Article
|
Open Access
Featured
-
-
Article
| Open Access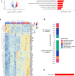 Accelerated DNA replication fork speed due to loss of R-loops in myelodysplastic syndromes with SF3B1 mutation
Accelerated DNA replication fork speed due to loss of R-loops in myelodysplastic syndromes with SF3B1 mutation
Here the authors find that erythroblasts of myelodysplastic syndromes with SF3B1 mutation leading to inefficient erythropoiesis show DNA replication stress with accelerated forks and reduced R-loops. Restoring R-loops by a histone deacetylase inhibitor rescues erythroid differentiation.
- David Rombaut
- , Carine Lefèvre
- & Michaela Fontenay
-
Article
| Open Access FANCJ promotes PARP1 activity during DNA replication that is essential in BRCA1 deficient cells
FANCJ promotes PARP1 activity during DNA replication that is essential in BRCA1 deficient cells
Here the authors show that PARPi efficacy along with the fitness of BRCA1 deficient cells relies on FANCJ, which maintains S-phase PARP1 activity by preventing its sequestration with MSH2 on G-quadruplexes.
- Ke Cong
- , Nathan MacGilvary
- & Sharon B. Cantor
-
Article
| Open Access ZNF827 is a single-stranded DNA binding protein that regulates the ATR-CHK1 DNA damage response pathway
ZNF827 is a single-stranded DNA binding protein that regulates the ATR-CHK1 DNA damage response pathway
Here, the authors characterise the zinc finger protein ZNF827 as a single stranded DNA binding protein that accumulates at stalled replication forks to activate the ATR-CHK1 pathway and engage homologous-recombination mediated DNA repair.
- Sile F. Yang
- , Christopher B. Nelson
- & Hilda A. Pickett
-
Article
| Open Access RTF2 controls replication repriming and ribonucleotide excision at the replisome
RTF2 controls replication repriming and ribonucleotide excision at the replisome
Ribonucleotides are incorporated into DNA during every S phase. Here, the authors show that replisome protein RTF2 localizes RNase H2 to the replisome, promoting ribonucleotide removal by RNase H2 when replication is ongoing but interfering with PRIM1-dependent restart following fork stalling.
- Brooke A. Conti
- , Penelope D. Ruiz
- & Agata Smogorzewska
-
Article
| Open Access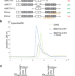 TopBP1 utilises a bipartite GINS binding mode to support genome replication
TopBP1 utilises a bipartite GINS binding mode to support genome replication
Effective and regulated activation of the Mcm2-7 helicase underlies faithful genome replication. Here the authors reveal mechanistic detail how the pre-loading complex proteins TopBP1 and GINS interact and, thus, how the helicase activator GINS loads on Mcm2-7 during replication origin firing.
- Matthew Day
- , Bilal Tetik
- & Dominik Boos
-
Article
| Open Access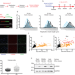 Oncogenic c-Myc induces replication stress by increasing cohesins chromatin occupancy in a CTCF-dependent manner
Oncogenic c-Myc induces replication stress by increasing cohesins chromatin occupancy in a CTCF-dependent manner
Here the authors report that oncogenic c-Myc induces replication stress via increasing the amount of cohesins bound to chromatin at CTCF sites.
- Silvia Peripolli
- , Leticia Meneguello
- & Robertus A. M. de Bruin
-
Article
| Open Access The chromatin-associated lncREST ensures effective replication stress response by promoting the assembly of fork signaling factors
The chromatin-associated lncREST ensures effective replication stress response by promoting the assembly of fork signaling factors
Replication stress represents a major threat to genome integrity of normal and cancer cells. Here, the authors find that the long non-coding RNA lncREST affects the replication stress response through interaction with nucleolin. This interaction bridges the recruitment of replication factors to stressed chromatin.
- Luisa Statello
- , José Miguel Fernandez-Justel
- & Maite Huarte
-
Article
| Open Access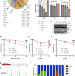 FLIP(C1orf112)-FIGNL1 complex regulates RAD51 chromatin association to promote viability after replication stress
FLIP(C1orf112)-FIGNL1 complex regulates RAD51 chromatin association to promote viability after replication stress
Recombination is essential for life. Here, the authors characterize FLIP as a novel regulator of the key recombination protein RAD51’s functions. FLIP loss caused marked sensitivity to DNA damage, increased DNA breakage and defective replication.
