Article
|
Open Access
Featured
-
-
Article
| Open Access Enhancing CRISPR-Cas9 gRNA efficiency prediction by data integration and deep learning
Enhancing CRISPR-Cas9 gRNA efficiency prediction by data integration and deep learning
High-quality gRNA activity data is needed for accurate on-target efficiency predictions. Here the authors generate activity data for over 10,000 gRNA and build a deep learning model CRISPRon for improved performance predictions.
- Xi Xiang
- , Giulia I. Corsi
- & Yonglun Luo
-
Article
| Open Access Pareto optimality between growth-rate and lag-time couples metabolic noise to phenotypic heterogeneity in Escherichia coli
Pareto optimality between growth-rate and lag-time couples metabolic noise to phenotypic heterogeneity in Escherichia coli
It is unclear how noise in gene expression propagates to phenotypic heterogeneity in clonal bacterial populations. Here, the authors explore how variability in central sugar metabolism in E. coli can mediate and promote population diversification.
- Diego Antonio Fernandez Fuentes
- , Pablo Manfredi
- & Mattia Zampieri
-
Article
| Open Access The quantitative metabolome is shaped by abiotic constraints
The quantitative metabolome is shaped by abiotic constraints
Evolution selects for the fittest but must operate within the realm of the physically possible. Here, the authors present a theoretical framework that allows them to explore how ten abiotic constraints can shape the operation, regulation, and adaptation of metabolism in E. coli.
- Amir Akbari
- , James T. Yurkovich
- & Bernhard O. Palsson
-
Article
| Open Access Longitudinal analysis of blood markers reveals progressive loss of resilience and predicts human lifespan limit
Longitudinal analysis of blood markers reveals progressive loss of resilience and predicts human lifespan limit
Aging is associated with an increased risk of chronic diseases and functional decline. Here, the authors investigate the fluctuations of physiological indices along aging trajectories and observed a characteristic decrease in the organism state recovery rate.
- Timothy V. Pyrkov
- , Konstantin Avchaciov
- & Peter O. Fedichev
-
Article
| Open Access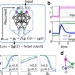 Finding gene network topologies for given biological function with recurrent neural network
Finding gene network topologies for given biological function with recurrent neural network
Networks are useful ways to describe interactions between molecules in a cell, but predicting the real topology of large networks can be challenging. Here, the authors use deep learning to predict the topology of networks that perform biologically-plausible functions.
- Jingxiang Shen
- , Feng Liu
- & Chao Tang
-
Article
| Open Access Synthetic neural-like computing in microbial consortia for pattern recognition
Synthetic neural-like computing in microbial consortia for pattern recognition
Complex biological systems have individual cells acting collectively to solve complex tasks. Here the authors implement neural network-like computing in a bacterial consortia to recognise patterns.
- Ximing Li
- , Luna Rizik
- & Ramez Daniel
-
Article
| Open Access Yeast cell fate control by temporal redundancy modulation of transcription factor paralogs
Yeast cell fate control by temporal redundancy modulation of transcription factor paralogs
How dynamic transcription factors temporally interact to regulate stress survival in yeast is currently unclear. Here the authors integrate single-cell imaging, RNA-seq, and modeling to identify a new cell fate control mechanism mediated by temporal redundancy modulation during yeast stress response.
- Yan Wu
- , Jiaqi Wu
- & Yihan Lin
-
Article
| Open Access The transcriptional landscape of a rewritten bacterial genome reveals control elements and genome design principles
The transcriptional landscape of a rewritten bacterial genome reveals control elements and genome design principles
Rewriting genomes allows for complete annotation of gene regulatory elements. Here the authors compare endogenous and rewritten segments of a genome and find extensive transcriptional changes, based on which they formulate design principles that aid in the programming of biological systems.
- Mariëlle J. F. M. van Kooten
- , Clio A. Scheidegger
- & Beat Christen
-
Article
| Open Access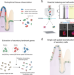 Clump sequencing exposes the spatial expression programs of intestinal secretory cells
Clump sequencing exposes the spatial expression programs of intestinal secretory cells
Combining scRNA-seq with spatial information to enable the reconstruction of spatially-resolved cell atlases is challenging for rare cell types. Here the authors present ClumpSeq, an approach for sequencing small clumps of tissue attached cells, and apply it to establish spatial atlases for all secretory cell types in the small intestine.
