Article
|
Open Access
Featured
-
-
Article
| Open Access Inhibition of fatty acid oxidation enables heart regeneration in adult mice
Inhibition of fatty acid oxidation enables heart regeneration in adult mice
Inhibition of the fatty acid oxidation metabolic pathway through inactivation of Cpt1b enhances cardiomyocyte survival and proliferation and allows heart regeneration in adult mice.
- Xiang Li
- , Fan Wu
- & Thomas Braun
-
Article |
 Acetyl-methyllysine marks chromatin at active transcription start sites
Acetyl-methyllysine marks chromatin at active transcription start sites
Cellular lysine residues can be both methylated and acetylated on the same sidechain to form Nε-acetyl-Nε-methyllysine (Kacme), which is found on histone H4 across a range of species and across mammalian tissues and is associated with active chromatin.
- William J. Lu-Culligan
- , Leah J. Connor
- & Matthew D. Simon
-
Article
| Open Access Epigenetic dysregulation from chromosomal transit in micronuclei
Epigenetic dysregulation from chromosomal transit in micronuclei
Missegregated chromosomes that are sequestrated in micronuclei are subject to changes in histone modifications leading to abnormalities in chromatin accessibility that remain long after the chromosomes have been reincorporated into the primary nucleus.
- Albert S. Agustinus
- , Duaa Al-Rawi
- & Samuel F. Bakhoum
-
Article |
 Structure of the NuA4 acetyltransferase complex bound to the nucleosome
Structure of the NuA4 acetyltransferase complex bound to the nucleosome
The cryo-electron microscopy structure of NuA4 from Saccharomyces cerevisiae bound to the nucleosome illustrates how NuA4 is assembled and provides mechanistic insights into nucleosome recognition and transcription co-activation by a histone acetyltransferase.
- Keke Qu
- , Kangjing Chen
- & Zhucheng Chen
-
Article
| Open Access Rixosomal RNA degradation contributes to silencing of Polycomb target genes
Rixosomal RNA degradation contributes to silencing of Polycomb target genes
The rixosome associates with Polycomb repressive complexes and chromatin and has a role in silencing of Polycomb target gene expression in human cells via degradation of nascent RNA transcripts.
- Haining Zhou
- , Chad B. Stein
- & Danesh Moazed
-
Article |
 Sex-specific chromatin remodelling safeguards transcription in germ cells
Sex-specific chromatin remodelling safeguards transcription in germ cells
Following global DNA demethylation, mouse gonadal primordial germ cells undergo remodelling of repressive chromatin modifications, resulting in a sex-specific signature that is required to safeguard the transcriptional program.
- Tien-Chi Huang
- , Yi-Fang Wang
- & Petra Hajkova
-
Article |
 Activation of homologous recombination in G1 preserves centromeric integrity
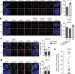
Activation of homologous recombination in G1 preserves centromeric integrity
Centromeres are able to recruit the homologous recombination machinery during G1 via CENP-A and HJURP, thereby preserving centromeric integrity even in the absence of a sister chromatid.
- Duygu Yilmaz
- , Audrey Furst
- & Evi Soutoglou
-
Article |
 UTX condensation underlies its tumour-suppressive activity
UTX condensation underlies its tumour-suppressive activity
Phase separation properties are a major determinant of UTX activity in chromatin regulation in tumour suppression, and are dependent on a core intrinsically disordered region of the protein.
- Bi Shi
- , Wei Li
- & Hao Jiang
-
Article |
 Elevated NSD3 histone methylation activity drives squamous cell lung cancer
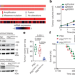
Elevated NSD3 histone methylation activity drives squamous cell lung cancer
The histone H3K36 methyltransferase NSD3, which is associated with the common 8p11–12 chromosomal amplification, is an oncogenic driver in lung squamous cell carcinoma.
- Gang Yuan
- , Natasha M. Flores
- & Or Gozani
-
Article |
 METTL3 regulates heterochromatin in mouse embryonic stem cells
METTL3 regulates heterochromatin in mouse embryonic stem cells
Binding of METTL3 to chromatin is enriched over IAP family endogenous retroviral elements in mouse embryonic stem cells, helping to ensure the integrity of heterochromatin at these elements.
- Wenqi Xu
- , Jiahui Li
- & Hongjie Shen
-
Article |
 Molecular basis of nucleosomal H3K36 methylation by NSD methyltransferases
Molecular basis of nucleosomal H3K36 methylation by NSD methyltransferases
Cryo-electron microscopy structures of the nucleosome-bound NSD2 and NSD3 histone methyltransferases reveal the molecular basis of their histone modification activity, and show how mutations in these proteins can lead to oncogenesis.