- Jessica D. Tischler
- , Hiroshi Tsuchida
- & Richard O. Adeyemi
-
Article
| Open Access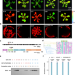 Primase promotes the competition between transcription and replication on the same template strand resulting in DNA damage
Primase promotes the competition between transcription and replication on the same template strand resulting in DNA damage
Resolving R-loops caused by transcription-replication conflicts (TRCs) is vital to genome stability in organisms. Here, the authors show that the chloroplast-localized primase ATH intensifies template strand competition and exacerbates the Head-On TRCs induced DNA damage.
- Weifeng Zhang
- , Zhuo Yang
- & Qianwen Sun
-
Article
| Open Access Structural basis for DNA proofreading
Structural basis for DNA proofreading
Here, the authors use cryo-EM to capture nine intermediates along the DNA proofreading pathway using human mitochondrial DNA Polymerase Gamma. The results provide a step-by-step view of the DNA proofreading at single-nucleotide resolution.
- Gina Buchel
- , Ashok R. Nayak
- & Dmitry Temiakov
-
Article
| Open Access Molecular basis for proofreading by the unique exonuclease domain of Family-D DNA polymerases
Molecular basis for proofreading by the unique exonuclease domain of Family-D DNA polymerases
Family D replicative DNA polymerases (PolD) contain a unique proofreading active site. Here, the authors present structures of PolD and enzymatic studies, revealing an unanticipated correction mechanism that extends the repertoire of protein domains known to be involved in DNA proofreading.
- Leonardo Betancurt-Anzola
- , Markel Martínez-Carranza
- & Ludovic Sauguet
-
Article
| Open Access The bacterial replication origin BUS promotes nucleobase capture
The bacterial replication origin BUS promotes nucleobase capture
Here, the authors show that a DnaA:origin complex promotes specific nucleobase capture from a single DNA strand. It is proposed that this mechanism may play a key role stimulating opening of bacterial chromosome origins.
- Simone Pelliciari
- , Salomé Bodet-Lefèvre
- & Heath Murray
-
Article
| Open Access RIF1 regulates early replication timing in murine B cells
RIF1 regulates early replication timing in murine B cells
Here the authors show that in activated B cells, RIF1 primarily binds early-replicating active chromatin and promotes early replication. RIF1 and MCM proteins establish early replication timing signatures genome-wide and ensure early replication of highly transcribed genes.
- Daniel Malzl
- , Mihaela Peycheva
- & Rushad Pavri
-
Article
| Open Access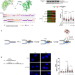 CaMKK2 and CHK1 phosphorylate human STN1 in response to replication stress to protect stalled forks from aberrant resection
CaMKK2 and CHK1 phosphorylate human STN1 in response to replication stress to protect stalled forks from aberrant resection
Here the authors show that the calcium-sensing kinase CaMKK2 phosphorylates STN1 in response to replication stress and elevated cytosolic calcium concentration to protect stalled replication forks from aberrant MRE11 degradation. Cancer-associated STN1 mutations abolish STN1 phosphorylation, resulting in fork instability.
- Rishi Kumar Jaiswal
- , Kai-Hang Lei
- & Weihang Chai
-
Article
| Open Access Arabidopsis telomerase takes off by uncoupling enzyme activity from telomere length maintenance in space
Arabidopsis telomerase takes off by uncoupling enzyme activity from telomere length maintenance in space
Telomeres are proposed to be sentinels for stress. Here, the authors report a strong induction of telomerase in space-flown Arabidopsis without telomere length changes. Instead, telomerase activity is inversely correlated with genome oxidation
- Borja Barbero Barcenilla
- , Alexander D. Meyers
- & Dorothy E. Shippen
-
Article
| Open Access Nuclear actin polymerization rapidly mediates replication fork remodeling upon stress by limiting PrimPol activity
Nuclear actin polymerization rapidly mediates replication fork remodeling upon stress by limiting PrimPol activity
How nuclear architecture assists the replication stress response is still largely unknown. Here the authors show that nuclear actin polymerization rapidly extends upon mild DNA damage. By limiting Primpol activity, this response mediates fork slowing and reversal, protecting chromosome stability.
