Featured
-
-
Article
| Open Access Tankyrase inhibition preserves osteoarthritic cartilage by coordinating cartilage matrix anabolism via effects on SOX9 PARylation
Tankyrase inhibition preserves osteoarthritic cartilage by coordinating cartilage matrix anabolism via effects on SOX9 PARylation
Osteoarthritis results from the progressive destruction of cartilage matrix. Here, Kim et al. identify tankyrase as a regulator of cartilage matrix anabolism, and find that tankyrase inhibition, by preventing SOX9 PARylation, protects from cartilage destruction in a mouse model of osteoarthritis.
- Sukyeong Kim
- , Sangbin Han
- & Jin-Hong Kim
-
Article
| Open Access High-performance chemical- and light-inducible recombinases in mammalian cells and mice
High-performance chemical- and light-inducible recombinases in mammalian cells and mice
The availability of high performance recombinases with low basal activity and high dynamic range is limited. Here the authors present a library of over 20 orthogonal split recombinases that can be induced by small molecules, light and temperature in vivo.
- Benjamin H. Weinberg
- , Jang Hwan Cho
- & Wilson W. Wong
-
Article
| Open Access Oncolytic adenovirus programmed by synthetic gene circuit for cancer immunotherapy
Oncolytic adenovirus programmed by synthetic gene circuit for cancer immunotherapy
It is difficult to improve the efficacy of oncolytic virotherapy due to immune system responses and limited understanding of population dynamics. Here the authors use synthetic biology gene circuits to control adenoviral replication and release of immunomodulators in hepatocellular carcinoma cells.
- Huiya Huang
- , Yiqi Liu
- & Zhen Xie
-
Article
| Open Access A multiscale signalling network map of innate immune response in cancer reveals cell heterogeneity signatures
A multiscale signalling network map of innate immune response in cancer reveals cell heterogeneity signatures
The complexity of the innate immune response to cancer makes interpretation of large data sets challenging. Here, the authors provide an integrated multi-scale map of signalling networks representing the different immune cells and their interactions and show its utility for data interpretation.
- Maria Kondratova
- , Urszula Czerwinska
- & Inna Kuperstein
-
Article
| Open Access Transcriptional programming using engineered systems of transcription factors and genetic architectures
Transcriptional programming using engineered systems of transcription factors and genetic architectures
Successful approaches for controlling gene expression modulate mRNA synthesis by coupling it to inducible transcription effectors. Here the authors design 27 non-natural and non-synonymous transcription factors.
- Ronald E. Rondon
- , Thomas M. Groseclose
- & Corey J. Wilson
-
Article
| Open Access Single-cell transcriptomics of human T cells reveals tissue and activation signatures in health and disease
Single-cell transcriptomics of human T cells reveals tissue and activation signatures in health and disease
Immune cells are shaped by the tissue environment, yet the states of healthy human T cells are mainly studied in the blood. Here, the authors perform single cell RNA-seq of T cells from tissues and blood of healthy donors and show its utility as a reference map for comparison of human T cell states in disease.
- Peter A. Szabo
- , Hanna Mendes Levitin
- & Peter A. Sims
-
Article
| Open Access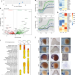 A genome-wide assessment of the ancestral neural crest gene regulatory network
A genome-wide assessment of the ancestral neural crest gene regulatory network
An understanding of the ancestral state of the neural crest (NC) gene regulatory network (GRN) gives insight into vertebrate evolution. Here, the authors use transcriptomic and chromatin accessibility analyses of the lamprey NC, as well as cross-species enhancer assays, to identify GRN elements conserved throughout vertebrates.
- Dorit Hockman
- , Vanessa Chong-Morrison
- & Tatjana Sauka-Spengler
-
Article
| Open Access Bacterial variability in the mammalian gut captured by a single-cell synthetic oscillator
Bacterial variability in the mammalian gut captured by a single-cell synthetic oscillator
Synthetic gene oscillators can be used to control timed function and periodic expression of genes. Here the authors demonstrate in vivo implementation in the mammalian gut that can keep time over several days.
