Featured
-
-
Article |
 Accelerated cryo-EM-guided determination of three-dimensional RNA-only structures
Accelerated cryo-EM-guided determination of three-dimensional RNA-only structures
The Ribosolve pipeline combines single-particle cryo-EM, M2-seq biochemical analysis and Rosetta auto-DRRAFTER modeling to guide three-dimensional RNA structure determination.
- Kalli Kappel
- , Kaiming Zhang
- & Rhiju Das
-
Article |
 Measurement of atom resolvability in cryo-EM maps with Q-scores
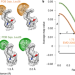
Measurement of atom resolvability in cryo-EM maps with Q-scores
Q-scores provide a quantitative metric for resolvability in cryo-EM maps, and can be used at the atom, residue or macromolecule scale.
- Grigore Pintilie
- , Kaiming Zhang
- & Wah Chiu
-
-
Article |
 Template-free detection and classification of membrane-bound complexes in cryo-electron tomograms
Template-free detection and classification of membrane-bound complexes in cryo-electron tomograms
A template-free image processing approach automatically detects and classifies membrane-bound protein complexes in cryo-electron tomograms of isolated endoplasmic reticulum and in intact cells.
- Antonio Martinez-Sanchez
- , Zdravko Kochovski
- & Vladan Lučić
-
Article |
 In situ structure determination at nanometer resolution using TYGRESS
In situ structure determination at nanometer resolution using TYGRESS
The unique advantages of single-particle cryo-electron microscopy and cryo-electron tomography are combined in a method called TYGRESS, here applied to determine the structure of the intact ciliary axoneme at a resolution of 12 Å.
- Kangkang Song
- , Zhiguo Shang
- & Daniela Nicastro
-
Article |
 Bottom-up structural proteomics: cryoEM of protein complexes enriched from the cellular milieu
Bottom-up structural proteomics: cryoEM of protein complexes enriched from the cellular milieu
An approach combining cryo-electron microscopy and mass spectrometry analysis of protein complexes enriched directly from cells enables structure determination of unknown complexes at atomic resolution.
- Chi-Min Ho
- , Xiaorun Li
- & Z. Hong Zhou
-
Article |
 A complete data processing workflow for cryo-ET and subtomogram averaging
A complete data processing workflow for cryo-ET and subtomogram averaging
An integrated pipeline for processing cryo-ET data implemented in EMAN2 streamlines data processing to minimize human bias, and improves the quality and resolution of resulting macromolecular structures, both in vitro and in cells.
- Muyuan Chen
- , James M. Bell
- & Steven J. Ludtke
-
Article |
 Real-time cryo-electron microscopy data preprocessing with Warp
Real-time cryo-electron microscopy data preprocessing with Warp
The user-friendly software tool Warp enables automated, on-the-fly preprocessing of cryo-EM data, including motion correction, defocus estimation, particle picking and image denoising.
- Dimitry Tegunov
- & Patrick Cramer
-
Article |
 Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs
Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs
The challenge of accurate particle picking in cryo-EM analysis is addressed with Topaz, a neural-network-based algorithm that shows advantages over other tools, especially in picking unusually shaped particles.
- Tristan Bepler
- , Andrew Morin
- & Bonnie Berger
-
News & Views |
 Electrons see the light
Electrons see the light
Focused laser light can manipulate the phase of electrons and solve the long-standing phase contrast conundrum in electron microscopy.
- Radostin Danev
-
Article |
 A cryo-FIB lift-out technique enables molecular-resolution cryo-ET within native Caenorhabditis elegans tissue
A cryo-FIB lift-out technique enables molecular-resolution cryo-ET within native Caenorhabditis elegans tissue
A technique to ‘lift out’ samples of interest from high-pressure-frozen specimens expands applications of cryo-electron tomography to multicellular organisms and tissue.
- Miroslava Schaffer
- , Stefan Pfeffer
- & Juergen M. Plitzko
-
Article |
 Protein secondary structure detection in intermediate-resolution cryo-EM maps using deep learning
Protein secondary structure detection in intermediate-resolution cryo-EM maps using deep learning
Emap2sec uses a deep convolutional neural network to assign protein secondary structures in intermediate-resolution (5–10 Å) cryo-EM maps.