- Rita Manco
- , Inna Averbukh
- & Shalev Itzkovitz
-
Article
| Open Access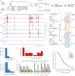 Tissue context determines the penetrance of regulatory DNA variation
Tissue context determines the penetrance of regulatory DNA variation
The functional consequences of variation in human regulatory DNA depend on the local chromatin environment and the cell/tissue context. Here the authors use highly diverged hybrid mice to study genetic effects on DNA accessibility in vivo across multiple cell and tissue types.
- Jessica M. Halow
- , Rachel Byron
- & Matthew T. Maurano
-
Article
| Open Access Programming gene expression in multicellular organisms for physiology modulation through engineered bacteria
Programming gene expression in multicellular organisms for physiology modulation through engineered bacteria
Manipulating animal physiology can be difficult because of the complexity of multicellular systems. Here the authors use engineered bacteria to modulate Caenorhabditis elegans gene expression through genetic circuit controlled RNAi.
- Baizhen Gao
- & Qing Sun
-
Article
| Open Access Automatic synchronisation of the cell cycle in budding yeast through closed-loop feedback control
Automatic synchronisation of the cell cycle in budding yeast through closed-loop feedback control
It is difficult to synchronize the cell cycle in a population of yeast cells for extended periods of time. Here the authors use a cybergenetic system with inbuilt feedback to synchronize a population of modified yeast.
- Giansimone Perrino
- , Sara Napolitano
- & Diego di Bernardo
-
Article
| Open Access Protein design and variant prediction using autoregressive generative models
Protein design and variant prediction using autoregressive generative models
The ability to design functional sequences is central to protein engineering and biotherapeutics. Here the authors introduce a deep generative alignment-free model for sequence design applied to highly variable regions and design and test a diverse nanobody library with improved properties for selection experiments.
- Jung-Eun Shin
- , Adam J. Riesselman
- & Debora S. Marks
-
Article
| Open Access The VRNetzer platform enables interactive network analysis in Virtual Reality
The VRNetzer platform enables interactive network analysis in Virtual Reality
Data-rich networks can be difficult to interpret beyond a certain size. Here, the authors introduce a platform that uses virtual reality to allow the visual exploration of large networks, while interfacing with data repositories and other analytical methods to improve the interpretation of big data.
- Sebastian Pirch
- , Felix Müller
- & Jörg Menche
-
Article
| Open Access Improving cell-free glycoprotein synthesis by characterizing and enriching native membrane vesicles
Improving cell-free glycoprotein synthesis by characterizing and enriching native membrane vesicles
Cell-free gene expression systems are an attractive platform for biomanufacturing and synthetic biology. Here the authors characterize native membrane vesicles in E. coli extracts for improved glycoengineering.
- Jasmine M. Hershewe
- , Katherine F. Warfel
- & Michael C. Jewett
-
Article
| Open Access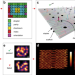 An alternative approach to nucleic acid memory
An alternative approach to nucleic acid memory
Encoding data in DNA is a promising approach to high density data storage. Here the authors present a prototype sequencing-free method that uses the spatial orientation of DNA strands with super-resolution microscopy readout.
- George D. Dickinson
- , Golam Md Mortuza
- & William L. Hughes
-
Article
| Open Access multiSLIDE is a web server for exploring connected elements of biological pathways in multi-omics data
multiSLIDE is a web server for exploring connected elements of biological pathways in multi-omics data
The integration and interpretation of different omics data types is an ongoing challenge for biologists. Here, the authors present a web-based, interactive tool called multiSLIDE for the visualization of protein, phosphoprotein, and RNA data presented as interlinked heatmaps.
- Soumita Ghosh
- , Abhik Datta
- & Hyungwon Choi
-
Article
| Open Access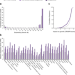 The landscape of molecular chaperones across human tissues reveals a layered architecture of core and variable chaperones
The landscape of molecular chaperones across human tissues reveals a layered architecture of core and variable chaperones
Tissue-specific differences in protein folding capacities are poorly understood. Here, the authors show that the human chaperone system consists of ubiquitous core chaperones and tissue-specific variable chaperones, perturbation of which leads to tissue-specific phenotypes.
