« Prev Next »
Two early developmental processes — oocyte (egg) maturation and early embryonic development — present excellent opportunities to study the cell division cycle and cell proliferation control. Primary oocytes undergo meiosis to produce matured oocytes. Then, upon fertilization, the eggs divide at incredible speeds by alternating DNA synthesis and mitosis without noticeable G1 or G2 phases. During these rapid cell divisions, the cell does not pause to transcribe mRNA. Instead, new proteins that are necessary to support this massive growth phase are translated solely from the stored maternal mRNAs. For example, soon after fertilization of sea urchin oocytes, the rate of protein synthesis increases by 10–30 fold. Similarly, in mouse, frog, clam, and several other organisms, the pattern of protein synthesis also changes dramatically. What is the mechanism in the egg — or for that matter any cell — that distinguishes between mRNA that should be translated right away and mRNA that needs to wait for translation?
What Keeps Maternal mRNAs Dormant and What Activates Them?
In the 1960s and 1970s, scientists asked this question and came up with two possible answers. In 1966, Alexander Spirin proposed that the dormant maternal mRNAs remained obscured for translation by "masking" proteins that prevented mRNA translation. The masking hypothesis also suggested that upon stimulation such as fertilization, the masking proteins were removed from the dormant mRNAs so that they were incorporated into polysomes (poly-ribosomes) for active translation.
An alternative hypothesis dealt with the change of maternal mRNAs themselves. In the late 1960s through early 1970s, results from several groups indicated that some cellular RNAs from mouse cells or some viral RNAs from virus-infected human cells contain polyadenylic acid sequences (stretches of adenine nucleotides) that allow them to bind to cellulose bound with oligo-thymidine (since adenosine can pair with thymidines). These polyadenylic acid sequences, called poly(A) tails, were longer in the nucleus and slightly shorter in the cytoplasm, and were measured after digestion with ribonucleases A and T1.
But how does the longer mRNA get out of the nucleus and why is it shortened right after it exits? James Darnell and associates used a pulse or continuous labeling technique with radioactive phosphate or adenosine and discovered that heterogeneous nuclear RNA (hnRNA) was the precursor of cytoplasmic mRNA and that the poly(A), which was attached to the end of the mRNA after transcription, helped export the mRNA to the cytoplasm (Darnell et al. 1971). Once the mRNA got into the cytoplasmic space, the shorter poly(A) tail was not sufficient to permit protein translation. It turns out that the length of the poly(A) tails in the cytoplasm is extremely important.
Lengthening the mRNA Poly(A) Tails in Sea Urchin Oocytes
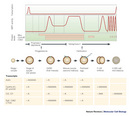
![A line graph shows the changes in polyadenylation of the MRNA in sea urchins as embryogenesis proceeds from an unfertilized egg to an embryo with four cells. The level of polyadenylic acid [Poly(A)] increases after fertilization, between the single cell and the two cell stages.](/scitable/content/ne0000/ne0000/ne0000/ne0000/14263635/slater_240333a0_f3_2_1.jpg)
Another scenario would have been that the mRNA poly(A) tail was fully synthesized prior to nuclear export and was shortened in the cytoplasm. This was actually confirmed in later experiments. Nonetheless, Slater's conclusion that the poly(A) tails were elongated by polyadenylation in the cytoplasm of the oocytes after fertilization was still valid and intriguing and set further experiments on the right path.
Poly(A) Length and Translatability of the mRNA: A Clam Story
The low resolution of the assay and the measurement in bulk were somewhat limited for drawing firm conclusions, so the results were controversial. Rosenthal et al. decided to use gene-specific probes to examine individual RNA species that were isolated from oocytes and early embryos in Spisula (commonly known as triangle shells or surf clams) (Rosenthal et al. 1983). By comparing proteins, which were labeled in oocytes and early embryos with 35S-methionine, Rosenthal's group discovered that several proteins only became highly expressed after fertilization. They identified the cDNAs for two such proteins (called A and C) by "hybrid-selection" to probe the RNA isolated from oocytes and embryos. They separated each RNA preparation using oligo-thymidine cellulose into two pools: the bound fraction containing RNA with polyA longer than 15-20 adenines and the unbound fraction contained RNA with no or very short poly(A). The poly(A) and non-poly(A) fractions were then electrophoresed on agarose gels and probed with protein A cDNA labeled with 32P. They found that the mRNA for protein A was in the non-poly(A) fraction of oocytes but it was in the poly(A) fraction of embryos. Furthermore, the protein A mRNA in the poly(A) faction was longer than that in the non-poly(A) fraction, and the length difference disappeared when poly(A) sequences were digested away with RNase H and oligo-thymidines. To verify which RNA was being translated in vivo, they sedimented cytoplasmic lysates through sucrose gradients and probed each fraction for the presence of protein A mRNA. They found that, whereas Protein A mRNA in the oocyte lysate was not associated with a polysome, the longer polyadenylated mRNA in the embryo lysate (2-cell stage, about 18 hours after fertilization) was polysome associated. The mRNA for protein C behaved the same way as protein A mRNA in aforementioned assays. Thus, they discovered a strong correlation between polyA tail lengthening and translatability.
