene switches to limit Mycoplasma growth for biosafety applications
A genetic toolkit and gene switches to limit Mycoplasma growth for biosafety applications
Mycoplasmas are minimal cell model organisms but lack genetic tools. Here the authors provide a robust genetic toolkit for Mycoplasma demonstrating gene circuit engineering applications.
- Alicia Broto
- , Erika Gaspari
- & Mark Isalan
-
Article
| Open Access acorde unravels functionally interpretable networks of isoform co-usage from single cell data
acorde unravels functionally interpretable networks of isoform co-usage from single cell data
Alternative splicing (AS) is a highly-regulated post-transcriptional mechanism known to modulate isoform expression within genes and contribute to cell-type identity. Here, the authors present acorde, a pipeline that successfully leverages bulk long reads and single-cell data to confidently detect alternative isoform co-expression relationships.
- Angeles Arzalluz-Luque
- , Pedro Salguero
- & Ana Conesa
-
Review Article
| Open Access Current progress and open challenges for applying deep learning across the biosciences
Current progress and open challenges for applying deep learning across the biosciences
Deep learning has enabled advances in understanding biology. In this review, the authors outline advances, and limitations of deep learning in five broad areas and the future challenges for the biosciences.
- Nicolae Sapoval
- , Amirali Aghazadeh
- & Todd J. Treangen
-
Article
| Open Access Robust and tunable signal processing in mammalian cells via engineered covalent modification cycles
Robust and tunable signal processing in mammalian cells via engineered covalent modification cycles
Phosphorylation networks are frequently at the heart of complex cellular decision making. Here the authors engineer synthetic phosphorylation devices with feedback regulation in mammalian cells and demonstrate how to use these to achieve tunable and robust control of cell behaviours.
- Ross D. Jones
- , Yili Qian
- & Ron Weiss
-
Article
| Open Access Ultrasound-controllable engineered bacteria for cancer immunotherapy
Ultrasound-controllable engineered bacteria for cancer immunotherapy
Synthetic biology has enabled the design of strategies for bacteria-based cancer immunotherapy. Here the authors report the development of focused ultrasound-activatable therapeutic bacteria engineered to express anti-CTLA-4 and anti-PD-L1 nanobodies for cancer immunotherapy.
- Mohamad H. Abedi
- , Michael S. Yao
- & Mikhail G. Shapiro
-
Article
| Open Access Integrative network analysis of early-stage lung adenocarcinoma identifies aurora kinase inhibition as interceptor of invasion and progression
Integrative network analysis of early-stage lung adenocarcinoma identifies aurora kinase inhibition as interceptor of invasion and progression
The molecular factors that drive invasiveness and metastasis in lung adenocarcinoma (LUAD) are not completely understood. Here, the authors use an integrative network approach to identify a gene signature of invasiveness in LUAD, and reveal Aurora kinases as master regulators of invasion.
- Seungyeul Yoo
- , Abhilasha Sinha
- & Charles A. Powell
-
Article
| Open Access Expanding biochemical knowledge and illuminating metabolic dark matter with ATLASx
Expanding biochemical knowledge and illuminating metabolic dark matter with ATLASx
“Mapping the dark matter of metabolism remains an open challenge that can be addressed globally and systematically by existing computational solutions. Here the authors present ATLASx, a repository of known and predicted enzymatic reaction, connecting millions of compounds to help synthetic biologists and metabolic engineers to design and explore metabolic pathways.”
- Homa MohammadiPeyhani
- , Jasmin Hafner
- & Vassily Hatzimanikatis
-
Article
| Open Access Design of stable and self-regulated microbial consortia for chemical synthesis
Design of stable and self-regulated microbial consortia for chemical synthesis
Stability and tunability are two desirable properties of microbial consortia-based bioproduction. Here, the authors integrate a caffeate-responsive biosensor into two and three strains coculture system to achieve autonomous regulation of strain ratios for coniferol and silybin/isosiltbin production, respectively.
- Xianglai Li
- , Zhao Zhou
- & Qipeng Yuan
-
Article
| Open Access Machine learning discovery of missing links that mediate alternative branches to plant alkaloids
Machine learning discovery of missing links that mediate alternative branches to plant alkaloids
Producing plant secondary metabolites by microbes is limited by the known enzymatic reactions. Here, the authors apply machine learning to predict missing link enzymes of benzylisoquinoline alkaloid (BIA) biosynthesis in Papaver somniferum, and validate the specialized activities through heterologous production.
- Christopher J. Vavricka
- , Shunsuke Takahashi
- & Tomohisa Hasunuma
-
Article
| Open Access Fully-automated and ultra-fast cell-type identification using specific marker combinations from single-cell transcriptomic data
Fully-automated and ultra-fast cell-type identification using specific marker combinations from single-cell transcriptomic data
Cell types are typically identified in single cell transcriptomic data by manual annotation of cell clusters using established marker genes. Here the authors present a fully-automated computational platform that can quickly and accurately distinguish between cell types.
- Aleksandr Ianevski
- , Anil K. Giri
- & Tero Aittokallio
-
Article
| Open Access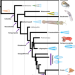 Fossil coleoid cephalopod from the Mississippian Bear Gulch Lagerstätte sheds light on early vampyropod evolution
Fossil coleoid cephalopod from the Mississippian Bear Gulch Lagerstätte sheds light on early vampyropod evolution
The authors describe a new cephalopod from the Carboniferous (Mississippian) Bear Gulch Lagerstätte of Montana, USA. This specimen extends the fossil record of vampyropods back by ~82 million years and changes our understanding of their evolution.
- Christopher D. Whalen
- & Neil H. Landman
Browse broader subjects
Browse narrower subjects
- Bayesian inference
- Biobricks
- Biochemical networks
- Bioenergetics
- Cellular noise
- Complexity
- Computer modelling
- Computer science
- Control theory
- Criticality
- Differential equations
- DNA computing and cryptography
- Dynamic networks
- Dynamical systems
- Emergence
- Evolvability
- Genetic circuit engineering
- Genetic interaction
- Genomic engineering
- Information theory
- Logic gates
- Metabolic engineering
- Modularity
- Molecular engineering
- Molecular fluctuations
- Multicellular systems
- Multistability
- Nonlinear dynamics
- Numerical simulations
- Oscillators
- Population dynamics
- Programming language
- Protein engineering
- Regulatory networks
- Reverse engineering
- Robustness
- Signal processing
- Single-cell imaging
- Software
- Standardization
- Stochastic modelling
- Stochastic networks
- Synthetic biology
- Systems analysis
- Time series

 Endothelial cell heterogeneity and microglia regulons revealed by a pig cell landscape at single-cell level
Endothelial cell heterogeneity and microglia regulons revealed by a pig cell landscape at single-cell level