Article
|
Open Access
Featured
-
-
Article
| Open Access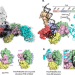 Replication fork binding triggers structural changes in the PriA helicase that govern DNA replication restart in E. coli
Replication fork binding triggers structural changes in the PriA helicase that govern DNA replication restart in E. coli
The mechanism of replication restart initiation by the bacterial DNA replication restart proteins PriA and PriB is resolved, revealing a switch-like restructuring of PriA triggered by replication fork binding that mediates PriA/PriB complex assembly.
- Alexander T. Duckworth
- , Peter L. Ducos
- & James L. Keck
-
Article
| Open Access DNA-binding mechanism and evolution of replication protein A
DNA-binding mechanism and evolution of replication protein A
Here the authors present the structure of Replication Protein A (RPA) in Archaea. The RPA structure from P. abyssi has been determined in presence and absence of DNA, providing insights into the evolution of this replication factor in eukaryotes
- Clément Madru
- , Markel Martínez-Carranza
- & Ludovic Sauguet
-
Article
| Open Access The in vivo measurement of replication fork velocity and pausing by lag-time analysis
The in vivo measurement of replication fork velocity and pausing by lag-time analysis
Lag-time analysis was developed to measure in vivo replisome dynamics. Observed dynamics are both locus and cell-cycle dependent: Pauses of seconds are observed at wild-type ribosomal DNA loci, as well as temporal fork velocity oscillations.
- Dean Huang
- , Anna E. Johnson
- & Paul A. Wiggins
-
Article
| Open Access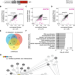 Genetic requirements for repair of lesions caused by single genomic ribonucleotides in S phase
Genetic requirements for repair of lesions caused by single genomic ribonucleotides in S phase
RNase H2 removes mutagenic rNMPs from genomic DNA. The authors demonstrate how nicked rNMPs are repaired when they are encountered in S phase. This study unveils genetic interactions that could potentially be exploited in RNase H2-deficient pathologies.
- Natalie Schindler
- , Matthias Tonn
- & Brian Luke
-
Article
| Open Access Global landscape of replicative DNA polymerase usage in the human genome
Global landscape of replicative DNA polymerase usage in the human genome
Profiling of human DNA polymerase Polε and Polα demonstrates their roles in leading and lagging strand DNA synthesis, and their independent measures allowed accurate predictions of replication dynamics and effects of transcription.
- Eri Koyanagi
- , Yoko Kakimoto
- & Yasukazu Daigaku
-
Article
| Open Access The non-catalytic role of DNA polymerase epsilon in replication initiation in human cells
The non-catalytic role of DNA polymerase epsilon in replication initiation in human cells
DNA polymerase epsilon has a critical role in DNA replication initiation. Here, the authors show that in human cancer cells POLE is dispensable for the replicative helicase assembly but not for replication initiation, which requires the non-catalytic domain of POLE1.
- Sameera Vipat
- , Dipika Gupta
- & Tatiana N. Moiseeva
-
Article
| Open Access Cooperative assembly of p97 complexes involved in replication termination
Cooperative assembly of p97 complexes involved in replication termination
This study describes how p97Ufd1-Npl4 and the UBA-UBX protein Ubxn7 disassemble vertebrate replisomes during replication termination, and it provides novel insights into how p97 complexes assemble with UBA-UBX proteins on ubiquitylated substrates
- Olga V. Kochenova
- , Sirisha Mukkavalli
- & Johannes C. Walter
-
Article
| Open Access Solving the MCM paradox by visualizing the scaffold of CMG helicase at active replisomes
Solving the MCM paradox by visualizing the scaffold of CMG helicase at active replisomes
For several decades the MCM2-7 proteins, the core of the DNA replicative helicase, eluded detection at DNA replication sites. Here, the authors solve this conundrum by gene editing, which enables visualization of replication dynamics in living cells.
- Hana Polasek-Sedlackova
- , Thomas C. R. Miller
- & Jiri Lukas
-
Article
| Open Access The mechanism of replication stalling and recovery within repetitive DNA
The mechanism of replication stalling and recovery within repetitive DNA
DNA replication of repetitive sequences was recreated in a test tube using purified components. DNA alone was sufficient to induce stalling. Both stalling and recovery were dictated by the capacity of DNA to fold into unusual secondary structures.
- Corella S. Casas-Delucchi
- , Manuel Daza-Martin
- & Gideon Coster
-
Article
| Open Access Stable inheritance of H3.3-containing nucleosomes during mitotic cell divisions
Stable inheritance of H3.3-containing nucleosomes during mitotic cell divisions
How nucleosome assembly of parental histones is regulated following DNA replication is still an open question. Here the authors show that unlike deposition of new histones H3.1 and H3.3 that utilizes different histone chaperones, parental H3.1 and H3.3 are both stably inherited during mitotic cell division in mouse embryonic stem cells, and this involves histone chaperones Mcm2, Pole3 and Pole4.