- Sai Raghavendra Maddhuri Venkata Subramaniya
- , Genki Terashi
- & Daisuke Kihara
-
-
Perspective |
 Software tools for automated transmission electron microscopy
Software tools for automated transmission electron microscopy
Py-EM and SerialEM enable automated microscope control for high-throughput data acquisition in diverse transmission electron microscopy imaging experiments.
- Martin Schorb
- , Isabella Haberbosch
- & David N. Mastronarde
-
Perspective |
 The cryo-EM method microcrystal electron diffraction (MicroED)
The cryo-EM method microcrystal electron diffraction (MicroED)
This paper reviews the cryo-EM technique of microcrystal electron diffraction (MicroED), providing a broad overview of the technique, the unique structures determined, and the opportunities for future development.
- Brent L. Nannenga
- & Tamir Gonen
-
-
Editorial |
Challenges for cryo-EM
Two community challenges assess the correctness of cryo-EM structures; future challenges should help determine the most appropriate structure validation methods.
-
Article |
 A particle-filter framework for robust cryo-EM 3D reconstruction
A particle-filter framework for robust cryo-EM 3D reconstruction
A particle-filter algorithm for single-particle cryo-electron microscopy, implemented in a tool called THUNDER, provides high-dimensional parameter estimation, improving the obtainable resolution for several protein structures.
- Mingxu Hu
- , Hongkun Yu
- & Xueming Li
-
Brief Communication |
 A fully automatic method yielding initial models from high-resolution cryo-electron microscopy maps
A fully automatic method yielding initial models from high-resolution cryo-electron microscopy maps
A fully automated method for modeling protein and protein–RNA complex structure from cryo-EM data, requiring minimal user intervention, is described.
- Thomas C. Terwilliger
- , Paul D. Adams
- & Oleg V. Sobolev
-
Article |
 De novo computational RNA modeling into cryo-EM maps of large ribonucleoprotein complexes
De novo computational RNA modeling into cryo-EM maps of large ribonucleoprotein complexes
DRRAFTER, a method for RNA modeling into cryo-EM maps, generates accurate models for diverse RNA–protein complexes.
- Kalli Kappel
- , Shiheng Liu
- & Rhiju Das
-
Article |
 emClarity: software for high-resolution cryo-electron tomography and subtomogram averaging
emClarity: software for high-resolution cryo-electron tomography and subtomogram averaging
A software tool, emClarity, implements several improvements in cryo-electron tomography image-processing algorithms to achieve sub-nanometer resolution for diverse macromolecular structures.
- Benjamin A. Himes
- & Peijun Zhang
-
This Month |
 Bridget Carragher
Bridget Carragher
Speeding up spot-to-plunge in cryo-EM and how to keep a lab talking.
- Vivien Marx
-
Brief Communication |
 Reducing effects of particle adsorption to the air–water interface in cryo-EM
Reducing effects of particle adsorption to the air–water interface in cryo-EM
Reducing the length of time that protein particles spend on a sample grid prior to freezing mitigates deleterious effects caused by particle adsorption to the air–water interface in single-particle cryo-EM.
- Alex J. Noble
- , Hui Wei
- & Bridget Carragher
-
Research Highlight |
Taking inventory with shotgun EM
The combination of single-particle electron microscopy and mass spectrometry shows potential for surveys of both the structure and the identity of protein complexes in the cell.
- Allison Doerr
-
-
Tools in Brief |
cisTEM software for cryo-EM
-
Tools in Brief |
Local resolution of cryo-EM maps with MonoRes
-
Methods in Brief |
Improving the efficiency of cryo-EM
-
Brief Communication |
 Achieving better-than-3-Å resolution by single-particle cryo-EM at 200 keV
Achieving better-than-3-Å resolution by single-particle cryo-EM at 200 keV
Cryo-EM-based structure determination of macromolecular complexes at near-atomic resolution is possible using a mid-range 200-keV transmission electron microscope instrument.
- Mark A Herzik Jr
- , Mengyu Wu
- & Gabriel C Lander
-
Research Highlights |
 Seeing DNA
Seeing DNA
DNA and chromatin structures can be visualized in situ with electron tomography.
- Zachary J Lapin
-
Brief Communication |
 Addressing preferred specimen orientation in single-particle cryo-EM through tilting
Addressing preferred specimen orientation in single-particle cryo-EM through tilting
The preferred specimen orientation problem limits accuracy and resolution in structure determination by cryo-EM. Collecting data at defined sample tilts yielded near-atomic-resolution structures for the influenza hemagglutinin trimer and ribosomal biogenesis intermediates.
