
Examples of using SSD, an object-detection algorithm described in a highly cited artificial intelligence paper.
Credit: Wei Liu et al. European Conference on Computer Vision (2016)
Google Scholar reveals its most influential papers for 2021
Early clinical observations of COVID-19 and its mortality risk factors among the most cited output, while a five-year-old AI paper continues to command attention.
24 August 2021

Wei Liu et al. European Conference on Computer Vision (2016)
Examples of using SSD, an object-detection algorithm described in a highly cited artificial intelligence paper.
COVID-19-related papers have eclipsed artificial intelligence research in the annual listing of the most highly-cited publications in the Google Scholar database. The most highly cited COVID-19 paper, published in The Lancet in early 2020, has garnered more than 30,000 citations to date (see below for paper summary).
But, in the database of almost 400 million academic papers and other scholarly literature, even it fell a long way short of the most highly cited paper of the last five years, ‘Deep Residual Learning for Image Recognition’, published in Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition by a team from Microsoft in 2016.
The five-year-old paper’s astonishing ascendancy continues, from 25,256 citations in 2019 to 49,301 citations in 2020 to 82,588 citations in 2021. We wrote about it last year here.
The 2021 Google Scholar Metrics ranking tracks papers published between 2016 and 2020, and includes citations from all articles that were indexed in Google Scholar as of July 2020. Google Scholar is the largest database in the world of its kind.
Below we describe selections from Google Scholar’s most highly-cited articles for 2021. COVID-19 research dominated new arrivals in the list, but we’re also featuring a popular AI paper from 2016, and research that provides an economical shortcut to seeing patterns of human genetic variation, also from 2016.
See our coverage of the 2019 and 2020 lists.
‘Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China’
30,529 citations
Published in February 2020, this is one of the earliest papers to describe the clinical characteristics of COVID-19. It was authored by researchers in China and doctors working in hospitals in Wuhan, the city where COVID-19 was first detected in late 2019.
The team, from institutions such as the Jin Yin-tan Hospital in Wuhan and China-Japan Friendship Hospital in Beijing, reviewed the clinical and nursing reports, chest X-rays and lab results of the first 41 COVID-19 patients. They noted that the novel virus acts similarly to SARS and MERS, in that it causes pneumonia, but is different in that it seldom manifests as a runny nose or intestinal symptoms.
The final sentences of the paper call for robust and rapid testing, because of the likelihood of the disease spreading out of control:
“Reliable quick pathogen tests and feasible differential diagnosis based on clinical description are crucial for clinicians in their first contact with suspected patients. Because of the pandemic potential of 2019-nCoV, careful surveillance is essential to monitor its future host adaption, viral evolution, infectivity, transmissibility, and pathogenicity.”
The paper has been referenced or cited in almost 100 policy documents to date, including several released by the World Health Organization on topics such as mask-wearing and clinical care of patients with severe symptoms.
‘Clinical Characteristics of Coronavirus Disease 2019 in China’
New England Journal of Medicine
19,656 citations
Published online in February 2020, this study was a retrospective review of medical records for 1,099 COVID-19 cases reported to the National Health Commission of the People's Republic of China between 11 December 2019 and 29 January 2020.
The team, which included almost 40 researchers from China from institutions such as the Guangzhou Medical University in Guangzhou and Wuhan Jinyintan Hospital in Wuhan, accessed electronic medical records from 552 hospitals in mainland China to summarise exposure risk, signs and symptoms, laboratory and radiologic findings related to COVID-19 infection.
The study garnered a lot of media attention based on the evidence it put forward that men might be more severely impacted by disease – 58% of the patient cohort were male.
However, as Sharon Begley reported for STAT, “It’s possible the apparent sex imbalance reflects patterns of travel and contacts that make men more likely to be exposed to carriers of the virus, not any inherent biological differences. It’s also possible the apparent worse disease severity in men could skew the data.”
A paper published in JAMA around the same time by researchers in the United States reported that, among hospitalized patients, there is “a slight predominance of men”.
A Nature Communications meta-analysis, published in December 2020, looked at 92 studies covering more than three million patients and concluded that, while males and females appeared to be susceptible to infection, men were 2.84 times more likely to be end up in intensive care and 1.39 times more likely to die from the disease.
‘Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study’
17,047 citations
Published in March 2020, The Lancet described this study as the first time researchers have examined risk factors associated with severe symptoms and death in hospitalised or deceased patients. Of the 191 patients studied, 137 were discharged from hospital and 54 died.
The study, by researchers from hospitals in China, also presented new data on viral shedding – information that informed early understanding of how the virus spreads and can be detected over the cause of infection.