- David T. Riglar
- , David L. Richmond
- & Pamela A. Silver
-
Article
| Open Access Sustained elevation of MG53 in the bloodstream increases tissue regenerative capacity without compromising metabolic function
Sustained elevation of MG53 in the bloodstream increases tissue regenerative capacity without compromising metabolic function
MG53 is a protein that regulates the cell membrane repair process, and it’s been suggested that it might play a role in diabetes. Here, the authors demonstrate that circulating MG53 functions as a myokine to facilitate tissue injury-repair and regeneration without impacting glucose handling.
- Zehua Bian
- , Qiang Wang
- & Jianjie Ma
-
Article
| Open Access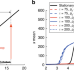 Transient hysteresis and inherent stochasticity in gene regulatory networks
Transient hysteresis and inherent stochasticity in gene regulatory networks
Cell fate commitment is understood in terms of bistable regulatory circuits with hysteresis, but inherent stochasticity in gene expression is incompatible with hysteresis. Here, the authors quantify how, under slow dynamics, the dependency of the non-stationary solutions on the initial state of the cells can lead to transient hysteresis.
- M. Pájaro
- , I. Otero-Muras
- & A. A. Alonso
-
Article
| Open Access Dissecting splicing decisions and cell-to-cell variability with designed sequence libraries
Dissecting splicing decisions and cell-to-cell variability with designed sequence libraries
Alternative splicing is regulated by multiple mechanisms. Here the authors employed designed splice site libraries and massively parallel reporter assays to dissect the regulatory complexity and cell-to-cell variability of splicing decisions and to build accurate predictive models.
- Martin Mikl
- , Amit Hamburg
- & Eran Segal
-
Article
| Open Access Mouse embryo geometry drives formation of robust signaling gradients through receptor localization
Mouse embryo geometry drives formation of robust signaling gradients through receptor localization
How receptor localization affects morphogen gradient formation during embryonic development is unclear. Here, the authors study the relationship between the BMP gradient, receptor localization, and compartmentalized geometry in the early mouse embryo, using experimental data and computational simulation.
- Zhechun Zhang
- , Steven Zwick
- & Sharad Ramanathan
-
Article
| Open Access Systematic identification of metabolites controlling gene expression in E. coli
Systematic identification of metabolites controlling gene expression in E. coli
Interactions between metabolites and transcription factors are known to control gene expression but analyzing these events at genome-scale is challenging. Here, the authors integrate dynamic metabolome and transcriptome data from E.coli to predict regulatory metabolite-transcription factor interactions.
- Martin Lempp
- , Niklas Farke
- & Hannes Link
-
Article
| Open Access Massive computational acceleration by using neural networks to emulate mechanism-based biological models
Massive computational acceleration by using neural networks to emulate mechanism-based biological models
Mechanistic models provide valuable insights, but large-scale simulations are computationally expensive. Here, the authors show that it is possible to explore the dynamics of a mechanistic model over a large set of parameters by training an artificial neural network on a smaller set of simulations.
- Shangying Wang
- , Kai Fan
- & Lingchong You
-
Article
| Open Access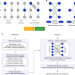 Discovering genetic interactions bridging pathways in genome-wide association studies
Discovering genetic interactions bridging pathways in genome-wide association studies
Genetic interactions may contribute to phenotypic traits but are challenging to decipher. Here, the authors develop BridGE, a computational approach for identifying pathways connected by genetic interactions from GWAS data.
- Gang Fang
- , Wen Wang
- & Chad L. Myers
-
Article
| Open Access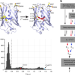 Learning the pattern of epistasis linking genotype and phenotype in a protein
Learning the pattern of epistasis linking genotype and phenotype in a protein
Epistasis underlies the complexity of genotype-phenotype maps. Here, the authors analyze 8,192 mutants that link two phenotypically distinct variants of the Entacmaea quadricolor fluorescent protein, and show the existence, but also the sparsity, of high-order epistatic interactions.
- Frank J. Poelwijk
- , Michael Socolich
- & Rama Ranganathan
-
Article
| Open Access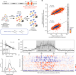 The mutational landscape of a prion-like domain
The mutational landscape of a prion-like domain
TDP43 aggregates are a hallmark of amyotrophic lateral sclerosis. By using deep mutagenesis to measure the toxicity of more than 50,000 mutations in the prion domain of TDP43, the authors conclude that mutations that increase toxicity promote formation of liquid-like condensates, while aggregation of TDP43 is protective for the cell.