- Netta Shemesh
- , Juman Jubran
- & Esti Yeger-Lotem
-
Article
| Open Access Differential controls of MAIT cell effector polarization by mTORC1/mTORC2 via integrating cytokine and costimulatory signals
Differential controls of MAIT cell effector polarization by mTORC1/mTORC2 via integrating cytokine and costimulatory signals
Mucosal-associated invariant T (MAIT) cells are key in immunity and diseases, but how their effector polarization is controlled is still unclear. Here, the authors show that an IL-1β/IL-23/mTORC2 axis is essential for the induction of IL-17-producing MAIT17, while an IL-2/IL-15/mTORC1 axis is important for the homeostasis of IFN-γ-producing MAIT1.
- Huishan Tao
- , Yun Pan
- & Xiao-Ping Zhong
-
Article
| Open Access Single strain control of microbial consortia
Single strain control of microbial consortia
Engineered microbial communities can divide labour between their members and interface with natural microbiomes. Here the authors demonstrate how a single toxin producing engineered strain can tune the composition of a two-strain community.
- Alex J. H. Fedorec
- , Behzad D. Karkaria
- & Chris P. Barnes
-
Article
| Open Access Single-cell analyses of Crohn’s disease tissues reveal intestinal intraepithelial T cells heterogeneity and altered subset distributions
Single-cell analyses of Crohn’s disease tissues reveal intestinal intraepithelial T cells heterogeneity and altered subset distributions
Crohn’s disease results from transmural inflammation in the gut, but analyses of local immune populations are still lacking. Here, the authors show, by combining multiple single-cell approaches, that intraepithelial and lamina propria T cells are heterogenous, show unique phenotypes, and exhibit altered subsets upon inflammation.
- Natalia Jaeger
- , Ramya Gamini
- & Marco Colonna
-
Article
| Open Access A rationally engineered decoder of transient intracellular signals
A rationally engineered decoder of transient intracellular signals
Cells encode information by modulating signal dynamics. Here the authors developed a rapid prototyping tool, TopoDesign, to engineer a synthetic short-pulse decoder in yeast.
- Claude Lormeau
- , Fabian Rudolf
- & Jörg Stelling
-
Article
| Open Access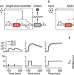 Harnessing the central dogma for stringent multi-level control of gene expression
Harnessing the central dogma for stringent multi-level control of gene expression
Inducible gene expression systems should minimise leaky output and offer a large achievable range of expression. Here, the authors regulate transcription and translation together to suppress noise and create digital-like responses, while maintaining a large expression range in vivo and in vitro.
- F. Veronica Greco
- , Amir Pandi
- & Thomas E. Gorochowski
-
Article
| Open Access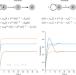 Nonlinear delay differential equations and their application to modeling biological network motifs
Nonlinear delay differential equations and their application to modeling biological network motifs
Network motif models focus on small sub-networks in biological systems to quantitatively describe overall behavior but they often overlook time delays. Here, the authors systematically examine the most common network motifs via delay differential equations (DDE), often leading to more concise descriptions.
- David S. Glass
- , Xiaofan Jin
- & Ingmar H. Riedel-Kruse
-
Article
| Open Access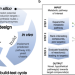 A computational workflow for the expansion of heterologous biosynthetic pathways to natural product derivatives
A computational workflow for the expansion of heterologous biosynthetic pathways to natural product derivatives
The top down cheminformatics method is usually used for the reconstitution of heterologous pathway to produce plant natural products. Here, the authors report a bottom up computational workflow for the identification of potential products and the enzymes required to make them in a noscapine pathway in yeast.
- Jasmin Hafner
- , James Payne
- & Christina Smolke
-
Article
| Open Access dCas9 regulator to neutralize competition in CRISPRi circuits
dCas9 regulator to neutralize competition in CRISPRi circuits
CRISPRi allows for the simultaneous control of many genes, however the sgRNAs compete for binding to dCas9. Here the authors design a dCas9 concentration regulator to allow independent regulation of multiple genes.
- Hsin-Ho Huang
- , Massimo Bellato
- & Domitilla Del Vecchio
-
Article
| Open Access Interactions between timing and transmissibility explain diverse flavivirus dynamics in Fiji
Interactions between timing and transmissibility explain diverse flavivirus dynamics in Fiji
Dengue and Zika virus are closely related flaviviruses but can have contrasting transmission dynamics in the same populations. Here, the authors use a model combining serological, surveillance and viral sequence data to explain differences in transmission dynamics in Fiji.
