Article |
Featured
-
-
Article |
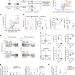 cBAF complex components and MYC cooperate early in CD8+ T cell fate
cBAF complex components and MYC cooperate early in CD8+ T cell fate
cBAF is a negative determinant of memory T cell fate and the manipulation of cBAF early in T cell differentiation can improve cancer immunotherapy.
- Ao Guo
- , Hongling Huang
- & Douglas R. Green
-
Article |
 Structure of human chromatin-remodelling PBAF complex bound to a nucleosome
Structure of human chromatin-remodelling PBAF complex bound to a nucleosome
Cryo-electron microscopy structures of the 12-member PBAF complex provide insights into nucleosome recognition by the complex and the role of mutations in human disease.
- Junjie Yuan
- , Kangjing Chen
- & Zhucheng Chen
-
Article |
 Brahma safeguards canalization of cardiac mesoderm differentiation
Brahma safeguards canalization of cardiac mesoderm differentiation
The BAF chromatin-remodelling complex ATPase gene Brm safeguards cell identity during directed cardiogenesis of mouse embryonic stem cells.
- Swetansu K. Hota
- , Kavitha S. Rao
- & Benoit G. Bruneau
-
Article
| Open Access Targeting SWI/SNF ATPases in enhancer-addicted prostate cancer
Targeting SWI/SNF ATPases in enhancer-addicted prostate cancer
PROTAC degrader–induced SWI/SNF inactivation abolishes DNA accessibility at enhancer elements of oncogenes and also tempers supra-physiologic expression of driver transcription factors, resulting in potent inhibition of tumour growth in mouse models.
- Lanbo Xiao
- , Abhijit Parolia
- & Arul M. Chinnaiyan
-
Article |
 Centromeres are dismantled by foundational meiotic proteins Spo11 and Rec8
Centromeres are dismantled by foundational meiotic proteins Spo11 and Rec8
The meiotic proteins Spo11 and Rec8, which ensure meiotic recombination and reductional chromosome segregation, have additional activities that challenge centromere stability by promoting centromeric nucleosome remodelling in both fission yeast and human cells.
- Haitong Hou
- , Eftychia Kyriacou
- & Julia Promisel Cooper
-
Article |
 Cryo-EM structure of SWI/SNF complex bound to a nucleosome
Cryo-EM structure of SWI/SNF complex bound to a nucleosome
The cryo-electron microscopy structure of the yeast SWI/SNF complex bound to a nucleosome substrate provides insights into the chromatin-remodelling function of this family of protein complexes and suggests mechanisms by which the mutated proteins may cause cancer.
- Yan Han
- , Alexis A Reyes
- & Yuan He
-
Article |
 Structure of SWI/SNF chromatin remodeller RSC bound to a nucleosome
Structure of SWI/SNF chromatin remodeller RSC bound to a nucleosome
The cryo-electron microscopy structure of the 16-subunit yeast SWI/SNF complex RSC in complex with a nucleosome substrate provides insights into the chromatin-remodelling function of this family of protein complexes.
- Felix R. Wagner
- , Christian Dienemann
- & Patrick Cramer
-
Letter |
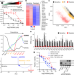 BORIS promotes chromatin regulatory interactions in treatment-resistant cancer cells
BORIS promotes chromatin regulatory interactions in treatment-resistant cancer cells
The CTCF paralogue BORIS is upregulated in transcriptionally reprogrammed neuroblastoma cells rendered resistant to targeted therapy, in which it promotes regulatory chromatin interactions that maintain the resistance phenotype.
- David N. Debruyne
- , Ruben Dries
- & Rani E. George
-
Letter |
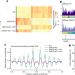 Mammalian ISWI and SWI/SNF selectively mediate binding of distinct transcription factors
Mammalian ISWI and SWI/SNF selectively mediate binding of distinct transcription factors
Genetic deletion of mammalian chromatin remodelling complexes reveals that ISWI and SWI/SNF are required for binding of specific transcription factors and that ISWI regulates nucleosome positioning and nuclear organization in stem cells.
- Darko Barisic
- , Michael B. Stadler
- & Dirk Schübeler
-
Letter |
 Mechanism of DNA translocation underlying chromatin remodelling by Snf2
Mechanism of DNA translocation underlying chromatin remodelling by Snf2
Cryo-EM structures of yeast Snf2 bound to the nucleosome in the presence of ADP or an ATP analogue reveal that Snf2 binding leads to distortion of the DNA, and a two-step mechanism underlying chromatin remodelling by Snf2 is proposed.
