Featured
-
-
Letter |
 Tyrosine phosphorylation of histone H2A by CK2 regulates transcriptional elongation
Tyrosine phosphorylation of histone H2A by CK2 regulates transcriptional elongation
A conserved tyrosine residue, Tyr 57, of histone H2A is phosphorylated by an unsuspected tyrosine kinase activity of casein kinase 2, influencing a series of histone marks associated with active transcription and regulating transcription elongation.
- Harihar Basnet
- , Xue B. Su
- & Michael G. Rosenfeld
-
Letter |
 Histone H2A.Z subunit exchange controls consolidation of recent and remote memory
Histone H2A.Z subunit exchange controls consolidation of recent and remote memory
The authors identify a specific histone variant as a memory-suppressor that is initially reduced in expression within the hippocampus during memory formation; as a memory is consolidated to the cortex, reduced histone association with specific plasticity genes is observed, promoting stabilization of the memory.
- Iva B. Zovkic
- , Brynna S. Paulukaitis
- & J. David Sweatt
-
Letter
| Open Access Comparative analysis of metazoan chromatin organization
Comparative analysis of metazoan chromatin organization
A large collection of new modENCODE and ENCODE genome-wide chromatin data sets from cell lines and developmental stages in worm, fly and human are analysed; this reveals many conserved features of chromatin organization among the three organisms, as well as notable differences in the composition and locations of repressive chromatin.
- Joshua W. K. Ho
- , Youngsook L. Jung
- & Peter J. Park
-
Letter |
 Contrasting roles of histone 3 lysine 27 demethylases in acute lymphoblastic leukaemia
Contrasting roles of histone 3 lysine 27 demethylases in acute lymphoblastic leukaemia
T-cell acute lymphoblastic leukaemia (T-ALL) is a haematological malignancy with a poor prognosis and no available targeted therapies; now two histone H3 lysine 27 demethylases, JMJD3 and UTX, are shown to have contrasting roles in human T-ALL cells and a mouse model of the disease, and a small molecule demethylase inhibitor is found to inhibit the growth of T-ALL cell lines, introducing a potential therapeutic avenue for acute leukaemia.
- Panagiotis Ntziachristos
- , Aristotelis Tsirigos
- & Iannis Aifantis
-
Letter |
 Selective transcriptional regulation by Myc in cellular growth control and lymphomagenesis
Selective transcriptional regulation by Myc in cellular growth control and lymphomagenesis
Global transcriptional and epigenomic analyses in diverse cell types reveal that the primary action of Myc is to up- and downregulate transcription of distinct groups of genes, rather than to amplify transcription of all active genes; general RNA amplification, when observed, is better explained as an indirect consequence of Myc’s action on cellular physiology.
- Arianna Sabò
- , Theresia R. Kress
- & Bruno Amati
-
Letter |
 Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA
Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA
A nucleosome-spacing mechanism for human ATP-dependent chromatin assembly and remodelling factor (ACF).
- William L. Hwang
- , Sebastian Deindl
- & Xiaowei Zhuang
-
Letter |
 ZMYND11 links histone H3.3K36me3 to transcription elongation and tumour suppression
ZMYND11 links histone H3.3K36me3 to transcription elongation and tumour suppression
Candidate tumour suppressor ZMYND11 specifically recognizes histone K36 trimethylation on the histone variant H3.3 and helps regulate transcription elongation.
- Hong Wen
- , Yuanyuan Li
- & Xiaobing Shi
-
Letter |
 Two independent transcription initiation codes overlap on vertebrate core promoters
Two independent transcription initiation codes overlap on vertebrate core promoters
The transcription start sites used during the maternal to zygotic transition in zebrafish are mapped, revealing that the transition is characterized by a switch between two different sequence signs to guide transcription initiation, which often co-exist in core promoters.