- Benedetta Bolognesi
- , Andre J. Faure
- & Ben Lehner
-
Article
| Open Access Extensive disruption of protein interactions by genetic variants across the allele frequency spectrum in human populations
Extensive disruption of protein interactions by genetic variants across the allele frequency spectrum in human populations
Low frequency coding single-nucleotide variants (SNVs) are predicted to disproportionately affect protein function. Here, the authors evaluate 2,009 missense SNVs across 2,185 protein-protein interactions using yeast two-hybrid and protein complementation assays and find that disruptive SNVs often occur in disease-associated genes.
- Robert Fragoza
- , Jishnu Das
- & Haiyuan Yu
-
Article
| Open Access Bacterial co-culture with cell signaling translator and growth controller modules for autonomously regulated culture composition
Bacterial co-culture with cell signaling translator and growth controller modules for autonomously regulated culture composition
To avoid metabolic overload and divide tasks, synthetic biologists are turning to microbial consortia engineering. Here the authors design a co-culture controller that autonomously regulates population composition.
- Kristina Stephens
- , Maria Pozo
- & William E. Bentley
-
Article
| Open Access Gene networks that compensate for crosstalk with crosstalk
Gene networks that compensate for crosstalk with crosstalk
Crosstalk between genetic circuits is a major challenge for engineering sophisticated networks. Here the authors design networks that compensate for crosstalk by integrating, not insulating, pathways.
- Isaak E. Müller
- , Jacob R. Rubens
- & Timothy K. Lu
-
Article
| Open Access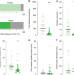 Rapid metabolic shifts occur during the transition between hunger and satiety in Drosophila melanogaster
Rapid metabolic shifts occur during the transition between hunger and satiety in Drosophila melanogaster
The relationship between metabolomic and behavioral changes is not well understood. Here, the authors analyze metabolome changes in D. melanogaster heads and bodies during hunger and satiety, and develop the Flyscape tool to visualize the resulting metabolic networks and integrate them with other omics data.
- Daniel Wilinski
- , Jasmine Winzeler
- & Monica Dus
-
Article
| Open Access Irrelevance of linear controllability to nonlinear dynamical networks
Irrelevance of linear controllability to nonlinear dynamical networks
Linear controllability theories have stimulated research on control of complex networks. Here the authors investigate the concordance between linear and nonlinear approaches in ranking the importance of nodes in nonlinear networks, and conclude that linear controllability may not be applicable.
- Junjie Jiang
- & Ying-Cheng Lai
-
Article
| Open Access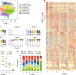 Heterogeneity of human bone marrow and blood natural killer cells defined by single-cell transcriptome
Heterogeneity of human bone marrow and blood natural killer cells defined by single-cell transcriptome
Natural killer (NK) cells are important innate immune cells with diverse functions. Here the authors use single-cell RNA-sequencing of purified human bone marrow and peripheral blood NK cells to define five populations of NK cells with distinct transcriptomic profile to further our understanding of NK development and heterogeneity.
- Chao Yang
- , Jason R. Siebert
- & Subramaniam Malarkannan
-
Article
| Open Access Changes in gene expression predictably shift and switch genetic interactions
Changes in gene expression predictably shift and switch genetic interactions
Non-additive genetic interactions are plastic and can complicate genetic prediction. Here, using deep mutagenesis of the lambda repressor, Li et al. reveal that changes in gene expression can alter the strength and direction of genetic interactions between mutations in many genes and develop mathematical models for predicting them.
- Xianghua Li
- , Jasna Lalić
- & Ben Lehner
-
Article
| Open Access Orthogonal monoterpenoid biosynthesis in yeast constructed on an isomeric substrate
Orthogonal monoterpenoid biosynthesis in yeast constructed on an isomeric substrate
Titers of monoterpenoids production in yeast are low due to the fact that the geranyl diphosphate (GPP)-based pathway can redirect metabolic fluxes to growth. Here, the authors build an orthogonal pathway by selecting the cis isomer of GPP as an alternative precursor and achieve high titer monoterpene production.
- Codruta Ignea
- , Morten H. Raadam
- & Sotirios C. Kampranis
-
Article
| Open Access Programmable biomolecular switches for rewiring flux in Escherichia coli
Programmable biomolecular switches for rewiring flux in Escherichia coli
Current flux rewiring technologies in metabolic engineering are mainly transcriptional regulation. Here, the authors build two sets of controllable protein units using engineered viral proteases and proteolytic signals, and utilize for increasing titers of shikimate and D-xylonate in E. coli.