Interestingly, they also observed deadenlyation of mRNA encoding alpha-tubulin — it had a long poly(A) tail and was translated in the oocytes, but the poly(A) tail was removed, and the mRNA was not associated with polysomes after fertilization. The study provided strong molecular evidence that fertilization of oocytes was accompanied by cytoplasmic polyadenylation and deadenylation of mRNA that determined the length of the poly(A) tail, and this in turn correlated with mRNA translatability.
Studies from Vertebrates and Mammals
The Controllers-RNA Elements and Binding Proteins
What molecular entities were involved in the regulation of the polyadenylation in the cytoplasm of oocytes? Joel Richter and his colleagues have made some very interesting observations by studying Xenopus oocyte and embryonic development. They found that several mRNAs that were polyadenylated in the cytoplasm of maturing oocytes contained a uridine-rich sequence (like UUUUUAU), which was essential for poly(A) elongation and translational activation. They called this sequence CPE for cytoplasmic polyadenylation element, and predicted that CPE would bind proteins involved in regulating polyadenylation.
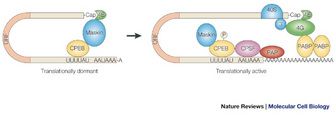
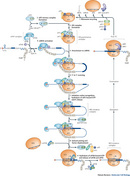
Subsequent research then detailed how CPEB was itself regulated. CPE is phosphorylated differentially during development, which alters the ability of CPE to interact with its protein partners and affects the protein complexes assembled near the mRNA 3' end. These 3' end protein complexes influence not only polyadenylation but also deadenylation and translation activation (Mendez & Richter 2001).
What Happens to the End Affects the Beginning
Several experiments using the yeast PAB1 gene, which encodes the poly(A) binding protein (PABP), supported this idea. For example, Sachs and Davis showed that depleting or altering PABP caused a reduced translation rate in yeast (Sachs & Davis 1989). The size of polysomes was not significantly changed but the amount of polysomes was dramatically reduced. They screened for additional mutations that would rescue the PAB1 mutant, and they isolated extragenic suppressors that mapped to genes with roles in the 60S ribosome function. Their study supported the notion that PABP bound to the poly(A) tail and enhanced translation of that mRNA. We now know that PABP interacts with eIF-4G and other associated translation proteins, and this interaction facilitates translation initiation at the mRNA 5' end (Figure 4).
Correlation Does Not Mean Causation
How much does polyadenylation of maternal mRNA contribute to the dramatic changes in protein synthesis in oocytes? Many studies demonstrate a strong correlation between poly(A) tail lengthening and protein synthesis, also known as polysome association. However, a high correlation between two observations is not sufficient to conclude that one causes the other. One also needs to consider the other variables (for example, two unrelated mechanisms could elicit a common molecular response without affecting each other directly) and the confounding effects. For example, in sea urchin, protein synthesis increased immediately after fertilization but the bulk of poly(A) lengthening occurred 30 to 90 minutes later. Some studies also indicate that the translation machinery itself is stimulated after fertilization. Nonetheless, a tremendous amount of literature depicts a picture that the poly(A) size control is critical for oocyte and embryonic development (Radford et al. 2008).
Summary
References and Recommended Reading
Darnell, J. E. et al. Polyadenylic acid sequences: role in conversion of nuclear RNA into messenger RNA. Science 174, 507–510 (1971).
Hake, L. E. & Richter, J. D. CPEB is a specificity factor that mediates cytoplasmic polyadenylation during Xenopus oocyte maturation. Cell 79, 617–627 (1994).
Mendez, R. & Richter, J. D. Translational control by CPEB: a means to the end. Nature Reviews Mol Cell Biol 2, 521–529 (2001).
Radford, H. E., Meijer, H.A. & de Moor, C.H. Translational control by cytoplasmic polyadenylation in Xenopus oocytes. Biochimica et biophysica acta 1779, 217–229 (2008).
Rosenthal, E. T., Tansey, T. R. & Ruderman, J. V. Sequence-specific adenylations and deadenylations accompany changes in the translation of maternal messenger RNA after fertilization of Spisula oocytes. Journal of molecular biology 166, 309–327 (1983).
Sachs, A. B. & Davis, R. W. The poly(A) binding protein is required for poly(A) shortening and 60S ribosomal subunit-dependent translation initiation. Cell 58, 857–867 (1989).
Slater, D.W., Slater, I. & Gillespie, D. Post-fertilization synthesis of polyadenylic acid in sea urchin embryos. Nature 240, 333–337 (1972).
Strickland, S. et al. Antisense RNA directed against the 3' noncoding region prevents dormant mRNA activation in mouse oocytes. Science 241, 680–684 (1988).



 Figure 1: Changes of Poly(A) titers after fertilization
Figure 1: Changes of Poly(A) titers after fertilization