- Alasdair D. Henderson
- , Mike Kama
- & Adam J. Kucharski
-
Article
| Open Access A computer-guided design tool to increase the efficiency of cellular conversions
A computer-guided design tool to increase the efficiency of cellular conversions
Transcription factor over-expression-based cellular conversion methods often endure low conversion efficiency. Here the authors show how to increase conversion efficiency by combining a computational method for prioritizing more efficient TF combinations with a transposon-based genomic integration system for delivery.
- Sascha Jung
- , Evan Appleton
- & Antonio del Sol
-
Article
| Open Access Aptardi predicts polyadenylation sites in sample-specific transcriptomes using high-throughput RNA sequencing and DNA sequence
Aptardi predicts polyadenylation sites in sample-specific transcriptomes using high-throughput RNA sequencing and DNA sequence
Short read RNA sequencing and DNA sequence contain useful information for profiling polyadenylation sites, but each also possesses inherent limitations when examined independently. Aptardi combines these data and significantly improves annotation of polyadenylation sites in the expressed transcriptome.
- Ryan Lusk
- , Evan Stene
- & Laura M. Saba
-
Article
| Open Access Protein context shapes the specificity of SH3 domain-mediated interactions in vivo
Protein context shapes the specificity of SH3 domain-mediated interactions in vivo
The SRC Homology 3 (SH3) domains mediate protein–protein interactions (PPIs). Here, the authors assess the SH3-mediated PPIs in yeast, and show that the identity of the protein itself and the position of the SH3 both affect the interaction specificity and thus the PPI-dependent cellular functions.
- Ugo Dionne
- , Émilie Bourgault
- & Christian R. Landry
-
Article
| Open Access Overcoming the design, build, test bottleneck for synthesis of nonrepetitive protein-RNA cassettes
Overcoming the design, build, test bottleneck for synthesis of nonrepetitive protein-RNA cassettes
Phage-coat proteins can be used to build synthetic biology parts. Here the authors use an oligo library and machine learning to predict and verify sequences based on binding scores.
- Noa Katz
- , Eitamar Tripto
- & Roee Amit
-
Comment
| Open Access Seeding the idea of encapsulating a representative synthetic metagenome in a single yeast cell
Seeding the idea of encapsulating a representative synthetic metagenome in a single yeast cell
Synthetic metagenomics could potentially unravel the complexities of microbial ecosystems by revealing the simplicity of microbial communities captured in a single cell. Conceptionally, a yeast cell carrying a representative synthetic metagenome could uncover the complexity of multi-species interactions, illustrated here with wine ferments.
- Ignacio Belda
- , Thomas C. Williams
- & Isak S. Pretorius
-
Article
| Open Access A self-organized synthetic morphogenic liposome responds with shape changes to local light cues
A self-organized synthetic morphogenic liposome responds with shape changes to local light cues
The authors generated a Synthetic Morphogenic Membrane System by encapsulating a dynamic microtubule aster and a light-inducible signaling system driven by GTP/ATP chemical potential into cell-sized liposomes. This reconstitution of artificial proto-cells reveals how non-equilibrium phenomena affect cellular information processing in morphogenesis.
- Konstantin Gavriljuk
- , Bruno Scocozza
- & Philippe I. H. Bastiaens
-
Article
| Open Access Evolution of the locomotor skeleton in Anolis lizards reflects the interplay between ecological opportunity and phylogenetic inertia
Evolution of the locomotor skeleton in Anolis lizards reflects the interplay between ecological opportunity and phylogenetic inertia
Both ecological opportunity and phenotypic modularity have been suggested to facilitate adaptive radiations. Feiner et al. show that Anolis lizards evolved a new modularity structure in their island adaptive radiation, but that this modularity did not produce the same extreme diversification when Anolis returned to the mainland.