- Wanqiu Li
- , Wei Tian
- & Zhanxin Wang
-
Article |
 H1 histones control the epigenetic landscape by local chromatin compaction
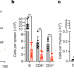
H1 histones control the epigenetic landscape by local chromatin compaction
Experiments using a conditional triple-knockout mouse strain show that histone H1 regulates the activity of chromatin domains by controlling chromatin compaction, genome architecture and histone methylation.
- Michael A. Willcockson
- , Sean E. Healton
- & Arthur I. Skoultchi
-
Article |
 Epigenetic gene silencing by heterochromatin primes fungal resistance
Epigenetic gene silencing by heterochromatin primes fungal resistance
Fission yeast grown in sublethal levels of caffeine develop heterochromatin-dependent epimutations conferring unstable heritable gene silencing that conveys resistance to caffeine, while remaining genetically wild type.
- Sito Torres-Garcia
- , Imtiyaz Yaseen
- & Robin C. Allshire
-
Article |
 Histone H3.3 phosphorylation amplifies stimulation-induced transcription
Histone H3.3 phosphorylation amplifies stimulation-induced transcription
The histone variant H3.3 is phosphorylated at Ser31 in induced genes, and this selective mark stimulates the histone methyltransferase SETD2 and ejects the ZMYND11 repressor, thus revealing a role for histone phosphorylation in amplifying de novo transcription.
- Anja Armache
- , Shuang Yang
- & Steven Z. Josefowicz
-
Article |
 Phase separation directs ubiquitination of gene-body nucleosomes
Phase separation directs ubiquitination of gene-body nucleosomes
The yeast E3 ligase Bre1 forms a core–shell condensate with the scaffold protein Lge1, implicating liquid–liquid phase separation as a mechanism in the ubiquitination of histone H2B along gene bodies.
- Laura D. Gallego
- , Maren Schneider
- & Alwin Köhler
-
Article |
 Structure of the transcription coactivator SAGA
Structure of the transcription coactivator SAGA
Structural studies on the yeast transcription coactivator complex SAGA (Spt–Ada–Gcn5–acetyltransferase) provide insights into the mechanism of initiation of regulated transcription by this multiprotein complex, which is conserved among eukaryotes.
- Haibo Wang
- , Christian Dienemann
- & Patrick Cramer
-
Article |
 Metabolic regulation of gene expression by histone lactylation
Metabolic regulation of gene expression by histone lactylation
The lactylation of lysine residues on histones in mammalian cells is stimulated by hypoxia and bacterial challenges, and increased histone lactylation induces genes involved in wound healing.
- Di Zhang
- , Zhanyun Tang
- & Yingming Zhao
-
Letter |
 Structural basis of nucleosome recognition and modification by MLL methyltransferases
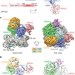
Structural basis of nucleosome recognition and modification by MLL methyltransferases
Cryo-electron microscopy structures of mixed-lineage leukaemia methyltransferases 1 and 3 associated with unmodified or mono-ubiquitinated nucleosome reveal the structural basis for the activity specificity and regulation of these enzymes.
- Han Xue
- , Tonghui Yao
- & Jing Huang
-
Letter |
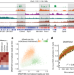 The histone mark H3K36me2 recruits DNMT3A and shapes the intergenic DNA methylation landscape
The histone mark H3K36me2 recruits DNMT3A and shapes the intergenic DNA methylation landscape
H3K36me2 targets DNMT3A to intergenic regions and this process, together with H3K36me3-mediated recruitment of DNMT3B, has a key role in establishing and maintaining genomic DNA methylation landscapes.
- Daniel N. Weinberg
- , Simon Papillon-Cavanagh
- & Chao Lu
-
Letter |
 Active chromatin marks drive spatial sequestration of heterochromatin in C. elegans nuclei
Active chromatin marks drive spatial sequestration of heterochromatin in C. elegans nuclei
MRG-1 indirectly promotes anchoring of chromatin in differentiated intestinal cells in Caenorhabditis elegans by sequestering the histone acetyltransferase CBP-1/p300.
- Daphne S. Cabianca
- , Celia Muñoz-Jiménez
- & Susan M. Gasser
-
Article |
 Transcription factor dimerization activates the p300 acetyltransferase
Transcription factor dimerization activates the p300 acetyltransferase
The activation of the histone acetyltransferase p300 depends on the activation and oligomerization status of transcription factor ligands.