- Maria Dilia Palumbieri
- , Chiara Merigliano
- & Massimo Lopes
-
Article
| Open Access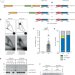 Gene duplication and deletion caused by over-replication at a fork barrier
Gene duplication and deletion caused by over-replication at a fork barrier
Gene duplications and deletions are important drivers of evolution and disease. Here, the authors show that excess DNA generated at a replication fork barrier can be integrated at a new genomic site causing both a gene duplication and a deletion.
- Judith Oehler
- , Carl A. Morrow
- & Matthew C. Whitby
-
Article
| Open Access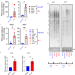 Alternative lengthening of telomeres (ALT) cells viability is dependent on C-rich telomeric RNAs
Alternative lengthening of telomeres (ALT) cells viability is dependent on C-rich telomeric RNAs
ALT cells use an alternative lengthening mechanism of telomeres and bear telomeric DNA damage with increased levels of damage-induced long non-coding RNA. Here the AUs show that antisense oligonucleotides (ASO) targeting such RNAs can induce ALT cancer cells selective cell death.
- Ilaria Rosso
- , Corey Jones-Weinert
- & Fabrizio d’Adda di Fagagna
-
Article
| Open Access Structural basis for stabilisation of the RAD51 nucleoprotein filament by BRCA2
Structural basis for stabilisation of the RAD51 nucleoprotein filament by BRCA2
Here the authors report the cryoEM structure of the RAD51 nucleoprotein filament bound to the C-terminal TR2 domain of BRCA2. The structure explains how BRCA2 stabilises the filament and uncovers a conserved mechanism of filament binding by recombination mediators.
- Robert Appleby
- , Luay Joudeh
- & Luca Pellegrini
-
Article
| Open Access A chromatinized origin reduces the mobility of ORC and MCM through interactions and spatial constraint
A chromatinized origin reduces the mobility of ORC and MCM through interactions and spatial constraint
Here the authors investigate the impact of chromatinizing origins of replication on ORC and MCM is at the single-molecule level. They find mobility of ORC reduced, but not its binding to the origin. MCM is both efficiently recruited and spatially confined to the origin.
- Humberto Sánchez
- , Zhaowei Liu
- & Nynke H. Dekker
-
Article
| Open Access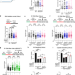 Multi-step processing of replication stress-derived nascent strand DNA gaps by MRE11 and EXO1 nucleases
Multi-step processing of replication stress-derived nascent strand DNA gaps by MRE11 and EXO1 nucleases
This study describes a multi-step nucleolytic processing of ssDNA gaps which converts them into cytotoxic double stranded breaks. Gaps can be induced not only by replication stress or genotoxic chemotherapy, but also by exposure to the environmental contaminant bisphenol A.
- Anastasia Hale
- , Ashna Dhoonmoon
- & George-Lucian Moldovan
-
Article
| Open Access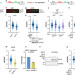 Interferon restores replication fork stability and cell viability in BRCA-defective cells via ISG15
Interferon restores replication fork stability and cell viability in BRCA-defective cells via ISG15
Here the authors show that the basal activation of the interferon/ISG15 pathway is required for the stability of nascent DNA during replication and its upregulation promotes viability, proliferation and acquisition of drug resistance in BRCA1/2 deficient cells.
- Ramona N. Moro
- , Uddipta Biswas
- & Lorenza Penengo
-
Article
| Open Access Synergism between CMG helicase and leading strand DNA polymerase at replication fork
Synergism between CMG helicase and leading strand DNA polymerase at replication fork
Coordinating the activities of replicative helicase and DNA polymerase(s) is crucial for DNA replication. Here, the authors show that DNA translocation by CMG is synchronized with the activities of Polε in the leading-strand replisome.
- Zhichun Xu
- , Jianrong Feng
- & Yuanliang Zhai
-
Article
| Open Access The COMPASS subunit Spp1 protects nascent DNA at the Tus/Ter replication fork barrier by limiting DNA availability to nucleases
The COMPASS subunit Spp1 protects nascent DNA at the Tus/Ter replication fork barrier by limiting DNA availability to nucleases
The Spp1 subunit of Set1C is recruited to the Tus/Ter replication fork barrier via its PHD domain and protects the replication fork by preventing excessive ssDNA formation, stressing the importance of chromatin structure in the replication stress response.