- Vanja Haberle
- , Nan Li
- & Boris Lenhard
-
Letter |
 Citrullination regulates pluripotency and histone H1 binding to chromatin
Citrullination regulates pluripotency and histone H1 binding to chromatin
This study shows that PADI4-mediated citrullination occurs during pluripotency and that citrullination of H1 results in loosening of chromatin compaction; furthermore, citrullination is shown to be important for the activation of stem-cell genes, for iPS cell reprogramming and to maintain pluripotent cells in the early mouse embryo.
- Maria A. Christophorou
- , Gonçalo Castelo-Branco
- & Tony Kouzarides
-
Article |
 ANP32E is a histone chaperone that removes H2A.Z from chromatin
ANP32E is a histone chaperone that removes H2A.Z from chromatin
Human protein ANP32E is a histone chaperone that promotes removal of H2A.Z from chromatin.
- Arnaud Obri
- , Khalid Ouararhni
- & Ali Hamiche
-
Letter |
 Glutamine methylation in histone H2A is an RNA-polymerase-I-dedicated modification
Glutamine methylation in histone H2A is an RNA-polymerase-I-dedicated modification
A description of a new histone modification, methylation of glutamine, on histone H2A in yeast and human cells.
- Peter Tessarz
- , Helena Santos-Rosa
- & Tony Kouzarides
-
Letter |
 A high-resolution map of the three-dimensional chromatin interactome in human cells
A high-resolution map of the three-dimensional chromatin interactome in human cells
A novel approach to analyse high-depth Hi-C data provides a comprehensive chromatin interaction map at approximately 5–10 kb resolution in human fibroblasts; this reveals that TNF-α-responsive enhancers are already in contact with target promoters before signalling and that this chromatin looping is a strong predictor of gene induction.
- Fulai Jin
- , Yan Li
- & Bing Ren
-
Letter |
 The pluripotent genome in three dimensions is shaped around pluripotency factors
The pluripotent genome in three dimensions is shaped around pluripotency factors
Using 4C technology, higher-order topological features of the pluripotent genome are identified; in pluripotent stem cells, Nanog clusters specifically with other pluripotency genes and this clustering is centred around Nanog-binding sites, suggesting that Nanog helps to shape the three-dimensional structure of the pluripotent genome and thereby contributes to the robustness of the pluripotent state.
- Elzo de Wit
- , Britta A. M. Bouwman
- & Wouter de Laat
-
Letter |
 PfSETvs methylation of histone H3K36 represses virulence genes in Plasmodium falciparum
PfSETvs methylation of histone H3K36 represses virulence genes in Plasmodium falciparum
The malaria parasite Plasmodium falciparum escapes immune detection by expressing one of 60 antigenically distinct var genes at any one time during the course of infection: here it is shown that the P. falciparum protein PfSETvs has a key role in var gene silencing through the trimethylation of histone H3K36.
- Lubin Jiang
- , Jianbing Mu
- & Louis H. Miller
-
Article |
 53BP1 is a reader of the DNA-damage-induced H2A Lys 15 ubiquitin mark
53BP1 is a reader of the DNA-damage-induced H2A Lys 15 ubiquitin mark
This study shows that 53BP1 recruitment to sites of DNA damage involves dual recognition of H4K20me2 and H2AK15 histone ubiquitination; the ubiquitin mark and the surrounding epitope on H2A are read by a region of 53BP1 designated the ubiquitination-dependent recruitment motif.
- Amélie Fradet-Turcotte
- , Marella D. Canny
- & Daniel Durocher
-
Letter |
 The bromodomain protein Brd4 insulates chromatin from DNA damage signalling
The bromodomain protein Brd4 insulates chromatin from DNA damage signalling
Isoform B of the chromatin-binding protein Brd4 acts to suppress DNA damage response signalling.
- Scott R. Floyd
- , Michael E. Pacold
- & Michael B. Yaffe
-
Letter |
 Structural basis of histone H2A–H2B recognition by the essential chaperone FACT
Structural basis of histone H2A–H2B recognition by the essential chaperone FACT
The crystal structure of the FACT histone chaperone domain Spt16M in complex with the H2A–H2B heterodimer is solved; Spt16M makes several interactions with histones and seems to block the interaction of H2B with DNA, which could explain how FACT destabilizes nucleosomes to promote transcription.