- Xiaowei Xu
- , Shoufu Duan
- & Zhiguo Zhang
-
Article
| Open Access Single-molecule imaging reveals replication fork coupled formation of G-quadruplex structures hinders local replication stress signaling
Single-molecule imaging reveals replication fork coupled formation of G-quadruplex structures hinders local replication stress signaling
In the genome, repetitive guanine-rich sequences have the potential to spontaneously fold into non-canonical DNA secondary structures known as G-quadruplex (G4). Using novel single-molecule imaging approaches, the authors reveal that G4 formation within active replication forks locally perturb replisome dynamics and damage response signaling, which require RPA and FANCJ for regulation.
- Wei Ting C. Lee
- , Yandong Yin
- & Eli Rothenberg
-
Article
| Open Access SDE2 integrates into the TIMELESS-TIPIN complex to protect stalled replication forks
SDE2 integrates into the TIMELESS-TIPIN complex to protect stalled replication forks
The fork protection complex (FPC), including the proteins TIMELESS and TIPIN, stabilizes the replisome to ensure unperturbed fork progression during DNA replication. Here the authors reveal that that SDE2, a PCNA-associated protein, plays an important role in maintaining active replication and protecting stalled forks by regulating the replication fork protection complex (FPC).
- Julie Rageul
- , Jennifer J. Park
- & Hyungjin Kim
-
Article
| Open Access Structural mechanism for replication origin binding and remodeling by a metazoan origin recognition complex and its co-loader Cdc6
Structural mechanism for replication origin binding and remodeling by a metazoan origin recognition complex and its co-loader Cdc6
The origin recognition complex (ORC) is essential for loading the Mcm2–7 replicative helicase onto DNA during DNA replication initiation. Here, the authors describe several cryo-electron microscopy structures of Drosophila ORC bound to DNA and its cofactor Cdc6 and also report an in vitro reconstitution system for Drosophila Mcm2–7 loading, revealing unexpected features of ORC’s DNA binding and remodeling mechanism during Mcm2–7 loading.
- Jan Marten Schmidt
- & Franziska Bleichert
-
Article
| Open Access DONSON and FANCM associate with different replisomes distinguished by replication timing and chromatin domain
DONSON and FANCM associate with different replisomes distinguished by replication timing and chromatin domain
Eukaryotic replisomes are multiprotein complexes. Here the authors reveal two distinct stressed replisomes, associated with DONSON and FANCM, displaying a bias in replication timing and chromatin domain.
- Jing Zhang
- , Marina A. Bellani
- & Michael M. Seidman
-
Article
| Open Access Ethanol exposure increases mutation rate through error-prone polymerases
Ethanol exposure increases mutation rate through error-prone polymerases
Whereas the toxic effects of ethanol are well-documented, the underlying mechanism is obscure. This study uses the eukaryotic model S. cerevisiae to reveal how exposure to sublethal ethanol concentrations causes DNA replication stress and an increased mutation rate.
- Karin Voordeckers
- , Camilla Colding
- & Kevin J. Verstrepen
-
Article
| Open Access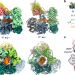 Structural basis for the increased processivity of D-family DNA polymerases in complex with PCNA
Structural basis for the increased processivity of D-family DNA polymerases in complex with PCNA
Replicative DNA polymerases (DNAPs) have evolved the ability to copy the genome with high processivity and fidelity. Here, the authors present a cryo-EM structure of the DNA-bound PolD–PCNA complex from Pyrococcus abyssi to reveal the molecular basis for the interaction and cooperativity between a replicative DNAP and PCNA.
- Clément Madru
- , Ghislaine Henneke
- & Ludovic Sauguet
-
Article
| Open Access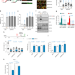 SPRTN protease and checkpoint kinase 1 cross-activation loop safeguards DNA replication
SPRTN protease and checkpoint kinase 1 cross-activation loop safeguards DNA replication
Cells deficient in SPRTN protease activity exhibit severe DNA-protein crosslink induced replication stress and genome instability. Here the author reveal a functional link between the SPRTN protease and the CHK1 kinase during physiological DNA replication.
- Swagata Halder
- , Ignacio Torrecilla
- & Kristijan Ramadan
-
Article
| Open Access DNA translocation mechanism of the MCM complex and implications for replication initiation
DNA translocation mechanism of the MCM complex and implications for replication initiation
Eukaryotes and archaea use a heximeric ring-shaped MCM helicase to unwind the DNA template during replication. Here the authors present a crystal structure of the MCM complex from archaeon S. solfataricus bound to single-stranded DNA, and to a combination of ADP, and ATP-mimic, ADP-BeF3.
- Martin Meagher
- , Leslie B. Epling
- & Eric J. Enemark
-
Article
| Open Access Structure of DNA-CMG-Pol epsilon elucidates the roles of the non-catalytic polymerase modules in the eukaryotic replisome
Structure of DNA-CMG-Pol epsilon elucidates the roles of the non-catalytic polymerase modules in the eukaryotic replisome
Eukaryotic origin firing depends on assembly of the Cdc45-MCM-GINS (CMG) helicase, which requires the leading-strand polymerase Pol ɛ. Here the authors present a structural analysis of a CMG Pol ɛ on a DNA fork, providing insight on the steps leading productive helicase engagement to the DNA junction.