- Nagham Ghaddar
- , Yves Corda
- & Vincent Géli
-
Article
| Open Access RNAPII-dependent ATM signaling at collisions with replication forks
RNAPII-dependent ATM signaling at collisions with replication forks
Deregulation of transcription by oncogenes leads to collisions of RNA Polymerase II (RNAPII) with DNA replication machinery (transcription-replication conflicts, TRCs). This study shows that RNAPII activates ATM kinase at TRCs providing a mechanism for replication fork stalling and ATM activation at TRCs.
- Elias Einig
- , Chao Jin
- & Nikita Popov
-
Article
| Open Access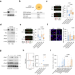 Trim33 masks a non-transcriptional function of E2f4 in replication fork progression
Trim33 masks a non-transcriptional function of E2f4 in replication fork progression
Here the authors show that under replicative stress the E2f4 transcription factor recruits the Recql DNA helicase to facilitate DNA replication. The Trim33 ubiquitin ligase targets E2f4 to limit its interactions with Recql and chromatin.
- Vanessa Rousseau
- , Elias Einig
- & Nikita Popov
-
Article
| Open Access TRAIP resolves DNA replication-transcription conflicts during the S-phase of unperturbed cells
TRAIP resolves DNA replication-transcription conflicts during the S-phase of unperturbed cells
The TRAIP E3 ubiquitin ligase is essential for genome integrity, mutations lead to primordial dwarfism in patients. Here, the authors show that TRAIP degradation in S-phase, results in cell arrest due to DNA damage caused by replication-transcription conflicts.
- Shaun Scaramuzza
- , Rebecca M. Jones
- & Agnieszka Gambus
-
Article
| Open Access NFIB facilitates replication licensing by acting as a genome organizer
NFIB facilitates replication licensing by acting as a genome organizer
The precise rule of replication origin selection and activation in metazoans remains unclear. Here, the authors identify NFIB as a genome organizer and replication pioneer by facilitating nucleosome remodeling and chromatin assembly of the pre-RC.
- Wenting Zhang
- , Yue Wang
- & Yongfeng Shang
-
Article
| Open Access C16orf72/HAPSTR1/TAPR1 functions with BRCA1/Senataxin to modulate replication-associated R-loops and confer resistance to PARP disruption
C16orf72/HAPSTR1/TAPR1 functions with BRCA1/Senataxin to modulate replication-associated R-loops and confer resistance to PARP disruption
Here the authors identify that C16orf72 regulates BRCA1/Senataxin to promote replication fork recovery. These proteins act together in a pathway parallel to PARP1 to suppress R-loop accumulation in response to replication stress and confer resistance to PARP inhibitors.
- Abhishek Bharadwaj Sharma
- , Muhammad Khairul Ramlee
- & Nicholas D. Lakin
-
Article
| Open Access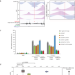 Dimeric G-quadruplex motifs-induced NFRs determine strong replication origins in vertebrates
Dimeric G-quadruplex motifs-induced NFRs determine strong replication origins in vertebrates
A program regulating replication origins ensures the exact duplication of vertebrate genomes. The authors identify a combination of guanine-rich motifs, known to form secondary DNA structures, which are sufficient to assemble efficient replication origins.
- Jérémy Poulet-Benedetti
- , Caroline Tonnerre-Doncarli
- & Marie-Noëlle Prioleau
-
Article
| Open Access ORC1 binds to cis-transcribed RNAs for efficient activation of replication origins
ORC1 binds to cis-transcribed RNAs for efficient activation of replication origins
Here the authors describe that the human origin recognition complex subunit 1 (ORC1) binds to RNAs transcribed from genes with origins of replication at their TSS impacting origin activation.
- Aina Maria Mas
- , Enrique Goñi
- & Maite Huarte
-
Article
| Open Access Structural basis of the T4 bacteriophage primosome assembly and primer synthesis
Structural basis of the T4 bacteriophage primosome assembly and primer synthesis
Here the authors present a methodical cryo-EM analysis to show how a T4 bacteriophage gp41 helicase hexamer and a gp61 primase assemble with a template-primer into an active primosome.