- Cong Gao
- , Jianshen Hou
- & Liming Liu
-
Article
| Open Access Harmonious genetic combinations rewire regulatory networks and flip gene essentiality
Harmonious genetic combinations rewire regulatory networks and flip gene essentiality
Studying how genetic variants in different genes interact and their combinatorial output is experimentally and analytically challenging. Here, the authors quantify the effects of more than 5000 mutation pairs in the yeast GAL regulatory system, finding that many combinations can be predicted with statistical models.
- Aaron M. New
- & Ben Lehner
-
Article
| Open Access Information-based centralization of locomotion in animals and robots
Information-based centralization of locomotion in animals and robots
Model-based centralization schemes, though able to quantify locomotion control in animals and bio-inspired robots, are limited to specific systems. Here, the authors report a generalized information-based centralization scheme that unifies existing models and can be applied to different systems.
- Izaak D. Neveln
- , Amoolya Tirumalai
- & Simon Sponberg
-
Article
| Open Access The pause-initiation limit restricts transcription activation in human cells
The pause-initiation limit restricts transcription activation in human cells
Gene activation requires an increase of successful initiation events. Here, by employing a genome-wide kinetic analysis of transcription, the authors showed that gene activation generally requires a decrease in RNA Polymerase II (Pol II) promoter-proximal pausing while transcription of enhancer elements is not limited by Pol II pausing.
- Saskia Gressel
- , Björn Schwalb
- & Patrick Cramer
-
Article
| Open Access Integrated TORC1 and PKA signaling control the temporal activation of glucose-induced gene expression in yeast
Integrated TORC1 and PKA signaling control the temporal activation of glucose-induced gene expression in yeast
Yeast cells respond to nutrients by altering expression of the protein synthesis genes and thus their growth rate. Here, the authors use microarrays to show that TORC1 controls gene expression during steady state growth while PKA speeds up expression changes when nutrient levels change.
- Joseph Kunkel
- , Xiangxia Luo
- & Andrew P. Capaldi
-
Article
| Open Access Microbial carbon use efficiency predicted from genome-scale metabolic models
Microbial carbon use efficiency predicted from genome-scale metabolic models
Microbial respiration releases carbon from the soil. Here, the authors estimate bacterial carbon use efficiency in soils for over 200 species using constraint-based modeling, incorporate the values into an ecosystem model, and find that shifts in community composition may impact carbon storage.
- Mustafa Saifuddin
- , Jennifer M. Bhatnagar
- & Adrien C. Finzi
-
Article
| Open Access A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism
A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism
Genome-scale metabolic models provide a platform to study metabolism through simulations and analysis of omics data. Here the authors introduce Yeast8 with its model ecosystem, a comprehensive computational resource for simulating the metabolism of Saccharomyces cerevisiae.
- Hongzhong Lu
- , Feiran Li
- & Jens Nielsen
-
Article
| Open Access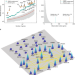 A genome-wide positioning systems network algorithm for in silico drug repurposing
A genome-wide positioning systems network algorithm for in silico drug repurposing
Identification of disease modules in the human interactome can guide more efficacious therapeutic selections. Here, the authors introduce a network-based methodology to identify individualized disease modules by mapping patients’ DNA and RNA sequencing profiles to the interactome, enabling prediction of cancer type-specific drug responses.
- Feixiong Cheng
- , Weiqiang Lu
- & Joseph Loscalzo
-
Article
| Open Access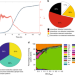 Regulatory mechanisms underlying coordination of amino acid and glucose catabolism in Escherichia coli
Regulatory mechanisms underlying coordination of amino acid and glucose catabolism in Escherichia coli
Bacteria must adapt their metabolism in the face of dynamically changing nutrient availability. Here, using their constraint-based modeling approach the authors analyze E. coli exometabolome data during growth in complex medium, revealing temporal coordination of glucose and amino acid catabolism.