“The extended viral shedding noted in our study has important implications for guiding decisions around isolation precautions and antiviral treatment in patients with confirmed COVID-19 infection,” said co-lead author, Bin Cao, from the China-Japan Friendship Hospital and Capital Medical University in Beijing.
“However, we need to be clear that viral shedding time should not be confused with other self-isolation guidance for people who may have been exposed to COVID-19 but do not have symptoms, as this guidance is based on the incubation time of the virus.”
‘A Novel Coronavirus from Patients with Pneumonia in China, 2019’
The New England journal of medicine
16,194 citations
On 31 December 2019, the Chinese Center for Disease Control and Prevention (China CDC) dispatched a rapid response team to accompany health authorities in Hubei province and Wuhan city in conducting COVID-19 investigations.
This study, published in January 2020, reported the results of that investigation, including the clinical features of the pneumonia of two patients.
Described by Jose Manuel Jimenez-Guardeño, a researcher in the Department of Infectious Diseases at King's College London, UK and colleagues in an article for The Conversation as “the article that released this virus to the world”, the paper details how the virus was isolated from patients with pneumonia in Wuhan in cell cultures.
“In fact, actual photographs of SARS-CoV-2 were shown to the world for the first time here,” say Jimenez-Guardeño and his co-authors.
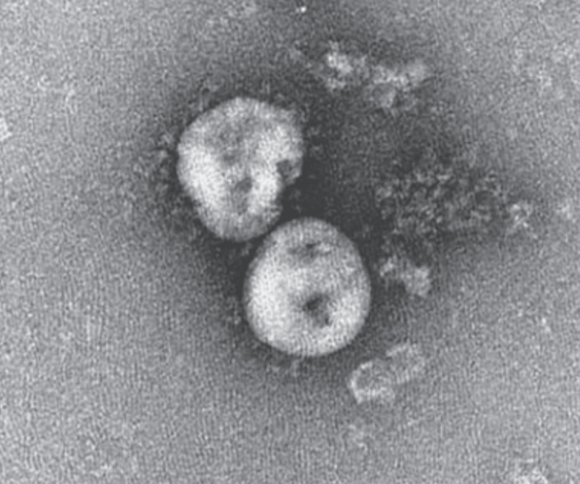 SARS-CoV-2 viral particles, Na Zhu et al., 2020.
SARS-CoV-2 viral particles, Na Zhu et al., 2020.
The study authors urged that more epidemiologic investigations were needed in order to characterize transmission modes, reproduction intervals and other characteristics of the virus to inform strategies to control and stop its spread.
‘SSD: Single Shot MultiBox Detector’
European Conference on Computer Vision
15,368 citations
A change of pace from recent COVID-19 studies, this paper, led by Wei Liu from the University of North Carolina at Chapel Hill and published in 2016, remains one of the most highly cited in the field of artificial intelligence (AI). It describes a new method for detecting objects in images or video footage using a single deep neural network – a set of AI algorithms inspired by the neurological processes that fire in the human cerebral cortex.
The approach, called the Single Shot MultiBox Detector, or SSD, has been described as faster than Faster R-CNN – another object detection technology that was described in a very highly cited paper published in 2015 (see our coverage here).
SSD works by dividing the image into a grid, with each grid cell responsible for detecting objects within that part of the image. As the name indicates, the network is able to identify all objects within an image in a single pass, allowing for real-time analysis.
SSD is now one of a handful of object detection technologies that are now available. YOLO (You Only Look Once) is a similar single-shot object detection algorithm, whereas R-CNN and Faster R-CNN use a two-step approach, which involves first identifying the regions where objects might be, and then detecting them.
‘Analysis of protein-coding genetic variation in 60,706 humans’
7,696 citations
Led by Monkol Lek from the University of Sydney in Australia and Daniel MacArthur from the Broad Institute of MIT and Harvard University, this 2016 paper presents an open-access catalogue of more than 60,000 human exome sequences (exomes are the coding portions of genes) from people of European, African, South Asian, East Asian, and Latinx ancestry.
The collection was compiled as part of the Exome Aggregation Consortium project, run by an international group of researchers with a focus on exome sequencing. As exomes only make up about 2% of the human genome, the approach has been praised for being able to highlight patterns of genetic variation, including known disease-related variants, in a more cost-effective way than whole-genome sequencing.
Presented at a 2015 genomics conference, the catalogue encompasses 7.4 million genetic variants, which can be used to identify those connected to rare diseases. “Large-scale reference datasets of human genetic variation are critical for the medical and functional interpretation of DNA sequence changes,” Lek said when the paper was published.