- Meijing Li
- , Xian Xia
- & Zhucheng Chen
-
Letter |
 Chromatin analysis in human early development reveals epigenetic transition during ZGA
Chromatin analysis in human early development reveals epigenetic transition during ZGA
By applying an optimized ATAC-seq protocol to human early embryos, distinct accessible chromatin landscapes are found before and after zygotic genome activation, revealing a marked epigenetic transition during zygotic genome activation and putative regulatory elements wiring human early development.
- Jingyi Wu
- , Jiawei Xu
- & Yingpu Sun
-
Letter |
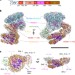 Structure and regulation of the human INO80–nucleosome complex
Structure and regulation of the human INO80–nucleosome complex
Cryo-electron microscopy structure of the human INO80 chromatin remodeller in complex with a bound nucleosome reveals that its motor domains are located at the DNA wrap around the histone core.
- Rafael Ayala
- , Oliver Willhoft
- & Xiaodong Zhang
-
Letter |
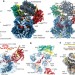 Structural basis for ATP-dependent chromatin remodelling by the INO80 complex
Structural basis for ATP-dependent chromatin remodelling by the INO80 complex
Cryo-electron microscopy structures of the evolutionarily conserved core of a fungal INO80 complex bound to the nucleosomal substrate reveal the mechanism underlying nucleosome sliding and histone editing used by this ATP-dependent chromatin remodeller.
- Sebastian Eustermann
- , Kevin Schall
- & Karl-Peter Hopfner
-
Letter |
 The cis-regulatory dynamics of embryonic development at single-cell resolution
The cis-regulatory dynamics of embryonic development at single-cell resolution
An improved assay for chromatin accessibility at single-cell resolution in Drosophila melanogaster embryos enables identification of developmental-stage- and cell-lineage-specific patterns of chromatin-level transcriptional regulation.
- Darren A. Cusanovich
- , James P. Reddington
- & Eileen E. M. Furlong
-
Letter |
 Nucleosome–Chd1 structure and implications for chromatin remodelling
Nucleosome–Chd1 structure and implications for chromatin remodelling
A cryo-electron microscopy structure of the chromatin remodelling factor Chd1 bound to a nucleosome leads to a model for DNA translocation by its ATPase motor.
- Lucas Farnung
- , Seychelle M. Vos
- & Patrick Cramer
-
Letter |
 ISWI chromatin remodellers sense nucleosome modifications to determine substrate preference
ISWI chromatin remodellers sense nucleosome modifications to determine substrate preference
A high-throughput approach using a DNA-barcoded nucleosome library shows that ISWI chromatin remodellers can distinguish between differently modified nucleosomes.
- Geoffrey P. Dann
- , Glen P. Liszczak
- & Tom W. Muir
-
Article |
 Mechanism of chromatin remodelling revealed by the Snf2-nucleosome structure
Mechanism of chromatin remodelling revealed by the Snf2-nucleosome structure
The cryo-electron microscopy structure of Saccharomyces cerevisiae Snf2 chromatin remodeller bound to a nucleosome and a proposed mechanism for DNA translocation by Snf2 are presented.
- Xiaoyu Liu
- , Meijing Li
- & Zhucheng Chen
-
Letter |
 Structure and regulation of the chromatin remodeller ISWI
Structure and regulation of the chromatin remodeller ISWI
The crystal structures of ISWI, the catalytic subunit of several chromatin remodelling complexes, and its complex with a histone H4 peptide are reported.
- Lijuan Yan
- , Li Wang
- & Zhucheng Chen
-
Letter |
 Broad histone H3K4me3 domains in mouse oocytes modulate maternal-to-zygotic transition
Broad histone H3K4me3 domains in mouse oocytes modulate maternal-to-zygotic transition
Three papers in this issue of Nature use highly sensitive ChIP–seq assays to describe the dynamic patterns of histone modifications during early mouse embryogenesis, showing that oocytes have a distinctive epigenome and providing insights into how the maternal gene expression program transitions to the zygotic program.
- John Arne Dahl
- , Inkyung Jung
- & Arne Klungland
-
Letter |
 Genome-wide nucleosome specificity and function of chromatin remodellers in ES cells
Genome-wide nucleosome specificity and function of chromatin remodellers in ES cells
Genome-wide binding profiles for eight different chromatin remodellers in mouse embryonic stem (ES) cells are determined at single nucleosome resolution; each remodeller binds at specific nucleosome positions relative to the start of genes, and the same remodeller acts as a positive or negative regulator of transcription depending on the promoter chromatin organization and epigenetic marking of the gene it binds.