- Mattia Zampieri
- , Manuel Hörl
- & Uwe Sauer
-
Perspective
| Open Access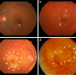 A systems biology approach towards understanding and treating non-neovascular age-related macular degeneration
A systems biology approach towards understanding and treating non-neovascular age-related macular degeneration
No effective therapies exist for dry age-related macular degeneration. In this perspective, the authors propose that research should emphasize system biology approaches that integrate various ‘omics’ data into mathematical models to establish pathogenic mechanisms on which to design novel treatments, and identify biomarkers that predict disease progression and therapeutic response.
- James T. Handa
- , Cathy Bowes Rickman
- & Lindsay A. Farrer
-
Article
| Open Access Estimating dispensable content in the human interactome
Estimating dispensable content in the human interactome
The fraction of protein-protein interactions (PPIs) that can be disrupted without fitness effect is unknown. Here, the authors model how disease-causing mutations and common mutations carried by healthy people perturb the interactome, and estimate that <20% of human PPIs are completely dispensable.
- Mohamed Ghadie
- & Yu Xia
-
Article
| Open Access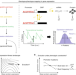 Empirical mean-noise fitness landscapes reveal the fitness impact of gene expression noise
Empirical mean-noise fitness landscapes reveal the fitness impact of gene expression noise
Quantifying the effects of noise in gene expression is difficult since noise and mean expression are coupled. Here the authors determine fitness landscapes in mean-noise expression space to uncouple these two parameters and show that changes in noise and mean expression are similarly detrimental to fitness.
- Jörn M. Schmiedel
- , Lucas B. Carey
- & Ben Lehner
-
Article
| Open Access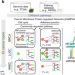 Membrane protein-regulated networks across human cancers
Membrane protein-regulated networks across human cancers
Membrane proteins have been implicated in cancers, but studying the downstream effects of their perturbation remains challenging. Here, the authors map the membrane protein-regulated network of 15 cancers, a resource for prognostic biomarker development and druggable target identification.
- Chun-Yu Lin
- , Chia-Hwa Lee
- & Jinn-Moon Yang
-
Article
| Open Access Environmental conditions shape the nature of a minimal bacterial genome
Environmental conditions shape the nature of a minimal bacterial genome
Minimal bacterial genomes still contain hundreds of genes of unknown function. Here the authors use in silico annotation methods and identify the environmental factors shaping a minimal genome.
- Magdalena Antczak
- , Martin Michaelis
- & Mark N. Wass
-
Article
| Open Access Optogenetic control of Bacillus subtilis gene expression
Optogenetic control of Bacillus subtilis gene expression
Bacillus subtilis has complex spatial and temporal gene expression patterns but currently lacks optogenetic tools to explore these processes. Here the authors import and debug a cyanobacterial green light sensor pathway and show that it enables precise optical control of gene expression.
- Sebastian M. Castillo-Hair
- , Elliot A. Baerman
- & Jeffrey J. Tabor
-
Article
| Open Access Pooled clone collections by multiplexed CRISPR-Cas12a-assisted gene tagging in yeast
Pooled clone collections by multiplexed CRISPR-Cas12a-assisted gene tagging in yeast
Construction of yeast libraries is time-consuming, costly and limited to the genetic background of the chosen strain. Here the authors present CASTLING which uses CRISPR-Cas12a and oligonucleotide pools to rapidly generate pooled libraries with large insertions such as fluorescent protein tags.
- Benjamin C. Buchmuller
- , Konrad Herbst
- & Michael Knop
-
Article
| Open Access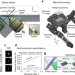 Multi-immersion open-top light-sheet microscope for high-throughput imaging of cleared tissues
Multi-immersion open-top light-sheet microscope for high-throughput imaging of cleared tissues
Light-sheet microscopes are increasingly used for imaging cleared tissues, but have imposed constraints on sample geometries and protocols. Here the authors present a multi-immersion open-top light-sheet microscope to overcome these limitations and enable high-throughput imaging of samples processed with various clearing protocols.
- Adam K. Glaser
- , Nicholas P. Reder
- & Jonathan T. C. Liu
-
Article
| Open Access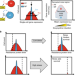 Role of network-mediated stochasticity in mammalian drug resistance
Role of network-mediated stochasticity in mammalian drug resistance
The role of gene expression noise in the evolution of drug resistance in mammalian cells is unclear. Here, by uncoupling noise from mean expression of a drug resistance gene in CHO cells the authors show that noisy expression aids adaptation to high drug levels, but delays it at low drug levels.