- Maria Hondele
- , Tobias Stuwe
- & Andreas G. Ladurner
-
Letter |
 BAF complexes facilitate decatenation of DNA by topoisomerase IIα
BAF complexes facilitate decatenation of DNA by topoisomerase IIα
Mutations in the subunits of BAF chromatin-remodelling complexes are frequently found in human cancer; here deletion of BAF subunits or expression of mutants of the ATPase subunit BRG1 attenuates genome-wide binding of topoisomerase IIα, resulting in tangled chromosomes, anaphase bridges and G2/M arrest.
- Emily C. Dykhuizen
- , Diana C. Hargreaves
- & Gerald R. Crabtree
-
Letter |
 Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1
Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1
In Arabidopsis, RNA-directed DNA methylation is a poorly understood gene silencing pathway in which small interfering RNAs generated by RNA polymerase IV (Pol-IV) target a DNA methyltransferase to its sites of action; here structural and genomic analyses demonstrate that SHH binds chromatin via repressive histone modifications and recruits Pol-IV to enable siRNA production.
- Julie A. Law
- , Jiamu Du
- & Steven E. Jacobsen
-
Letter |
 A conformational switch in HP1 releases auto-inhibition to drive heterochromatin assembly
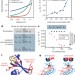
A conformational switch in HP1 releases auto-inhibition to drive heterochromatin assembly
The Schizosaccharomyces pombe HP1 protein, Swi6, is shown to exist in an auto-inhibited state when unbound to chromatin, switching to a spreading-competent state upon binding to the HK9 methyl mark; disrupting this switch affects heterochromatin assembly and gene silencing.
- Daniele Canzio
- , Maofu Liao
- & Geeta J. Narlikar
-
Letter |
 Regulation of ISWI involves inhibitory modules antagonized by nucleosomal epitopes
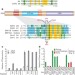
Regulation of ISWI involves inhibitory modules antagonized by nucleosomal epitopes
Two separate regulatory regions on the Drosophila chromatin remodeller ISWI are defined, AutoN and NegC, which negatively regulate ATP hydrolysis and the coupling of ATP hydrolysis to productive DNA translocation, respectively; epitopes on nucleosomes activate ISWI by inhibiting these negative regulatory domains, ensuring that remodelling occurs only in the appropriate chromatin context.
- Cedric R. Clapier
- & Bradley R. Cairns
-
Article |
 DAXX envelops a histone H3.3–H4 dimer for H3.3-specific recognition
DAXX envelops a histone H3.3–H4 dimer for H3.3-specific recognition
The crystal structures of the histone chaperone DAXX histone-binding domain bound to a histone H3.3–H4 dimer are described; DAXX wraps around the H3.3–H4 dimer and alters its conformation.
- Simon J. Elsässer
- , Hongda Huang
- & Dinshaw J. Patel
-
Letter |
 The yeast Fun30 and human SMARCAD1 chromatin remodellers promote DNA end resection
The yeast Fun30 and human SMARCAD1 chromatin remodellers promote DNA end resection
Fun30 and SMARCAD1 are identified as chromatin remodellers that promote DNA end resection during DNA repair and preserve genome stability in yeast and humans, respectively.
- Thomas Costelloe
- , Raphaël Louge
- & Bertrand Llorente
-
Letter |
 The Fun30 nucleosome remodeller promotes resection of DNA double-strand break ends
The Fun30 nucleosome remodeller promotes resection of DNA double-strand break ends
Nucleolytic degradation of 5′ strands at DNA double-strand breaks in yeast is shown to be facilitated by the nucleosome remodeller Fun30, particularly within chromatin bound by the checkpoint adaptor protein known to inhibit resection, Rad9.
- Xuefeng Chen
- , Dandan Cui
- & Grzegorz Ira
-
Article
| Open Access The accessible chromatin landscape of the human genome
The accessible chromatin landscape of the human genome
An extensive map of human DNase I hypersensitive sites, markers of regulatory DNA, in 125 diverse cell and tissue types is described; integration of this information with other ENCODE-generated data sets identifies new relationships between chromatin accessibility, transcription, DNA methylation and regulatory factor occupancy patterns.