- Xiang Feng
- , Michelle M. Spiering
- & Huilin Li
-
Article
| Open Access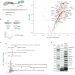 Nuclear myosin VI maintains replication fork stability
Nuclear myosin VI maintains replication fork stability
Whether actin and associated molecules have roles in the nucleus is an active area of study. Here Shi et al. report a nuclear function of the actin-based motor myosin VI in protecting stalled replication forks from nuclease-mediated degradation.
- Jie Shi
- , Kristine Hauschulte
- & Hans-Peter Wollscheid
-
Article
| Open Access Molecular choreography of primer synthesis by the eukaryotic Pol α-primase
Molecular choreography of primer synthesis by the eukaryotic Pol α-primase
DNA Polymerase α has separate RNA and DNA Polymerase subunits to form a hybrid RNA-DNA primer of unique length. Cryo-EM structures of each stage in primer synthesis reveal large movements among the four subunits and explain how primer length may be limited.
- Zuanning Yuan
- , Roxana Georgescu
- & Michael E. O’Donnell
-
Article
| Open Access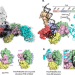 Replication fork binding triggers structural changes in the PriA helicase that govern DNA replication restart in E. coli
Replication fork binding triggers structural changes in the PriA helicase that govern DNA replication restart in E. coli
The mechanism of replication restart initiation by the bacterial DNA replication restart proteins PriA and PriB is resolved, revealing a switch-like restructuring of PriA triggered by replication fork binding that mediates PriA/PriB complex assembly.
- Alexander T. Duckworth
- , Peter L. Ducos
- & James L. Keck
-
Article
| Open Access DNA-binding mechanism and evolution of replication protein A
DNA-binding mechanism and evolution of replication protein A
Here the authors present the structure of Replication Protein A (RPA) in Archaea. The RPA structure from P. abyssi has been determined in presence and absence of DNA, providing insights into the evolution of this replication factor in eukaryotes
- Clément Madru
- , Markel Martínez-Carranza
- & Ludovic Sauguet
-
Article
| Open Access Replication-associated formation and repair of human topoisomerase IIIα cleavage complexes
Replication-associated formation and repair of human topoisomerase IIIα cleavage complexes
This study provides evidence that human topoisomerase IIIα (TOP3A) is closely associated with active replisomes. The authors uncover TOP3A DNA cleavage complexes (TOP3Accs) repair mechanisms to ensure normal DNA replication and genome integrity.
- Liton Kumar Saha
- , Sourav Saha
- & Yves Pommier
-
Article
| Open Access The in vivo measurement of replication fork velocity and pausing by lag-time analysis
The in vivo measurement of replication fork velocity and pausing by lag-time analysis
Lag-time analysis was developed to measure in vivo replisome dynamics. Observed dynamics are both locus and cell-cycle dependent: Pauses of seconds are observed at wild-type ribosomal DNA loci, as well as temporal fork velocity oscillations.
- Dean Huang
- , Anna E. Johnson
- & Paul A. Wiggins
-
Article
| Open Access Excessive reactive oxygen species induce transcription-dependent replication stress
Excessive reactive oxygen species induce transcription-dependent replication stress
Excessive oxidative stress is widely perceived as a key factor in cancer progression. Here, the authors reveal that oxidative stress induces transcription-dependent replication fork stalling that appears to be a major source of chromosomal rearrangements found in human cancers.
- Martin Andrs
- , Henriette Stoy
- & Pavel Janscak
-
Article
| Open Access DNA replication initiation factor RECQ4 possesses a role in antagonizing DNA replication initiation
DNA replication initiation factor RECQ4 possesses a role in antagonizing DNA replication initiation
RECQ4 mutations contribute to multiple developmental diseases and tumorigenesis. Here the authors describe how a highly oncogenic RECQ4 mutation alters the control of DNA synthesis, leading to abnormal DNA content and cell growth.
- Xiaohua Xu
- , Chou-Wei Chang
- & Yilun Liu
-
Article
| Open Access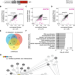 Genetic requirements for repair of lesions caused by single genomic ribonucleotides in S phase
Genetic requirements for repair of lesions caused by single genomic ribonucleotides in S phase
RNase H2 removes mutagenic rNMPs from genomic DNA. The authors demonstrate how nicked rNMPs are repaired when they are encountered in S phase. This study unveils genetic interactions that could potentially be exploited in RNase H2-deficient pathologies.