- Maud de Dieuleveult
- , Kuangyu Yen
- & Matthieu Gérard
-
Article |
 Synaptic, transcriptional and chromatin genes disrupted in autism
Synaptic, transcriptional and chromatin genes disrupted in autism
Whole-exome sequencing in a large autism study identifies over 100 autosomal genes that are likely to affect risk for the disorder; these genes, which show unusual evolutionary constraint against mutations, carry de novo loss-of-function mutations in over 5% of autistic subjects and many function in synaptic, transcriptional and chromatin-remodelling pathways.
- Silvia De Rubeis
- , Xin He
- & Joseph D. Buxbaum
-
Letter |
 Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA
Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA
A nucleosome-spacing mechanism for human ATP-dependent chromatin assembly and remodelling factor (ACF).
- William L. Hwang
- , Sebastian Deindl
- & Xiaowei Zhuang
-
Letter |
 The bromodomain protein Brd4 insulates chromatin from DNA damage signalling
The bromodomain protein Brd4 insulates chromatin from DNA damage signalling
Isoform B of the chromatin-binding protein Brd4 acts to suppress DNA damage response signalling.
- Scott R. Floyd
- , Michael E. Pacold
- & Michael B. Yaffe
-
Letter |
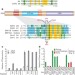 Regulation of ISWI involves inhibitory modules antagonized by nucleosomal epitopes
Regulation of ISWI involves inhibitory modules antagonized by nucleosomal epitopes
Two separate regulatory regions on the Drosophila chromatin remodeller ISWI are defined, AutoN and NegC, which negatively regulate ATP hydrolysis and the coupling of ATP hydrolysis to productive DNA translocation, respectively; epitopes on nucleosomes activate ISWI by inhibiting these negative regulatory domains, ensuring that remodelling occurs only in the appropriate chromatin context.
- Cedric R. Clapier
- & Bradley R. Cairns
-
Letter |
 The yeast Fun30 and human SMARCAD1 chromatin remodellers promote DNA end resection
The yeast Fun30 and human SMARCAD1 chromatin remodellers promote DNA end resection
Fun30 and SMARCAD1 are identified as chromatin remodellers that promote DNA end resection during DNA repair and preserve genome stability in yeast and humans, respectively.
- Thomas Costelloe
- , Raphaël Louge
- & Bertrand Llorente
-
Research Highlights |
Cancer gene shifts chromatin
-
Letter |
 Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma
Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma
Recurrent histone mutations are linked to paediatric glioblastoma multiforme, an aggressive type of brain tumour.
- Jeremy Schwartzentruber
- , Andrey Korshunov
- & Nada Jabado
-
Letter |
 Structure and mechanism of the Swi2/Snf2 remodeller Mot1 in complex with its substrate TBP
Structure and mechanism of the Swi2/Snf2 remodeller Mot1 in complex with its substrate TBP
- Petra Wollmann
- , Sheng Cui
- & Karl-Peter Hopfner
-
Article |
 Structure and mechanism of the chromatin remodelling factor ISW1a
Structure and mechanism of the chromatin remodelling factor ISW1a
- Kazuhiro Yamada
- , Timothy D. Frouws
- & Timothy J. Richmond
-
-
Letter |
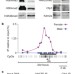 CpG islands influence chromatin structure via the CpG-binding protein Cfp1
CpG islands influence chromatin structure via the CpG-binding protein Cfp1
Most human gene promoters are embedded within CpG islands that lack DNA methylation and coincide with sites at which histone H3 lysine 4 is trimethylated (H3K4me3 sites). Here, a zinc-finger protein, Cfp1, is found to be associated with non-methylated CpG islands and H3K4me3 sites throughout the genome in the mouse brain. A primary function of non-methylated CpG islands might be to genetically determine the local chromatin modification state by interaction with Cfp1 and perhaps other CpG-binding proteins.
- John P. Thomson
- , Peter J. Skene
- & Adrian Bird
-
Letter |
 Chromatin signature of embryonic pluripotency is established during genome activation
Chromatin signature of embryonic pluripotency is established during genome activation
To study the changes in chromatin structure that accompany zygotic genome activation and pluripotency during the maternal–zygotic transition (MZT), the genomic locations of histone H3 modifications and RNA polymerase II have been mapped during this transition in zebrafish embryos. H3 lysine 27 trimethylation and H3 lysine 4 trimethylation are only detected after MZT; evidence is provided that the bivalent chromatin domains found in cultured embryonic stem cells also exist in embryos.
- Nadine L. Vastenhouw
- , Yong Zhang
- & Alexander F. Schier
-

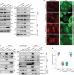 Mitotic bookmarking by SWI/SNF subunits
Mitotic bookmarking by SWI/SNF subunits Brain function and chromatin plasticity
Brain function and chromatin plasticity Chromatin remodelling during development
Chromatin remodelling during development