- Robert E. Thurman
- , Eric Rynes
- & John A. Stamatoyannopoulos
-
Article |
 Embryonic stem cell potency fluctuates with endogenous retrovirus activity
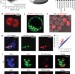
Embryonic stem cell potency fluctuates with endogenous retrovirus activity
A rare cell subpopulation within mouse embryonic stem cell cultures is identified that exhibits properties of two-cell (2C) embryos; the interconversion of ES cells to 2C cells correlates with endogenous retroviral activity.
- Todd S. Macfarlan
- , Wesley D. Gifford
- & Samuel L. Pfaff
-
Article |
 A map of nucleosome positions in yeast at base-pair resolution
A map of nucleosome positions in yeast at base-pair resolution
A new technique for mapping nucleosomes genome-wide with single-base-pair accuracy, by chemical modification of engineered histones.
- Kristin Brogaard
- , Liqun Xi
- & Jonathan Widom
-
Research Highlights |
Cancer gene shifts chromatin
-
News & Views |
 How to duplicate a DNA package
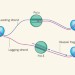
How to duplicate a DNA package
Cells replicate half of their genome as short fragments that are put together later on. The way in which this process is linked to the formation of DNA–protein complexes called nucleosomes is now becoming clearer. See Article p.434
- Alysia Vandenberg
- & Geneviève Almouzni
-
Article |
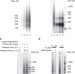 Intrinsic coupling of lagging-strand synthesis to chromatin assembly
Intrinsic coupling of lagging-strand synthesis to chromatin assembly
Genome-wide deep sequencing of Okazaki fragments in S. cerevisae reveals a connection between lagging-strand synthesis and chromatin assembly.
- Duncan J. Smith
- & Iestyn Whitehouse
-
Letter |
 Chromatin-modifying enzymes as modulators of reprogramming
Chromatin-modifying enzymes as modulators of reprogramming
Inhibition of DOT1L, the H3K79 histone methyltransferase, increases cell reprogramming and substituted for KLF4 and c-Myc, showing that chromatin-modifying enzymes act not only as facilitators but also as barriers to reprogramming.
- Tamer T. Onder
- , Nergis Kara
- & George Q. Daley
-
Letter |
 IDH mutation impairs histone demethylation and results in a block to cell differentiation
IDH mutation impairs histone demethylation and results in a block to cell differentiation
Cancer-associated IDH mutants that produce 2-hydroxyglutarate are shown to prevent the histone demethylation that is required for lineage-specific progenitor cells to differentiate into terminally differentiated cells.
- Chao Lu
- , Patrick S. Ward
- & Craig B. Thompson
-
Letter |
 Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma
Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma
Recurrent histone mutations are linked to paediatric glioblastoma multiforme, an aggressive type of brain tumour.
- Jeremy Schwartzentruber
- , Andrey Korshunov
- & Nada Jabado
-
Letter |
 GlcNAcylation of histone H2B facilitates its monoubiquitination
GlcNAcylation of histone H2B facilitates its monoubiquitination
Genome-wide analysis shows that H2B S112 O-linked to N-acetylglucosamine is frequently located near transcribed genes, suggesting that histone GlcNAcylation facilitates transcription of the genes.