- Yong Zi Tan
- , Philip R Baldwin
- & Dmitry Lyumkis
-
Methods in Brief |
Single-protein detection by cryo-EM
-
Research Highlights |
Zooming into the larval zebrafish brain
A serial-section electron microscopy data set of larval zebrafish brain—imaged at several scales—provides a resource for structure–function analyses of the animals' neural circuitry.
- Nina Vogt
-
Brief Communication |
 RosettaES: a sampling strategy enabling automated interpretation of difficult cryo-EM maps
RosettaES: a sampling strategy enabling automated interpretation of difficult cryo-EM maps
RosettaES, an algorithm that uses a fragment-based sampling strategy, improves macromolecular structure modeling from cryo-EM data at 3–5-Å resolution.
- Brandon Frenz
- , Alexandra C Walls
- & Frank DiMaio
-
Tools in Brief |
GPCR function insights by cryo-EM
-
Research Highlights |
Protein holography
Low-energy holography enables imaging single proteins and protein complexes.
- Zachary J Lapin
-
Correspondence |
 MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy
MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy
MotionCor2 software corrects for beam-induced sample motion, improving the resolution of cryo-EM reconstructions.
- Shawn Q Zheng
- , Eugene Palovcak
- & David A Agard
-
Brief Communication |
 Atomic-resolution structures from fragmented protein crystals with the cryoEM method MicroED
Atomic-resolution structures from fragmented protein crystals with the cryoEM method MicroED
Fragmentation of large, imperfect crystals into nanocrystals by sonication, vortexing, or vigorous pipetting facilitates atomic-resolution analysis by the cryo-EM method MicroED.
- M Jason de la Cruz
- , Johan Hattne
- & Tamir Gonen
-
Article |
 cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination
cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination
A software tool, cryoSPARC, addresses the speed bottleneck in cryo-EM image processing, enabling automated macromolecular structure determination in hours on a desktop computer without requiring a starting model.
- Ali Punjani
- , John L Rubinstein
- & Marcus A Brubaker
-
Method to Watch |
 Cryo-electron tomography
Cryo-electron tomography
Cryo-electron tomography may facilitate in situ structural biology on a proteomic scale.
- Allison Doerr
-
Research Highlights |
 Colorful electron microscopy
Colorful electron microscopy
A multicolor approach specifically labels multiple targets in electron microscopy images.
- Rita Strack
-
Advertising Feature: Application Note |
 Recent developments in FEI's in situ cryo-electron tomography workflow
Recent developments in FEI's in situ cryo-electron tomography workflow
- Alexander Rigort
-
Correspondence |
The democratization of cryo-EM
- David I Stuart
- , Sriram Subramaniam
- & Nicola G A Abrescia
-
Methods in Brief |
Cryo-EM with the Volta phase plate
-
Correspondence |
 EMPIAR: a public archive for raw electron microscopy image data
EMPIAR: a public archive for raw electron microscopy image data
- Andrii Iudin
- , Paul K Korir
- & Ardan Patwardhan
-
Article |
 A saposin-lipoprotein nanoparticle system for membrane proteins
A saposin-lipoprotein nanoparticle system for membrane proteins
A saposin protein–lipid nanoparticle system stabilizes diverse, fragile membrane proteins in a lipid environment for structural and functional studies.
- Jens Frauenfeld
- , Robin Löving
- & Pär Nordlund
-
Editorial |
Method of the Year 2015
The end of 'blob-ology': single-particle cryo-electron microscopy (cryo-EM) is now being used to solve macromolecular structures at high resolution.
-
Primer |
 Single-particle cryo-electron microscopy
Single-particle cryo-electron microscopy
A brief overview of how to solve a macromolecular structure using single-particle cryo-electron microscopy (cryo-EM).
- Allison Doerr
-
Historical Commentary |
 The development of cryo-EM into a mainstream structural biology technique
The development of cryo-EM into a mainstream structural biology technique
Single-particle cryo-electron microscopy (cryo-EM) has emerged over the last two decades as a technique capable of studying the structure of challenging systems. The author of this Commentary discusses some of the major historical landmarks in cryo-EM that have led to its present success.
- Eva Nogales

 Improvement of cryo-EM maps by density modification
Improvement of cryo-EM maps by density modification Structures in situ
Structures in situ