- Nathalie Feiner
- , Illiam S. C. Jackson
- & Tobias Uller
-
Article
| Open Access Epigenetic modulation reveals differentiation state specificity of oncogene addiction
Epigenetic modulation reveals differentiation state specificity of oncogene addiction
Epigenetic mechanisms associated with the differentiation state of cancer cells and their heterogeneity influence tumor responses to oncogene-targeted therapies. In this study, the authors perform an epigenetic compound screen and single-cell analysis in BRAF-mutant melanoma cells to identify compounds that block three distinct drug-tolerant epigenetic states associated with either one of the lysine-specific histone demethylases Kdm1a or Kdm4b, or BET proteins.
- Mehwish Khaliq
- , Mohan Manikkam
- & Mohammad Fallahi-Sichani
-
Article
| Open Access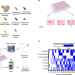 Complex yeast–bacteria interactions affect the yield of industrial ethanol fermentation
Complex yeast–bacteria interactions affect the yield of industrial ethanol fermentation
Industrial sugarcane ethanol fermentations are accomplished by a microbial community dominated by S. cerevisiae and co-occurring bacteria. Here, the authors investigate how microbial community composition contributes to community function and reveal the role of acetaldehyde in improving yeast growth rate and ethanol production.
- Felipe Senne de Oliveira Lino
- , Djordje Bajic
- & Morten Otto Alexander Sommer
-
Article
| Open Access Engineered yeast genomes accurately assembled from pure and mixed samples
Engineered yeast genomes accurately assembled from pure and mixed samples
The cost and complexity of whole genome sequencing limits its use in identifying and validating sequences used for genetic engineering and synthetic biology. Here the authors present Prymetime, an integrated workflow to sequence engineered strains and identify engineering in metagenomes.
- Joseph H. Collins
- , Kevin W. Keating
- & Eric M. Young
-
Article
| Open Access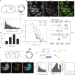 Single-cell measurement of plasmid copy number and promoter activity
Single-cell measurement of plasmid copy number and promoter activity
A quantitative assessment of promoter function can improve the precision of cellular engineering. Here the authors develop a method to simultaneously count plasmid DNA, RNA transcripts and protein expression in single living bacteria.
- Bin Shao
- , Jayan Rammohan
- & Christopher A. Voigt
-
Article
| Open Access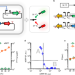 Reversible thermal regulation for bifunctional dynamic control of gene expression in Escherichia coli
Reversible thermal regulation for bifunctional dynamic control of gene expression in Escherichia coli
Genetic circuits can be built with bifunctional dynamic regulation of gene expression. Here the authors design a thermosensitive switch for spatial and temporal control of colony pattern, cell shape and polymer production.
- Xuan Wang
- , Jia-Ning Han
- & Guo-Qiang Chen
-
Review Article
| Open Access DNA stability: a central design consideration for DNA data storage systems
DNA stability: a central design consideration for DNA data storage systems
DNA has the potential to store vast amounts of data but it is subject to physical decay. In this Perspective, the authors propose that the stability of DNA should be a key consideration in how it is used for data storage.
- Karishma Matange
- , James M. Tuck
- & Albert J. Keung
-
Article
| Open Access Oscillations of Delta-like1 regulate the balance between differentiation and maintenance of muscle stem cells
Oscillations of Delta-like1 regulate the balance between differentiation and maintenance of muscle stem cells
The cell source and dynamics of Notch ligands during the regulation of muscle stem cells is unclear. Here, the authors show that the Notch ligand Dll1 has to oscillate in order to control the balance between self-renewal and differentiation of muscle stem cells, with Hes1 acting as transcriptional pacemaker for the oscillatory network.
- Yao Zhang
- , Ines Lahmann
- & Carmen Birchmeier
-
Article
| Open Access Quantifying information accumulation encoded in the dynamics of biochemical signaling
Quantifying information accumulation encoded in the dynamics of biochemical signaling
Understanding how cells discriminate between stimuli is an ongoing challenge. Here, the authors propose a mathematical framework for inferring the mutual information encoded in temporal signaling dynamics and use it to study how information is transmitted over time in response to different stimuli in NFκB, MAPK and p53 signaling pathways.
- Ying Tang
- , Adewunmi Adelaja
- & Alexander Hoffmann
-
Article
| Open Access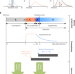 Harnessing peak transmission around symptom onset for non-pharmaceutical intervention and containment of the COVID-19 pandemic
Harnessing peak transmission around symptom onset for non-pharmaceutical intervention and containment of the COVID-19 pandemic
Transmission by pre-symptomatic and asymptomatic viral carriers makes intervention and containment of the COVID-19 extremely challenging. Here, the authors construct an epidemiological model that focuses on transmission around the symptom onset, exploring specific transmission control measures.