- Ryoji Fujiki
- , Waka Hashiba
- & Shigeaki Kato
-
Article |
 BRCA1 tumour suppression occurs via heterochromatin-mediated silencing
BRCA1 tumour suppression occurs via heterochromatin-mediated silencing
- Quan Zhu
- , Gerald M. Pao
- & Inder M. Verma
-
Letter |
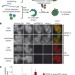 In vitro centromere and kinetochore assembly on defined chromatin templates
In vitro centromere and kinetochore assembly on defined chromatin templates
- Annika Guse
- , Christopher W. Carroll
- & Aaron F. Straight
-
Article
| Open Access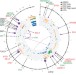 Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma
Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma
- Ryan D. Morin
- , Maria Mendez-Lago
- & Marco A. Marra
-
Letter |
 A Polycomb-based switch underlying quantitative epigenetic memory
A Polycomb-based switch underlying quantitative epigenetic memory
- Andrew Angel
- , Jie Song
- & Martin Howard
-
Letter |
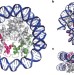 Crystal structure of the human centromeric nucleosome containing CENP-A
Crystal structure of the human centromeric nucleosome containing CENP-A
- Hiroaki Tachiwana
- , Wataru Kagawa
- & Hitoshi Kurumizaka
-
Letter |
 Structure and mechanism of the Swi2/Snf2 remodeller Mot1 in complex with its substrate TBP
Structure and mechanism of the Swi2/Snf2 remodeller Mot1 in complex with its substrate TBP
- Petra Wollmann
- , Sheng Cui
- & Karl-Peter Hopfner
-
Letter |
 Coordination of DNA replication and histone modification by the Rik1–Dos2 complex
Coordination of DNA replication and histone modification by the Rik1–Dos2 complex
- Fei Li
- , Rob Martienssen
- & W. Zacheus Cande
-
Letter |
 Determinants of nucleosome organization in primary human cells
Determinants of nucleosome organization in primary human cells
- Anton Valouev
- , Steven M. Johnson
- & Arend Sidow
-
Article |
 Structure and mechanism of the chromatin remodelling factor ISW1a
Structure and mechanism of the chromatin remodelling factor ISW1a
- Kazuhiro Yamada
- , Timothy D. Frouws
- & Timothy J. Richmond
-
Article |
 Mapping and analysis of chromatin state dynamics in nine human cell types
Mapping and analysis of chromatin state dynamics in nine human cell types
- Jason Ernst
- , Pouya Kheradpour
- & Bradley E. Bernstein
-
Letter |
 A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression
A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression
Long intergenic non-coding RNAs (lincRNAs) have been implicated in both gene silencing and activation, and could be a means for long-range control of gene expression. Here a lincRNA termed HOTTIP is identified at the 5′ tip of the HOXA locus that coordinates the activation of multiple 5′ HOXA genes. Chromosomal looping brings HOTTIP into the proximity of its target genes, where it seems to be required to facilitate histone H3 lysine 4 trimethylation and gene transcription.
- Kevin C. Wang
- , Yul W. Yang
- & Howard Y. Chang
-
Letter |
 Structural basis for recognition of centromere histone variant CenH3 by the chaperone Scm3
Structural basis for recognition of centromere histone variant CenH3 by the chaperone Scm3
- Zheng Zhou
- , Hanqiao Feng
- & Yawen Bai
-
-
Letter |
 The histone variant macroH2A suppresses melanoma progression through regulation of CDK8
The histone variant macroH2A suppresses melanoma progression through regulation of CDK8
The histone variant mH2A is shown to be expressed at reduced levels in many melanomas. Loss of mH2A promotes tumour growth and metastasis via transcriptional upregulation of CDK8, a known oncogene. This study therefore reveals a new tumour suppression mechanism exerted by epigenetic modifications.
- Avnish Kapoor
- , Matthew S. Goldberg
- & Emily Bernstein
-
Letter |
 A unique chromatin signature uncovers early developmental enhancers in humans
A unique chromatin signature uncovers early developmental enhancers in humans
Identifying the genomic regulatory sequences, such as enhancers, that control early embryonic development remains a difficult challenge. Here, profiling of histone modifications and chromatin regulators in human embryonic stem cells (hESCs) reveals unique signatures that are used to identify over 2,000 putative enhancers. These enhancers are either active in the h ESCs or associated with early developmental genes.
- Alvaro Rada-Iglesias
- , Ruchi Bajpai
- & Joanna Wysocka

 Cohesin-dependent globules and heterochromatin shape 3D genome architecture in S. pombe
Cohesin-dependent globules and heterochromatin shape 3D genome architecture in S. pombe The Polycomb complex PRC2 and its mark in life
The Polycomb complex PRC2 and its mark in life