- Esther Ortega
- , Srinivasan Rengachari
- & Daniel Panne
-
Letter |
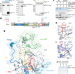 Automethylation-induced conformational switch in Clr4 (Suv39h) maintains epigenetic stability
Automethylation-induced conformational switch in Clr4 (Suv39h) maintains epigenetic stability
An autoinhibitory conformation of the histone H3K9 methyltransferase Clr4 of Schizosaccharomyces pombe helps to prevent aberrant heterochromatin formation and maintains epigenetic stability.
- Nahid Iglesias
- , Mark A. Currie
- & Danesh Moazed
-
Letter |
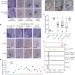 Embryonic epigenetic reprogramming by a pioneer transcription factor in plants
Embryonic epigenetic reprogramming by a pioneer transcription factor in plants
The seed-specific transcription factor LEC1 promotes an active chromatin state at the floral repressor FLC and activates its expression in the Arabidopsis pro-embryo, thus reversing the winter cold-induced silenced state that is inherited from gametes.
- Zeng Tao
- , Lisha Shen
- & Yuehui He
-
Letter |
 Polycomb-like proteins link the PRC2 complex to CpG islands
Polycomb-like proteins link the PRC2 complex to CpG islands
Crystal structures of the Polycomb-like proteins PHF1 and MTF2 with bound DNA and histone peptides show that extended homologous regions of the two proteins form a winged-helix structure that has an unexpected mechanism of binding to unmethylated CpG-containing DNA motifs.
- Haojie Li
- , Robert Liefke
- & Zhanxin Wang
-
Letter |
 Unique roles for histone H3K9me states in RNAi and heritable silencing of transcription
Unique roles for histone H3K9me states in RNAi and heritable silencing of transcription
Heterochromatin formation involves histone H3 methylation, with H3K9me2 defining a distinct heterochromatin state that is transcriptionally permissive and can couple with RNAi, and the transition to non-permissive H3K9me3 required for the epigenetic heritability of heterochromatin.
- Gloria Jih
- , Nahid Iglesias
- & Danesh Moazed
-
Letter |
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
Three papers in this issue of Nature use highly sensitive ChIP–seq assays to describe the dynamic patterns of histone modifications during early mouse embryogenesis, showing that oocytes have a distinctive epigenome and providing insights into how the maternal gene expression program transitions to the zygotic program.
- Bingjie Zhang
- , Hui Zheng
- & Wei Xie
-
Letter |
 Distinct features of H3K4me3 and H3K27me3 chromatin domains in pre-implantation embryos
Distinct features of H3K4me3 and H3K27me3 chromatin domains in pre-implantation embryos
Three papers in this issue of Nature use highly sensitive ChIP–seq assays to describe the dynamic patterns of histone modifications during early mouse embryogenesis, showing that oocytes have a distinctive epigenome and providing insights into how the maternal gene expression program transitions to the zygotic program.
- Xiaoyu Liu
- , Chenfei Wang
- & Shaorong Gao
-
Letter |
 The structural basis of modified nucleosome recognition by 53BP1
The structural basis of modified nucleosome recognition by 53BP1
A cryo-electron microscopy structure of the DNA damage repair protein 53BP1 bound to a nucleosome illuminates the way 53BP1 recognizes two types of histone modifications (a methyl group and a ubiquitin moiety), and provides insight into the highly specified recognition and recruitment of 53BP1 to modified chromatin.
- Marcus D. Wilson
- , Samir Benlekbir
- & Daniel Durocher
-
Letter |
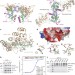 H4K20me0 marks post-replicative chromatin and recruits the TONSL–MMS22L DNA repair complex
H4K20me0 marks post-replicative chromatin and recruits the TONSL–MMS22L DNA repair complex
We have a limited understanding of how cells mark and identify newly replicated genomic loci that have a sister chromatid; here, unmethylated K20 in the tail of new histone H4 is shown to serve as a signature of post-replicative chromatin, which is specifically recognized by the homologous recombination complex TONSL–MMS22L.
- Giulia Saredi
- , Hongda Huang
- & Anja Groth
-
Article |
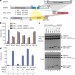 Structural basis for activity regulation of MLL family methyltransferases
Structural basis for activity regulation of MLL family methyltransferases
Crystal structures of the SET domains of MLL3 and a mutant MLL1 either unbound or complexed with domains from RBBP5 and ASH2L are determined; a combination of structural, biochemical and computational analyses reveals a two-step activation mechanism of MLL family proteins, which may be relevant for other histone methyltransferases.