- Kevin S. Farquhar
- , Daniel A. Charlebois
- & Gábor Balázsi
-
Article
| Open Access Community assessment to advance computational prediction of cancer drug combinations in a pharmacogenomic screen
Community assessment to advance computational prediction of cancer drug combinations in a pharmacogenomic screen
Resistance to first line treatment is a major hurdle in cancer treatment, that can be overcome with drug combinations. Here, the authors provide a large drug combination screen across cancer cell lines to benchmark crowdsourced methods and to computationally predict drug synergies.
- Michael P. Menden
- , Dennis Wang
- & Julio Saez-Rodriguez
-
Article
| Open Access Comprehensive study of nuclear receptor DNA binding provides a revised framework for understanding receptor specificity
Comprehensive study of nuclear receptor DNA binding provides a revised framework for understanding receptor specificity
The type II nuclear receptors (NRs) and the retinoid X receptor (RXR) form heterodimeric transcription factors to regulate development, metabolism, and inflammation. Here the authors employ protein-binding microarrays to comprehensively analyze the DNA binding of 12 NR:RXRα heterodimers, and report promiscuous NR-DNA binding.
- Ashley Penvose
- , Jessica L. Keenan
- & Trevor Siggers
-
Article
| Open Access Survival of the simplest in microbial evolution
Survival of the simplest in microbial evolution
In asexual populations selection at different genomic loci can interfere with each other. Here, using a biophysical model of molecular evolution the authors show that interference results in long-term degradation of molecular function, an effect that strongly depends on genome size.
- Torsten Held
- , Daniel Klemmer
- & Michael Lässig
-
Article
| Open Access Feed-forward regulation adaptively evolves via dynamics rather than topology when there is intrinsic noise
Feed-forward regulation adaptively evolves via dynamics rather than topology when there is intrinsic noise
Feed‐forward loops (FFLs) can filter out noise, but whether their overrepresentation in GRNs reflects adaptive evolution for this function is debated. Here, the authors develop a null model of regulatory evolution and find that FFLs evolve readily under selection for the noise filtering function.
- Kun Xiong
- , Alex K. Lancaster
- & Joanna Masel
-
Article
| Open Access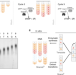 Terminator-free template-independent enzymatic DNA synthesis for digital information storage
Terminator-free template-independent enzymatic DNA synthesis for digital information storage
Adoption of DNA as a data storage medium could be accelerated with specialized synthesis processes and codecs. The authors describe TdT-mediated DNA synthesis in which data is stored in transitions between non-identical nucleotides and the use of synchronization markers to provide error tolerance.
- Henry H. Lee
- , Reza Kalhor
- & George M. Church
-
Article
| Open Access Theoretical analysis of Polycomb-Trithorax systems predicts that poised chromatin is bistable and not bivalent
Theoretical analysis of Polycomb-Trithorax systems predicts that poised chromatin is bistable and not bivalent
Polycomb and Trithorax group proteins regulate silent and active gene expression states, but also allow poised states in pluripotent cells. Here the authors present a mathematical model that integrates data on Polycomb/ Trithorax biochemistry into a single coherent framework which predicts that poised chromatin is not bivalent as previously proposed, but is bistable, meaning that the system switches frequently between stable active and silent states.
- Kim Sneppen
- & Leonie Ringrose
Browse broader subjects
Browse narrower subjects
- Bayesian inference
- Biobricks
- Biochemical networks
- Bioenergetics
- Cellular noise
- Complexity
- Computer modelling
- Computer science
- Control theory
- Criticality
- Differential equations
- DNA computing and cryptography
- Dynamic networks
- Dynamical systems
- Emergence
- Evolvability
- Genetic circuit engineering
- Genetic interaction
- Genomic engineering
- Information theory
- Logic gates
- Metabolic engineering
- Modularity
- Molecular engineering
- Molecular fluctuations
- Multicellular systems
- Multistability
- Nonlinear dynamics
- Numerical simulations
- Oscillators
- Population dynamics
- Programming language
- Protein engineering
- Regulatory networks
- Reverse engineering
- Robustness
- Signal processing
- Single-cell imaging
- Software
- Standardization
- Stochastic modelling
- Stochastic networks
- Synthetic biology
- Systems analysis
- Time series

 Biological process activity transformation of single cell gene expression for cross-species alignment
Biological process activity transformation of single cell gene expression for cross-species alignment