- Liang Tian
- , Xuefei Li
- & Lei-Han Tang
-
Article
| Open Access Robust inference of kinase activity using functional networks
Robust inference of kinase activity using functional networks
Kinases drive fundamental changes in cell state, but predicting kinase activity based on substrate-level changes can be challenging. Here the authors introduce a computational framework that utilizes similarities between substrates to robustly infer kinase activity.
- Serhan Yılmaz
- , Marzieh Ayati
- & Mehmet Koyutürk
-
Article
| Open Access Social networks predict the life and death of honey bees
Social networks predict the life and death of honey bees
Honey bee workers take on different tasks for the colony as they age. Here, the authors develop a method to extract a descriptor of the individuals’ social networks and show that interaction patterns predict task allocation and distinguish different developmental trajectories.
- Benjamin Wild
- , David M. Dormagen
- & Tim Landgraf
-
Article
| Open Access Single-cell transcriptional profiling of human thymic stroma uncovers novel cellular heterogeneity in the thymic medulla
Single-cell transcriptional profiling of human thymic stroma uncovers novel cellular heterogeneity in the thymic medulla
The thymus supports T cell immunity by providing the environment for thymocyte differentiation. Here the authors profile human thymic stroma at the single cell level, identifying ionocytes as a new medullary population and defining tissue specific antigen expression in multiple stromal cell types.
- Jhoanne L. Bautista
- , Nathan T. Cramer
- & Audrey V. Parent
-
Article
| Open Access Inference and analysis of cell-cell communication using CellChat
Inference and analysis of cell-cell communication using CellChat
Single-cell methods record molecule expressions of cells in a given tissue, but understanding interactions between cells remains challenging. Here the authors show by applying systems biology and machine learning approaches that they can infer and analyze cell-cell communication networks in an easily interpretable way.
- Suoqin Jin
- , Christian F. Guerrero-Juarez
- & Qing Nie
-
Article
| Open Access Construction of intracellular asymmetry and asymmetric division in Escherichia coli
Construction of intracellular asymmetry and asymmetric division in Escherichia coli
Establishing protein gradients for asymmetric cell division is fundamental across all kingdoms of life. Here the authors construct asymmetric cell division in E. coli by localizing the expression of RNA polymerase using an orthogonal unipolar scaffold, and restricting diffusion of its products.
- Da-Wei Lin
- , Yang Liu
- & Hsiao-Chun Huang
-
Article
| Open Access p53 dynamics vary between tissues and are linked with radiation sensitivity
p53 dynamics vary between tissues and are linked with radiation sensitivity
p53 mediates the response to irradiation, however, tissues with similar levels of p53 have different radiation sensitivities. Here, the authors show that the in vivo p53 dynamics varies in these tissues after radiation, and the use of Mdm2 inhibitor to sustain p53 activity enhances radiosensitivity.
- Jacob Stewart-Ornstein
- , Yoshiko Iwamoto
- & Galit Lahav
Browse broader subjects
Browse narrower subjects
- Bayesian inference
- Biobricks
- Biochemical networks
- Bioenergetics
- Cellular noise
- Complexity
- Computer modelling
- Computer science
- Control theory
- Criticality
- Differential equations
- DNA computing and cryptography
- Dynamic networks
- Dynamical systems
- Emergence
- Evolvability
- Genetic circuit engineering
- Genetic interaction
- Genomic engineering
- Information theory
- Logic gates
- Metabolic engineering
- Modularity
- Molecular engineering
- Molecular fluctuations
- Multicellular systems
- Multistability
- Nonlinear dynamics
- Numerical simulations
- Oscillators
- Population dynamics
- Programming language
- Protein engineering
- Regulatory networks
- Reverse engineering
- Robustness
- Signal processing
- Single-cell imaging
- Software
- Standardization
- Stochastic modelling
- Stochastic networks
- Synthetic biology
- Systems analysis
- Time series

 Design of synthetic human gut microbiome assembly and butyrate production
Design of synthetic human gut microbiome assembly and butyrate production