- Yanjing Li
- , Jianming Han
- & Ming Lei
-
Letter |
 Histone H1 couples initiation and amplification of ubiquitin signalling after DNA damage
Histone H1 couples initiation and amplification of ubiquitin signalling after DNA damage
At the initiation of DNA double-strand break repair, a number of ubiquitylation events occur; here, the RNF8 ubiquitin E3 ligase and the ubiquitin-conjugating E2 enzyme, UBC13, are shown to primarily modify H1-type linker histones, via a K63 linkage.
- Tina Thorslund
- , Anita Ripplinger
- & Niels Mailand
-
Article |
 Gain-of-function p53 mutants co-opt chromatin pathways to drive cancer growth
Gain-of-function p53 mutants co-opt chromatin pathways to drive cancer growth
A ChIP-seq analysis of the DNA-binding properties of mutant gain-of-function p53 protein compared to wild-type p53 reveals the gain-of-function proteins bind to and activate a distinct set of genes including chromatin modifying enzymes such as the histone methyltransferase MLL; small molecular inhibitors of MLL function may represent a new target for cancers with mutant p53.
- Jiajun Zhu
- , Morgan A. Sammons
- & Shelley L. Berger
-
Letter |
 Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation
Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation
Genome-wide localization and activity analysis of the de novo DNA methyltransferases DNMT3A and DNMT3B in mouse embryonic stem cells identifies overlapping and individual targeting preferences to the genome, including a role for DNMT3B in gene body methylation.
- Tuncay Baubec
- , Daniele F. Colombo
- & Dirk Schübeler
-
Letter |
 Tyrosine phosphorylation of histone H2A by CK2 regulates transcriptional elongation
Tyrosine phosphorylation of histone H2A by CK2 regulates transcriptional elongation
A conserved tyrosine residue, Tyr 57, of histone H2A is phosphorylated by an unsuspected tyrosine kinase activity of casein kinase 2, influencing a series of histone marks associated with active transcription and regulating transcription elongation.
- Harihar Basnet
- , Xue B. Su
- & Michael G. Rosenfeld
-
Letter |
 Contrasting roles of histone 3 lysine 27 demethylases in acute lymphoblastic leukaemia
Contrasting roles of histone 3 lysine 27 demethylases in acute lymphoblastic leukaemia
T-cell acute lymphoblastic leukaemia (T-ALL) is a haematological malignancy with a poor prognosis and no available targeted therapies; now two histone H3 lysine 27 demethylases, JMJD3 and UTX, are shown to have contrasting roles in human T-ALL cells and a mouse model of the disease, and a small molecule demethylase inhibitor is found to inhibit the growth of T-ALL cell lines, introducing a potential therapeutic avenue for acute leukaemia.
- Panagiotis Ntziachristos
- , Aristotelis Tsirigos
- & Iannis Aifantis
-
Letter |
 Selective transcriptional regulation by Myc in cellular growth control and lymphomagenesis
Selective transcriptional regulation by Myc in cellular growth control and lymphomagenesis
Global transcriptional and epigenomic analyses in diverse cell types reveal that the primary action of Myc is to up- and downregulate transcription of distinct groups of genes, rather than to amplify transcription of all active genes; general RNA amplification, when observed, is better explained as an indirect consequence of Myc’s action on cellular physiology.
- Arianna Sabò
- , Theresia R. Kress
- & Bruno Amati
-
Letter |
 ZMYND11 links histone H3.3K36me3 to transcription elongation and tumour suppression
ZMYND11 links histone H3.3K36me3 to transcription elongation and tumour suppression
Candidate tumour suppressor ZMYND11 specifically recognizes histone K36 trimethylation on the histone variant H3.3 and helps regulate transcription elongation.
- Hong Wen
- , Yuanyuan Li
- & Xiaobing Shi
-
Letter |
 Citrullination regulates pluripotency and histone H1 binding to chromatin
Citrullination regulates pluripotency and histone H1 binding to chromatin
This study shows that PADI4-mediated citrullination occurs during pluripotency and that citrullination of H1 results in loosening of chromatin compaction; furthermore, citrullination is shown to be important for the activation of stem-cell genes, for iPS cell reprogramming and to maintain pluripotent cells in the early mouse embryo.
- Maria A. Christophorou
- , Gonçalo Castelo-Branco
- & Tony Kouzarides
-
Letter |
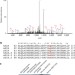 Glutamine methylation in histone H2A is an RNA-polymerase-I-dedicated modification
Glutamine methylation in histone H2A is an RNA-polymerase-I-dedicated modification
A description of a new histone modification, methylation of glutamine, on histone H2A in yeast and human cells.