- Natalie Schindler
- , Matthias Tonn
- & Brian Luke
-
Article
| Open Access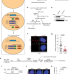 Mitotic DNA synthesis in response to replication stress requires the sequential action of DNA polymerases zeta and delta in human cells
Mitotic DNA synthesis in response to replication stress requires the sequential action of DNA polymerases zeta and delta in human cells
DNA replication stress can generate under-replicated DNA regions which is fixed by an atypical form of DNA repair synthesis in mitosis (MiDAS). Here the authors show that translesion and replicative DNA polymerases cooperate via the POLD3 subunit to complete MiDAS in human cells.
- Wei Wu
- , Szymon A. Barwacz
- & Ying Liu
-
Article
| Open Access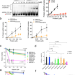 Replication gap suppression depends on the double-strand DNA binding activity of BRCA2
Replication gap suppression depends on the double-strand DNA binding activity of BRCA2
Here the authors demonstrate that the dsDNA binding function at the N-terminus of BRCA2 prevents nucleotide depletion-dependent replicative ssDNA gaps but not those induced by PARP inhibition. This function is impaired in breast-cancer variants affecting this region.
- Domagoj Vugic
- , Isaac Dumoulin
- & Aura Carreira
-
Article
| Open Access Global landscape of replicative DNA polymerase usage in the human genome
Global landscape of replicative DNA polymerase usage in the human genome
Profiling of human DNA polymerase Polε and Polα demonstrates their roles in leading and lagging strand DNA synthesis, and their independent measures allowed accurate predictions of replication dynamics and effects of transcription.
- Eri Koyanagi
- , Yoko Kakimoto
- & Yasukazu Daigaku
-
Article
| Open Access The non-catalytic role of DNA polymerase epsilon in replication initiation in human cells
The non-catalytic role of DNA polymerase epsilon in replication initiation in human cells
DNA polymerase epsilon has a critical role in DNA replication initiation. Here, the authors show that in human cancer cells POLE is dispensable for the replicative helicase assembly but not for replication initiation, which requires the non-catalytic domain of POLE1.
- Sameera Vipat
- , Dipika Gupta
- & Tatiana N. Moiseeva
-
Article
| Open Access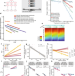 Cell division drives DNA methylation loss in late-replicating domains in primary human cells
Cell division drives DNA methylation loss in late-replicating domains in primary human cells
DNA methylation loss has been observed in aging tissues and cancers for decades. Researchers from Van Andel Institute have now provided experimental evidence that this process is directly driven by cell division.
- Jamie L. Endicott
- , Paula A. Nolte
- & Peter W. Laird
-
Article
| Open Access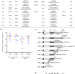 Pathogenic variants in SLF2 and SMC5 cause segmented chromosomes and mosaic variegated hyperploidy
Pathogenic variants in SLF2 and SMC5 cause segmented chromosomes and mosaic variegated hyperploidy
The SMC5/6 complex is critical for genome stability. Here, the authors identify mutations in SLF2 and SMC5 as cause of Atelís Syndrome characterized by microcephaly, short stature, anemia, segmented chromosomes and mosaic variegated hyperploidy.
- Laura J. Grange
- , John J. Reynolds
- & Grant S. Stewart
-
Article
| Open Access Cooperative assembly of p97 complexes involved in replication termination
Cooperative assembly of p97 complexes involved in replication termination
This study describes how p97Ufd1-Npl4 and the UBA-UBX protein Ubxn7 disassemble vertebrate replisomes during replication termination, and it provides novel insights into how p97 complexes assemble with UBA-UBX proteins on ubiquitylated substrates
- Olga V. Kochenova
- , Sirisha Mukkavalli
- & Johannes C. Walter
-
Article
| Open Access Profilin-1 regulates DNA replication forks in a context-dependent fashion by interacting with SNF2H and BOD1L
Profilin-1 regulates DNA replication forks in a context-dependent fashion by interacting with SNF2H and BOD1L
Subcellular localization plays an important yet underappreciated role in protein functions. Here the authors defined novel and context-dependent nuclear functions of the actin-binding factor profilin-1 in DNA replication fork dynamics and stability.
- Cuige Zhu
- , Mari Iwase
- & Jieya Shao

 Chromosome organization shapes replisome dynamics in Caulobacter crescentus
Chromosome organization shapes replisome dynamics in Caulobacter crescentus