- Panchali Goswami
- , Ferdos Abid Ali
- & Alessandro Costa
-
Article
| Open Access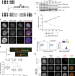 AND-1 fork protection function prevents fork resection and is essential for proliferation
AND-1 fork protection function prevents fork resection and is essential for proliferation
AND-1, the vertebrate orthologue of Ctf4, is a critical player during DNA replication and for maintenance of genome integrity. Here the authors use a conditional AND-1 depletion system in avian DT40 cells to reveal the consequences of the lack of AND-1 on cell proliferation and DNA replication.
- Takuya Abe
- , Ryotaro Kawasumi
- & Dana Branzei
-
Article
| Open Access SUMO2 conjugation of PCNA facilitates chromatin remodeling to resolve transcription-replication conflicts
SUMO2 conjugation of PCNA facilitates chromatin remodeling to resolve transcription-replication conflicts
Transcription-replication conflicts need to be resolved to minimize genome instability. Here the authors show that SUMO2-conjugated PCNA destabilizes RNAPII from chromatin, enhances replication progression and limits transcription-induced DNA damage at common fragile sites.
- Min Li
- , Xiaohua Xu
- & Yilun Liu
-
Article
| Open Access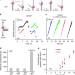 Helicase promotes replication re-initiation from an RNA transcript
Helicase promotes replication re-initiation from an RNA transcript
During DNA replication, replicative helicases play an essential role for DNA unwinding to occur. Here the authors find that bacteriophage T7 helicase is also involved in replication re-initiation by interacting with a non-replicating DNAP and increasing unwinding rate.
- Bo Sun
- , Anupam Singh
- & Michelle D. Wang
-
Article
| Open Access Chromatin conformation regulates the coordination between DNA replication and transcription
Chromatin conformation regulates the coordination between DNA replication and transcription
The maintenance of chromatin integrity during replication is critical for cell viability. Here the authors study how dividing cells respond to alterations in chromatin structure and find that these elicit a range of responses in the dynamics of DNA replication and consequences on replicative stress.
- Ricardo Almeida
- , José Miguel Fernández-Justel
- & María Gómez
-
Article
| Open Access Evidence that DNA polymerase δ contributes to initiating leading strand DNA replication in Saccharomyces cerevisiae
Evidence that DNA polymerase δ contributes to initiating leading strand DNA replication in Saccharomyces cerevisiae
DNA polymerases δ and ε (Pols δ and ε) are thought to be responsible for lagging and leading strand synthesis, respectively. Here the authors present evidence that Pol δ contributes to the initiation of leading strand replication in budding yeast by synthesizing DNA of both strands at replication origins.
- Marta A. Garbacz
- , Scott A. Lujan
- & Thomas A. Kunkel
-
Article
| Open Access Cdt1 stabilizes an open MCM ring for helicase loading
Cdt1 stabilizes an open MCM ring for helicase loading
The loading and activation of the Mcm2-7 replicative helicase couples cell cycle progression to DNA replication. Here the authors use X-ray crystallography and single-particle electron microscopy to demonstrate how Ctd1 functions to promote MCM loading onto DNA.
- Jordi Frigola
- , Jun He
- & John F. X. Diffley
-
Article
| Open Access Molecular basis for PrimPol recruitment to replication forks by RPA
Molecular basis for PrimPol recruitment to replication forks by RPA
PrimPol is a multifunctional replicative enzyme that can bypass DNA damage, as well as reprime replication restart. Here, the authors have elucidated how PrimPol is recruited to stalled replication forks via specific interactions with RPA, which stimulates its primase activity.
- Thomas A. Guilliam
- , Nigel C. Brissett
- & Aidan J. Doherty
-
Article
| Open Access Structural basis of human PCNA sliding on DNA
Structural basis of human PCNA sliding on DNA
DNA sliding clamps are ring-shaped proteins that encircle DNA and harbour polymerases and other factors that promote processive DNA replication. Here the authors use X-ray crystallography, NMR and MD simulations to propose a model for a PCNA sliding mechanism that relies on short-lived polar interactions.
- Matteo De March
- , Nekane Merino
- & Alfredo De Biasio
-
Article
| Open Access T7 replisome directly overcomes DNA damage
T7 replisome directly overcomes DNA damage
Genomic instability can result from stalled or collapsed replication fork at sites of unrepaired DNA lesions. Here the authors uncover a new lesion bypass pathway for the T7 replisome, where leading strand template lesions can be overcome through interaction between the replisome's helicase and polymerase components.
- Bo Sun
- , Manjula Pandey
- & Michelle D. Wang

 Synergism between CMG helicase and leading strand DNA polymerase at replication fork
Synergism between CMG helicase and leading strand DNA polymerase at replication fork