- Peter Tessarz
- , Helena Santos-Rosa
- & Tony Kouzarides
-
Letter |
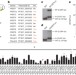 PfSETvs methylation of histone H3K36 represses virulence genes in Plasmodium falciparum
PfSETvs methylation of histone H3K36 represses virulence genes in Plasmodium falciparum
The malaria parasite Plasmodium falciparum escapes immune detection by expressing one of 60 antigenically distinct var genes at any one time during the course of infection: here it is shown that the P. falciparum protein PfSETvs has a key role in var gene silencing through the trimethylation of histone H3K36.
- Lubin Jiang
- , Jianbing Mu
- & Louis H. Miller
-
Article |
 53BP1 is a reader of the DNA-damage-induced H2A Lys 15 ubiquitin mark
53BP1 is a reader of the DNA-damage-induced H2A Lys 15 ubiquitin mark
This study shows that 53BP1 recruitment to sites of DNA damage involves dual recognition of H4K20me2 and H2AK15 histone ubiquitination; the ubiquitin mark and the surrounding epitope on H2A are read by a region of 53BP1 designated the ubiquitination-dependent recruitment motif.
- Amélie Fradet-Turcotte
- , Marella D. Canny
- & Daniel Durocher
-
Letter |
 Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1
Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1
In Arabidopsis, RNA-directed DNA methylation is a poorly understood gene silencing pathway in which small interfering RNAs generated by RNA polymerase IV (Pol-IV) target a DNA methyltransferase to its sites of action; here structural and genomic analyses demonstrate that SHH binds chromatin via repressive histone modifications and recruits Pol-IV to enable siRNA production.
- Julie A. Law
- , Jiamu Du
- & Steven E. Jacobsen
-
Article |
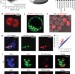 Embryonic stem cell potency fluctuates with endogenous retrovirus activity
Embryonic stem cell potency fluctuates with endogenous retrovirus activity
A rare cell subpopulation within mouse embryonic stem cell cultures is identified that exhibits properties of two-cell (2C) embryos; the interconversion of ES cells to 2C cells correlates with endogenous retroviral activity.
- Todd S. Macfarlan
- , Wesley D. Gifford
- & Samuel L. Pfaff
-
Letter |
 IDH mutation impairs histone demethylation and results in a block to cell differentiation
IDH mutation impairs histone demethylation and results in a block to cell differentiation
Cancer-associated IDH mutants that produce 2-hydroxyglutarate are shown to prevent the histone demethylation that is required for lineage-specific progenitor cells to differentiate into terminally differentiated cells.
- Chao Lu
- , Patrick S. Ward
- & Craig B. Thompson
-
Letter |
 GlcNAcylation of histone H2B facilitates its monoubiquitination
GlcNAcylation of histone H2B facilitates its monoubiquitination
Genome-wide analysis shows that H2B S112 O-linked to N-acetylglucosamine is frequently located near transcribed genes, suggesting that histone GlcNAcylation facilitates transcription of the genes.
- Ryoji Fujiki
- , Waka Hashiba
- & Shigeaki Kato
-
Article
| Open Access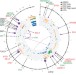 Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma
Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma
- Ryan D. Morin
- , Maria Mendez-Lago
- & Marco A. Marra
-
Letter |
 A Polycomb-based switch underlying quantitative epigenetic memory
A Polycomb-based switch underlying quantitative epigenetic memory
- Andrew Angel
- , Jie Song
- & Martin Howard
-
Letter |
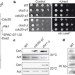 Coordination of DNA replication and histone modification by the Rik1–Dos2 complex
Coordination of DNA replication and histone modification by the Rik1–Dos2 complex
- Fei Li
- , Rob Martienssen
- & W. Zacheus Cande
-
Letter |
 Mechanism and regulation of acetylated histone binding by the tandem PHD finger of DPF3b
Mechanism and regulation of acetylated histone binding by the tandem PHD finger of DPF3b
The lysine residues of histone proteins can be acetylated or methylated, with important effects on gene expression. Until recently the protein modules that bind acetyl-lysine have been limited to bromodomains. However, the tandem plant homeodomain (PHD) finger of human DPF3b — which is involved in gene activation — has also been reported to bind to acetylated histones. Here, three-dimensional solution structures of DPF3b offer mechanistic insight into how this protein recognizes acetylation marks.
- Lei Zeng
- , Qiang Zhang
- & Ming-Ming Zhou

 Decoding chromatin states by proteomic profiling of nucleosome readers
Decoding chromatin states by proteomic profiling of nucleosome readers